Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 54(5); 2020 > Article
-
Original Article
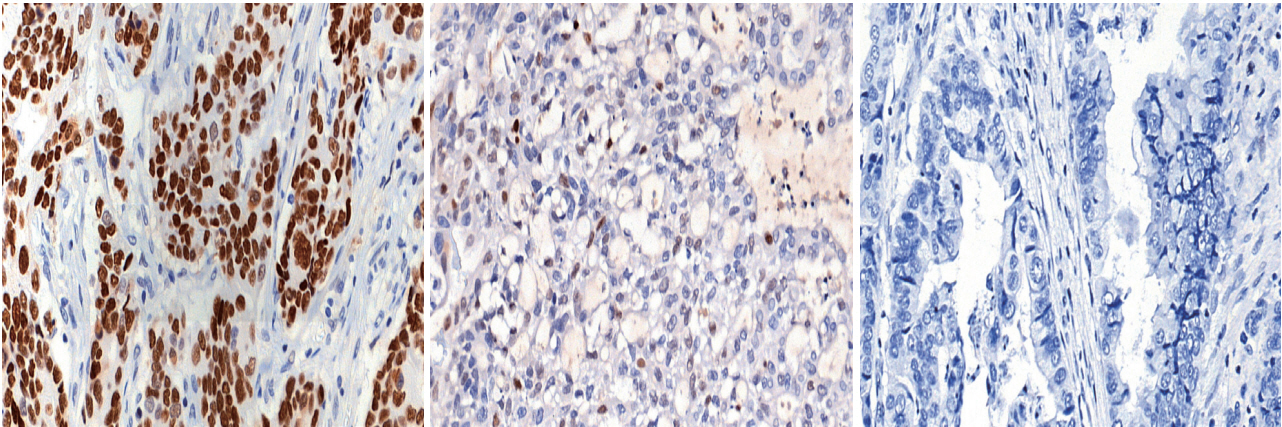
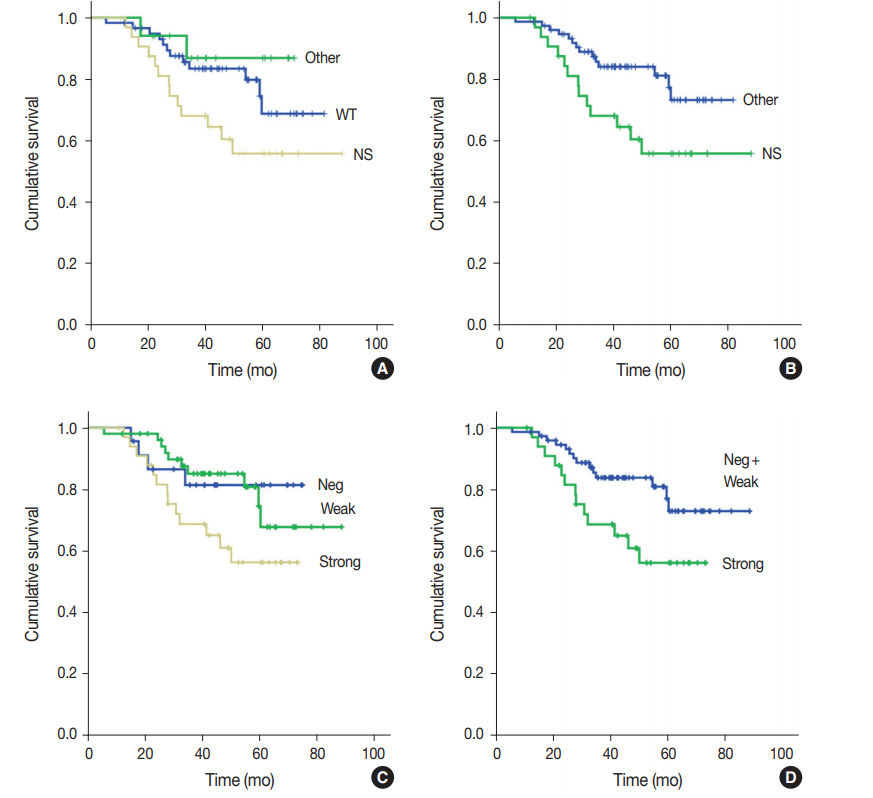
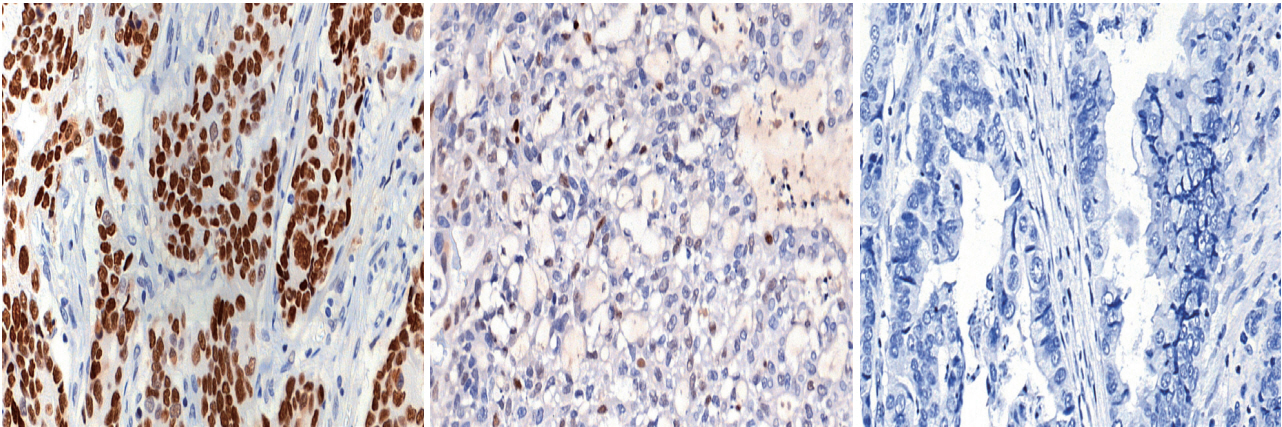
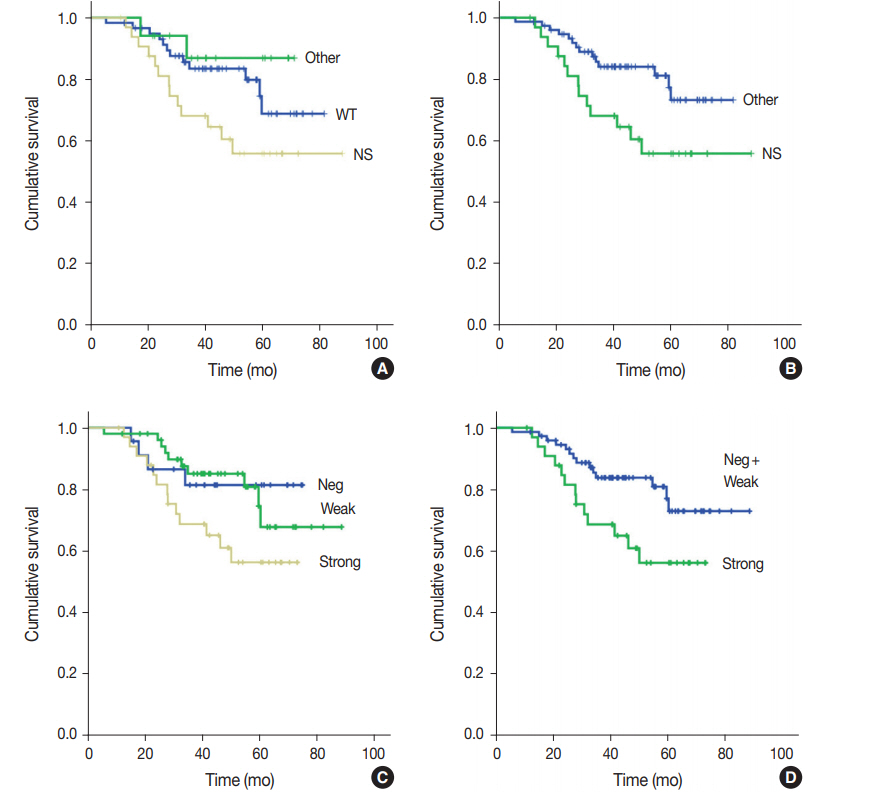
Prediction of TP53 mutations by p53 immunohistochemistry and their prognostic significance in gastric cancer -
Hye Jung Hwang1
 , Soo Kyung Nam1
, Soo Kyung Nam1 , Hyunjin Park2
, Hyunjin Park2 , Yujun Park1
, Yujun Park1 , Jiwon Koh3
, Jiwon Koh3 , Hee Young Na1
, Hee Young Na1 , Yoonjin Kwak3
, Yoonjin Kwak3 , Woo Ho Kim3
, Woo Ho Kim3 , Hye Seung Lee,1
, Hye Seung Lee,1
-
Journal of Pathology and Translational Medicine 2020;54(5):378-386.
DOI: https://doi.org/10.4132/jptm.2020.06.01
Published online: July 1, 2020
1Department of Pathology, Seoul National University Bundang Hospital, Seoul National University College of Medicine, Seongnam, Korea
2Department of Pathology, Gangnam Severance Hospital, Yonsei University College of Medicine, Seoul, Korea
3Department of Pathology, Seoul National University Hospital, Seoul National University College of Medicine, Seoul, Korea
- Corresponding Author: Hye Seung Lee, MD, PhD, Department of Pathology, Seoul National University Bundang Hospital, Seoul National University College of Medicine, 82 Gumi-ro 173beon-gil, Bundang-gu, Seongnam 13620, Korea Tel: +82-31-787-7714, Fax: +82-31-787-4012, E-mail: 'hye2@snu.ac.kr'
• Received: March 30, 2020 • Revised: May 22, 2020 • Accepted: June 1, 2020
© 2020 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations
Citations to this article as recorded by 

- Multiple approaches revealed MGc80‐3 as a somatic hybrid with HeLa cells rather than a gastric cancer cell line
Fang Cao, Hao Sun, Zhenli Yang, Yanhua Bai, Xiao Hu, Yuhong Hou, Xiaocui Bian, Yuqin Liu
International Journal of Cancer.2024; 154(1): 155. CrossRef - In Response to p53 Immunohistochemical Staining and TP53 Gene Mutations in Endometrial Cancer: Does Null Pattern Correlate With Prognosis?
Ikuko Sakamoto, Keiko Kagami, Takahiro Nozaki, Yosuke Hirotsu, Kenji Amemiya, Toshio Oyama, Masao Omata
American Journal of Surgical Pathology.2024; 48(3): 374. CrossRef - CHEK2 germline variants identified in familial nonmedullary thyroid cancer lead to impaired protein structure and function
Carolina Pires, Inês J. Marques, Mariana Valério, Ana Saramago, Paulo E. Santo, Sandra Santos, Margarida Silva, Margarida M. Moura, João Matos, Teresa Pereira, Rafael Cabrera, Diana Lousa, Valeriano Leite, Tiago M. Bandeiras, João B. Vicente, Branca M. Ca
Journal of Biological Chemistry.2024; 300(3): 105767. CrossRef - The spectrum of TP53 mutations in Rwandan patients with gastric cancer
Augustin Nzitakera, Jean Bosco Surwumwe, Ella Larissa Ndoricyimpaye, Schifra Uwamungu, Delphine Uwamariya, Felix Manirakiza, Marie Claire Ndayisaba, Gervais Ntakirutimana, Benoit Seminega, Vincent Dusabejambo, Eric Rutaganda, Placide Kamali, François Ngab
Genes and Environment.2024;[Epub] CrossRef - Application of NGS molecular classification in the diagnosis of endometrial carcinoma: A supplement to traditional pathological diagnosis
Qunxian Rao, Jianwei Liao, Yangyang Li, Xin Zhang, Guocai Xu, Changbin Zhu, Shengya Tian, Qiuhong Chen, Hui Zhou, Bingzhong Zhang
Cancer Medicine.2023; 12(5): 5409. CrossRef - Predictive value of p53 and AXL immunostaining for the efficacy of immune checkpoint inhibitor-based therapy after osimertinib treatment in patients with epidermal growth factor-mutant non-small cell lung cancer
Kenji Morimoto, Tadaaki Yamada, Ryo Sawada, Koichi Azuma, Yasuhiro Goto, Taishi Harada, Shinsuke Shiotsu, Nobuyo Tamiya, Yusuke Chihara, Takayuki Takeda, Osamu Hiranuma, Isao Hasegawa, Satomi Tanaka, Akihiro Yoshimura, Masahiro Iwasaku, Shinsaku Tokuda, Y
Cancer Immunology, Immunotherapy.2023; 72(6): 1699. CrossRef - Validation of p53 Immunohistochemistry (PAb240 Clone) in Canine Tumors with Next-Generation Sequencing (NGS) Analysis
Barbara Brunetti, Dario de Biase, Giulia Dellapina, Luisa Vera Muscatello, Francesco Ingravalle, Giorgia Tura, Barbara Bacci
Animals.2023; 13(5): 899. CrossRef - Mesonephric‐like adenocarcinoma of the female genital tract: novel observations and detailed molecular characterisation of mixed tumours and mesonephric‐like carcinosarcomas
Jelena Mirkovic, Ekaterina Olkhov‐Mitsel, Yutaka Amemiya, Maysa Al‐Hussaini, Sharon Nofech‐Mozes, Bojana Djordjevic, Rachel Kupets, Arun Seth, W Glenn McCluggage
Histopathology.2023; 82(7): 978. CrossRef - Clinicopathologic characterization of cervical metastasis from an unknown primary tumor: a multicenter study in Korea
Miseon Lee, Uiree Jo, Joon Seon Song, Youn Soo Lee, Chang Gok Woo, Dong-Hoon Kim, Jung Yeon Kim, Sun Och Yoon, Kyung-Ja Cho
Journal of Pathology and Translational Medicine.2023; 57(3): 166. CrossRef - P53 in Penile Squamous Cell Carcinoma: A Pattern-Based Immunohistochemical Framework with Molecular Correlation
Isabel Trias, Adela Saco, Lorena Marimon, Ricardo López del Campo, Carolina Manzotti, Oriol Ordi, Marta del Pino, Francisco M. Pérez, Naiara Vega, Silvia Alós, Antonio Martínez, Leonardo Rodriguez-Carunchio, Oscar Reig, Pedro Jares, Cristina Teixido, Tare
Cancers.2023; 15(10): 2719. CrossRef - p53/TP53 Status Assessment in Gastroesophageal Adenocarcinoma
Elisa Boldrin, Maria Assunta Piano, Francesco Bernaudo, Rita Alfieri, Maria Raffaella Biasin, Isabella Monia Montagner, Alice Volpato, Genny Mattara, Francesco Lamacchia, Giovanna Magni, Antonio Rosato, Antonio Scapinello, Pierluigi Pilati, Matteo Curtare
Cancers.2023; 15(10): 2783. CrossRef - Genomic profiling of dedifferentiated endometrial carcinomas arising in the background of high‐grade carcinoma: a targeted next‐generation sequencing study
Ekaterina Olkhov‐Mitsel, Aurelia Busca, Carlos Parra‐Herran, Yutaka Amemiya, Sharon Nofech‐Mozes, Bojana Djordjevic, Marisa R Nucci, Arun Seth, Jelena Mirkovic
Histopathology.2023; 83(3): 366. CrossRef -
Clinicopathologic Features and Prognostic Significance of Immunohistochemistry and In Situ Hybridization Based Molecular Classification in Gastric Carcinoma
Gizem Issin, İlyas Sayar, Fatih Demir, İrem Güvendir Bakkaloğlu, Mehmet Gamsizkan, Zeliha Yildiz, Ismail Yilmaz, Sevilay Akalp Özmen, Diren Vuslat Çağatay, Itır Ebru Zemheri, Murat Demiriz, Armağan Günal
Journal of Environmental Pathology, Toxicology and Oncology.2023; 42(4): 1. CrossRef - Clinicopathologic and Molecular Characterization of Anorectal Neuroendocrine Carcinomas Reveals Human Papillomavirus, p53, and c-Myc as Alternative Mechanisms of Carcinogenesis
Allison J. Cox, William E. Crowe, Qi Yang, Bin Zhang, Zoltán N. Oltvai, Xiaoyan Liao
Modern Pathology.2023; 36(11): 100295. CrossRef - Dedifferentiated Endometrial Carcinoma: A Rare Aggressive Neoplasm-Clinical, Morphological and Immunohistochemical Features
Giovanna Giordano, Elena Ferioli, Debora Guareschi, Alessandro Tafuni
Cancers.2023; 15(21): 5155. CrossRef - Characterization on the oncogenic effect of the missense mutations of p53 via machine learning
Qisheng Pan, Stephanie Portelli, Thanh Binh Nguyen, David B Ascher
Briefings in Bioinformatics.2023;[Epub] CrossRef - Adrenal Nodules Detected at Staging CT in Patients with Resectable Gastric Cancers Have a Low Incidence of Malignancy
Hae Young Kim, Won Chang, Yoon Jin Lee, Ji Hoon Park, Jungheum Cho, Hee Young Na, Hyungwoo Ahn, Sung Il Hwang, Hak Jong Lee, Young Hoon Kim, Kyoung Ho Lee
Radiology.2022; 302(1): 129. CrossRef - Intestinal-type gastric dysplasia in Helicobacter pylori-naïve patients
Kotaro Shibagaki, Ayako Itawaki, Yoichi Miyaoka, Kenichi Kishimoto, Yusuke Takahashi, Satoshi Kotani, Tsuyoshi Mishiro, Naoki Oshima, Kousaku Kawashima, Norihisa Ishimura, Hideyuki Onuma, Makoto Nagasaki, Mamiko Nagase, Asuka Araki, Kyuichi Kadota, Ryoji
Virchows Archiv.2022; 480(4): 783. CrossRef - Dedifferentiation-like tubular and solid carcinoma of the stomach shows phenotypic divergence and association with deficient SWI/SNF complex
Shih-Chiang Huang, Kuang-Hua Chen, Kwai-Fong Ng, I-Chieh Lin, Yi-Chun Chao, Ta-Sen Yeh, Huei-Chieh Chuang, Tse-Ching Chen
Virchows Archiv.2022; 480(4): 771. CrossRef - Distinct molecular phenotype and the potential prognostic value of immune prognostic index and tumor infiltrating lymphocytes in hepatoid adenocarcinoma of stomach
Muxing Kang, Xiaojing Ma, Jifei Shi, Guofeng Chen, Xiaoli Jin, Jun Wang, Lele Lin, Zhiwei Wu, Kaibo Chen, Jinghong Xu, Pintong Huang, Jian Chen
Translational Oncology.2022; 19: 101380. CrossRef - Evaluation of Tumor DNA Sequencing Results in Patients with Gastric and Gastroesophageal Junction Adenocarcinoma Stratified by TP53 Mutation Status
Anthony C Wood, Yonghong Zhang, Qianxing Mo, Ling Cen, Jacques Fontaine, Sarah E Hoffe, Jessica Frakes, Sean P Dineen, Jose M Pimiento, Christine M Walko, Rutika Mehta
The Oncologist.2022; 27(4): 307. CrossRef - Comprehensive Clinical Analysis of Gallbladder Neuroendocrine Neoplasms: A Large-Volume Multicenter Study During One Decade
Yangyang Wang, Bingfeng Huang, Qihan Fu, Jianing Wang, Mao Ye, Manyi Hu, Kai Qu, Kai Liu, Xiao Hu, Shumei Wei, Ke Sun, Wenbo Xiao, Bo Zhang, Haijun Li, Jingsong Li, Qi Zhang, Tingbo Liang
Annals of Surgical Oncology.2022; 29(12): 7619. CrossRef - Expression of SASP, DNA Damage Response, and Cell Proliferation Factors in Early Gastric Neoplastic Lesions: Correlations and Clinical Significance
Li Liang, Yijie Chai, Fei Chai, Haijing Liu, Ningning Ma, Hong Zhang, Shuang Zhang, Lin Nong, Ting Li, Bo Zhang
Pathology and Oncology Research.2022;[Epub] CrossRef - Systems biology and OMIC data integration to understand gastrointestinal cancers
Iasmin Moreira Costa Bispo, Henry Paul Granger, Palloma Porto Almeida, Patricia Belini Nishiyama, Leandro Martins de Freitas
World Journal of Clinical Oncology.2022; 13(10): 762. CrossRef - MicroRNA-552 expression in colorectal cancer and its clinicopathological significance
Joon Im, Soo Kyung Nam, Hye Seung Lee
Journal of Pathology and Translational Medicine.2021; 55(2): 125. CrossRef - Different effects of p53 protein overexpression on the survival of gastric cancer patients according to Lauren histologic classification: a retrospective study
Ki Wook Kim, Nayoung Kim, Yonghoon Choi, Won Seok Kim, Hyuk Yoon, Cheol Min Shin, Young Soo Park, Dong Ho Lee, Young Suk Park, Sang-Hoon Ahn, Do Joong Park, Hyung-Ho Kim, Hye Seung Lee, Ji-Won Kim, Jin Won Kim, Keun-Wook Lee, Won Chang, Ji Hoon Park, Yoon
Gastric Cancer.2021; 24(4): 844. CrossRef - The association between the expression of nuclear Yes-associated protein 1 (YAP1) and p53 protein expression profile in breast cancer patients
Yoon Jin Cha, Dooreh Kim, Soong June Bae, Sung Gwe Ahn, Joon Jeong, Min Kyung Cho, Pill Sun Paik, Tae-Kyung Yoo, Woo-Chan Park, Chang Ik Yoon, Elda Tagliabue
PLOS ONE.2021; 16(5): e0250986. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure

 E-submission
E-submission