Analysis of the molecular subtypes of preoperative core needle biopsy and surgical specimens in invasive breast cancer
Article information
Abstract
Background
Accurate molecular classification of breast core needle biopsy (CNB) tissue is important for determining neoadjuvant systemic therapies for invasive breast cancer. The researchers aimed to evaluate the concordance rate (CR) of molecular subtypes between CNBs and surgical specimens.
Methods
This study was conducted with invasive breast cancer patients who underwent surgery after CNB at Seoul St. Mary’s Hospital between December 2014 and December 2017. Estrogen receptor (ER), progesterone receptor (PR), human epidermal growth factor receptor 2 (HER2), and Ki67 were analyzed using immunohistochemistry. ER and PR were evaluated by Allred score (0–8). HER2 was graded from 0 to +3, and all 2+ cases were reflex tested with silver in situ hybridization. The labeling index of Ki67 was counted by either manual scoring or digital image analysis. Molecular subtypes were classified using the above surrogate markers.
Results
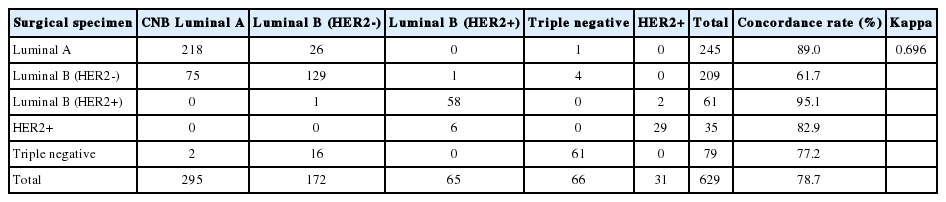
In total, 629 patients were evaluated. The CRs of ER, PR, HER2, and Ki67 were 96.5% (kappa, 0.883; p<.001), 93.0% (kappa, 0.824; p<.001), 99.7% (kappa, 0.988; p<.001), and 78.7% (kappa, 0.577; p<.001), respectively. Digital image analysis of Ki67 in CNB showed better concordance with Ki67 in surgical specimens (CR, 82.3%; kappa, 0.639 for digital image analysis vs. CR, 76.2%; kappa, 0.534 for manual counting). The CRs of luminal A, luminal B, HER2, and triple negative types were 89.0%, 70.0%, 82.9%, and 77.2%, respectively.
Conclusions
CNB was reasonably accurate for determining ER, PR, HER2, Ki67, and molecular subtypes. Using digital image analysis for Ki67 in CNB produced more accurate molecular classifications.
Breast cancer is a heterogeneous disease that is classified into four molecular subtypes: luminal A, luminal B, human epidermal growth factor receptor 2 (HER2) positive, and triple negative. Proper disease subtyping is important for predicting response to systemic therapy and prognosis. Immunohistochemistry for estrogen receptor (ER), progesterone receptor (PR), HER2, and Ki67 can be used to approximate molecular subtype classifications [1-3].
Endocrine therapy, cytotoxic therapy, and anti-HER2 therapy are used in the adjuvant setting for breast cancer, and molecular subtyping is used to decide which therapies are appropriate [3]. Neoadjuvant therapy is important in advanced breast cancer as it can reduce tumor size, facilitating breast conserving surgery, and decrease distant micrometastasis. Neoadjuvant therapy may also facilitate axillary preservation [4-7]. After receiving neoadjuvant therapy, some patients achieve pathologic complete response (pCR), at which point, core needle biopsy (CNB) tissue is the only specimen for obtaining information about prognostic markers that could impact therapeutic plans. Although some patients still have tumor burden after neoadjuvant therapy, CNB might be the only specimen available in which to evaluate histological markers because neoadjuvant therapy can alter histological grade, expression of hormonal receptors, HER2 status, and Ki67. Consequently, assessing the concordance rate (CR) of marker status between CNB and surgical specimen is crucial [8,9]. It has been reported that CRs of biomarker status between CNB and surgical specimen were high but variable: 79%–100% [10]. However, differences in Ki67 status according to manual or digital image analysis and the impact on subsequent molecular subtyping have not been adequately investigated. The aim of this study was to evaluate and analyze CRs of hormone receptors, HER2, Ki67, and molecular subtypes in invasive breast cancer patients between CNB and surgical specimens.
MATERIALS AND METHODS
Patient population and tissue samples
This study retroactively analyzed clinical data from cancer patients who underwent surgery after diagnosis by CNB at Seoul St. Mary’s Hospital. Clinicopathologic data of age; histologic type; grade; operation method; pTNM stage; immunohistochemical staining results of ER, PR, HER2, and Ki67; and whether digital image analysis was used to analyze results of Ki67 staining and silver in situ hybridization (SISH) for HER2 were obtained through medical records and pathologic reports. CNBs were mostly performed by ultrasound guidance with 14-gauge needles; 4–5 pieces were received for mass forming lesions and >7 pieces for microcalcifications. Especially when microcalcification was noted, an ultrasound-guided or stereotactic mammotome biopsy with 11-gauge needles was performed, and at least 12 pieces of core tissue were obtained for pathologic examination. All specimens were routinely processed and diagnosed according to national and international guidelines. Briefly, CNB specimens were fixed in 10% neutral formalin for 8–11 hours, embedded in paraffin, and sectioned at 4-μm thickness; hematoxylin and eosin (H&E) staining was subsequently performed. Surgical specimens were cut into 0.5–1-cm-thick sections after the operation and fixed in 10% neutral formalin for 8–24 hours; H&E staining was subsequently performed. Patients who were diagnosed with ductal carcinoma in situ, microinvasive breast cancer, or had received preoperative systemic therapy were excluded.
Evaluation of ER, PR, HER2, and Ki67
Immunohistochemistry (IHC) for ER, PR, HER2, and Ki67 was performed following the instructions of the pathology laboratory manual. Briefly, ER, PR, HER2, and Ki67 IHC staining was performed on an automated Ventana BenchmarkXT slide stainer (Ventana, Tucson, AZ, USA), using primary antibodies against ER (prediluted, SP1, Ventana), PR (prediluted, 1E2, Ventana), HER2 (prediluted, 4B5, Ventana), and Ki67 (prediluted, MIB-1, Ventana). The HER2 SISH assay was performed with INFORM HER2 DNA probes (Ventana) on the Ventana BenchMarkXT automated slide stainer according to the manufacturer’s protocols.
The Allred scoring system was used to interpret ER and PR staining [11]. The proportion of positive-stained tumor cells (the proportion score) was rated as follows: 0, no cells stained positive; 1, 0%–1% positive; 2, 1%–10% positive; 3, 10%–33% positive; 4, 33%–66% positive; and 5, 66%–100% positive. Intensity was scored on the basis of average staining intensity: 0, negative; 1, weak; 2, intermediate; and 3, strong. The sum of the proportion and intensity scores is referred to as the Allred score, and scores >2 are defined as positive [11]. HER2 status was scored from 0–3+ by IHC, where 0 was defined as no staining or membrane staining that was incomplete, faint, or barely perceptible in ≤10% of invasive tumor cells; 1+ was defined as >10% invasive tumor cells with incomplete membrane staining that was faint or barely perceptible; 2+ was defined as >10% invasive tumor cells with complete weak to moderate membrane staining (considered equivocal); and 3+ was defined as >10% invasive tumor cells with complete, intense membrane staining (positive according to American Society of Clinical Oncology [ASCO]/College of American Pathologists [CAP] guidelines) [12,13]. Cases of scores 0 and 1+ were considered negative, while those 2+ were further evaluated by reflex HER2 SISH to confirm HER2 gene amplification.
The Allred scoring system was used to interpret ER and PR staining [11]. The proportion of positive-stained tumor cells (the proportion score) was rated as follows: 0, no cells stained positive; 1, 0%–1% positive; 2, 1%–10% positive; 3, 10%–33% positive; 4, 33%–66% positive; and 5, 66%–100% positive. Intensity was scored on the basis of average staining intensity: 0, negative; 1, weak; 2, intermediate; and 3, strong. The sum of the proportion and intensity scores is referred to as the Allred score, and scores >2 are defined as positive [11]. HER2 status was scored from 0–3+ by IHC, where 0 was defined as no staining or membrane staining that was incomplete, faint, or barely perceptible in ≤10% of invasive tumor cells; 1+ was defined as >10% invasive tumor cells with incomplete membrane staining that was faint or barely perceptible; 2+ was defined as >10% invasive tumor cells with complete weak to moderate membrane staining (considered equivocal); and 3+ was defined as >10% invasive tumor cells with complete, intense membrane staining (positive according to American Society of Clinical Oncology [ASCO]/College of American Pathologists [CAP] guidelines) [12,13]. Cases of scores 0 and 1+ were considered negative, while those 2+ were further evaluated by reflex HER2 SISH to confirm HER2 gene amplification.The Allred scoring system was used to interpret ER and PR staining [11]. The proportion of positive-stained tumor cells (the proportion score) was rated as follows: 0, no cells stained positive; 1, 0%–1% positive; 2, 1%–10% positive; 3, 10%–33% positive; 4, 33%–66% positive; and 5, 66%–100% positive. Intensity was scored on the basis of average staining intensity: 0, negative; 1, weak; 2, intermediate; and 3, strong. The sum of the proportion and intensity scores is referred to as the Allred score, and scores >2 are defined as positive [11]. HER2 status was scored from 0–3+ by IHC, where 0 was defined as no staining or membrane staining that was incomplete, faint, or barely perceptible in ≤10% of invasive tumor cells; 1+ was defined as >10% invasive tumor cells with incomplete membrane staining that was faint or barely perceptible; 2+ was defined as >10% invasive tumor cells with complete weak to moderate membrane staining (considered equivocal); and 3+ was defined as >10% invasive tumor cells with complete, intense membrane staining (positive according to American Society of Clinical Oncology [ASCO]/College of American Pathologists [CAP] guidelines) [12,13]. Cases of scores 0 and 1+ were considered negative, while those 2+ were further evaluated by reflex HER2 SISH to confirm HER2 gene amplification.
Ki67 was evaluated according to the percentage of positively-stained invasive tumor cells of any intensity by pathologists manually or using an automated digital image analysis system. The “eyeballed” estimation method, which is approximate counting throughout the immunostained slide, was used for manual counting [14]. For Ki67 digital image analysis, slides were scanned by an iScan Coreo slide scanner with a 20× objective (Ventana), and invasive tumor components were analyzed using Virtuoso software (Ventana). At least three high-power fields (400×) including hot spots where the highest Ki67 staining area and two average intensity areas of Ki67 staining were selected. More than 1,000 tumor cells were counted according to the International Ki67 in Breast Cancer Working Group [15]. In the case of CNB with a heterogeneous Ki67 staining pattern, researchers attempted to select all tumor cells to evaluate Ki67 expression. Cases in which less than 500 tumor cells were present in CNB were excluded from digital image analysis. Digital image analysis of Ki67 was performed in 41% (260/629) of CNBs and all but 14 surgical specimens. The cutoff value for Ki67 was 20%.
Molecular subtype classifications
Molecular subtypes were classified as follows: luminal A: ER- and/or PR-positive, HER2-negative, Ki67 ≤20%; luminal B: ER- and/or PR-positive, HER2-negative, Ki67 >20% or ER- and/or PR-positive, HER2-positive, any Ki67; HER2-positive: ER- and PR-negative, HER2-positive; and triple-negative (basallike): ER-, PR-, and HER2-negative. Subsequently, luminal B was further divided as luminal B–HER2–negative (ER- and/or PR-positive, HER2-negative, Ki67 >20%) and luminal B–HER2–positive (ER- and/or PR-positive, HER2-positive, any Ki67) [2,3].
Statistical analysis
CRs of receptor status and molecular subtypes between CNB and surgical specimens were calculated as percentage and Kappa value. K-values <0.20 were correlated with poor agreement, 0.21–0.40 fair agreement, 0.41–0.60 moderate agreement, 0.61–0.80 good agreement, and 0.81–1.00 very good agreement. p-values were calculated using the chi-square test or Fisher exact test, and p-values of <0.05 were considered significant. Statistical analyses were performed using IBM SPSS Statistics software ver. 24.0 for Windows (IBM Corp., Armonk, NY, USA). Sankey diagrams depicting changes in Allred scores were computed in SankeyMATIC (http://sankeymatic.com).
Ethics statement
The protocol of the study was approved by the Institutional Review Board of The Catholic Medical Center, which waived the requirement for informed consent (approval KC19RESI0333).
RESULTS
Patient characteristics
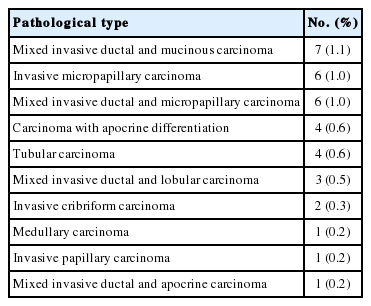
In total, 629 patients were included in the study. The median age was 53 (range, 23 to 89 years). All patients received either breast conserving surgery (70.4%) or mastectomy (29.6%). The most common pathological tumor type was invasive carcinoma of no special type (80.1%) (Tables 1, 2).
Concordance of ER, PR, HER2, and Ki67
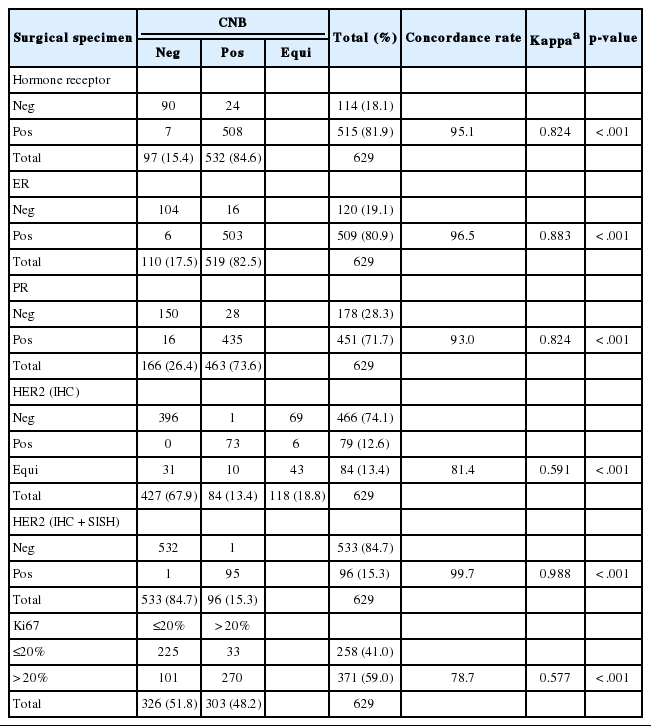
The CRs of surgical specimens with CNBs for ER and PR were 96.5% (kappa, 0.883; p<.001) and 93.0% (kappa, 0.824; p<.001), respectively (Table 3). The concordance of PR was slightly lower than that of ER. Changes in Allred score for ER and PR are shown in Figs. 1 and 2. The CR of surgical specimens with CNBs for hormone receptor status was 95.1%, with very good agreement (kappa, 0.824; p<.001). The CR of HER2 IHC was 81.4%, with moderate agreement (kappa, 0.591; p< .001). After reflex HER2 SISH, the CR of HER2 status was 99.7% (kappa, 0.988; p<.001). The CR for Ki67 was 78.7% (kappa, 0.577; p<.001, moderate agreement), which was lower than for hormone receptors and HER2 due to higher Ki67 expression in surgical specimens (Table 3).

Sankey diagrams depicting changes in Allred scores for estrogen receptor (ER) from core needle biopsy (CNB) to surgical specimen. EB, excisional biopsy (surgical specimen).
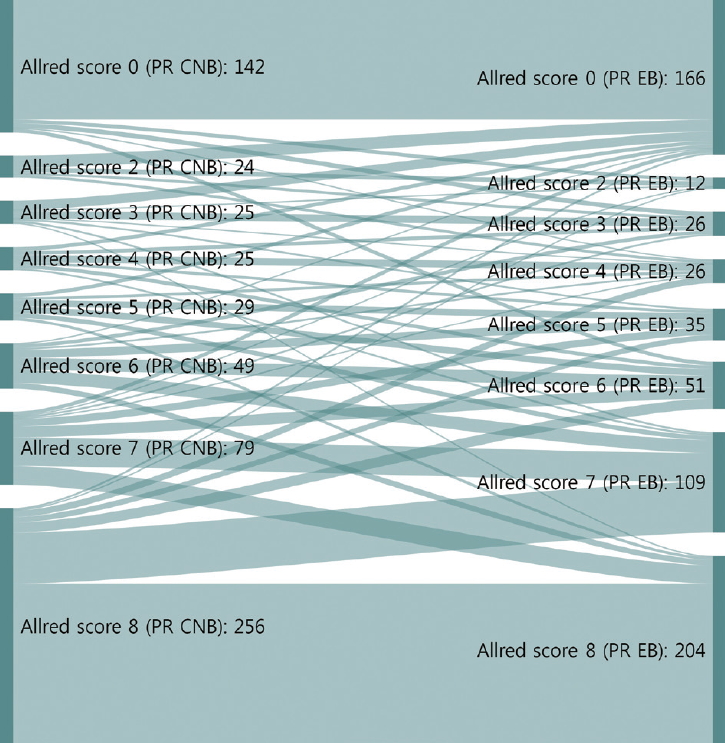
Sankey diagrams depicting changes in Allred scores for progesterone receptor (PR) from core needle biopsy (CNB) to surgical specimen. EB, excisional biopsy (surgical specimen).
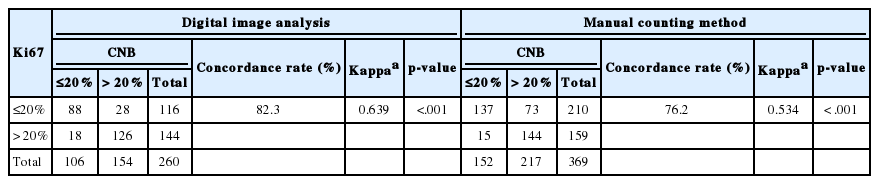
We next analyzed whether the Ki67 counting method (automated digital image analysis system or manual scoring) affected the CR for Ki67. Among the 629 cases, 260 CNBs were analyzed with a digital image system and 369 with manual counting. When CNB Ki67s were obtained using the digital image analysis system, the CR for Ki67 was 82.3% (kappa, 0.639; p < .001), and when CNB Ki67s were obtained by manual counting, the CR for Ki67 was 76.2% (kappa, 0.534; p<.001) (Table 4).
Concordance of molecular subtypes
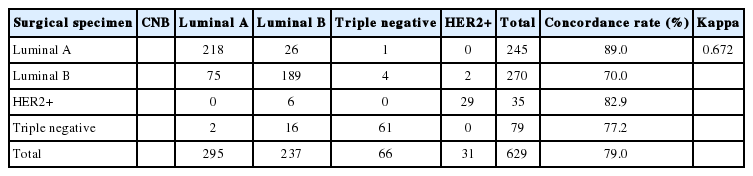
Molecular subtypes demonstrated good agreement (kappa, 0.672 in four subtypes and kappa, 0.696 in five subtypes). CRs of surgical specimens with CNBs for luminal A, luminal B, HER2, and triple negative were 89.0%, 70.0%, 82.9%, and 77.2%, respectively (kappa, 0.672; good agreement). The CRs of luminal A and HER2 showed significant agreement; however, luminal B showed the lowest CR among the molecular subtypes. When luminal B was subdivided into luminal B–HER2–negative and –HER2–positive, the CRs for these receptors between CNB and surgical specimens were 61.7% and 95.1%, respectively. Therefore, the especially high discrepancy seen for luminal B–HER2–negative occurred because molecular subtypes changed from luminal A due to low Ki67 in CNB (75 cases) to luminal B–HER2–negative after high Ki67 was found in surgical specimens (Tables 5, 6).
DISCUSSION
Neoadjuvant therapy in breast cancer can reduce tumor burden, which facilitates breast conserving surgery, preserves the axilla, and identifies response to systemic therapy before surgery [5-7]. Typically, treatment policies for neoadjuvant systemic therapy are decided according to the molecular subtype determined by ER, PR, HER2, and Ki67 status of CNB. These are also used to predict prognoses of breast cancer patients. In this study, the CRs of ER, PR, HER2, and Ki67 status in CNB specimens prior to surgery and those from surgical specimens were analyzed to confirm the accuracy of molecular classifications performed by CNB. ER and PR status showed high CRs of 96.5% (kappa, 0.883) and 93.0% (kappa, 0.824), respectively (Table 3). However, a greater distribution and bigger change of Allred scores from CNB to surgical specimen were noted for PR than ER (Figs. 1, 2). These findings are consistent with previous reports [16,17] and suggest that the reason behind these findings is heterogeneity of PR expression in tumor cells.
Identifying HER2-positive breast cancer patients by CNB is important, as neoadjuvant HER2-targeted therapy is an effective option for these patients. In this study, the CR for HER2 status, as determined by HER2 IHC or reflex HER2 SISH, was as high as 99.7% (kappa, 0.988), which was higher than most previous reports (ranging from 61%–97.3%) [10,18-29]. The reason for this high CR could partially be due to performing reflex HER2 SISH for all HER2 IHC equivocal cases to determine final HER2 status strictly following ASCO/CAP guidelines. Sufficient CNB specimens were obtained when radiological microcalcifications were noted. In our hospital, at least four core passes in CNB are usually obtained by ultrasound guidance with a 14-gauge needle. However, when radiological calcifications were identified, additional core passes (at least seven) were performed. Sometimes when scattered calcification was noted, ultrasound-guided mammotome biopsy was performed with an 11-gauge needle, and >12 core passes were obtained. It has been reported that radiologically recognized calcifications in breast cancer are associated with HER2 molecular subtype and pCR after neoadjuvant chemotherapy [30,31]. Another reason for collection of multiple cores in CNB is to overcome tumor heterogeneity. For accurate histologic diagnoses of breast cancer, at least four cores should be obtained using a 14-gauge needle [32]. It has been reported that the accuracy of histologic diagnoses (including tumor grade) plateaus at 74% when four passes are performed, while accuracy is only 32% with one pass [33]. Greer et al. [34] reported that the concordance for ER and PR between CNB and surgical specimen improved with increasing numbers of core passes. However, they also found the concordance of HER2 to be limited when tumor heterogeneity was present [34].
Although ER, PR, and HER2 status showed high CRs between CNB and surgical specimen, Ki67 revealed only moderate agreement. There were 75 cases classified as luminal A from CNB that were moved to luminal B after evaluating surgical specimens, as Ki67 was higher in the surgical specimen than in CNB. Additionally, a higher median value for Ki67 was identified in the surgical specimens (Tables 3, 5, 6). The tendency for a greater Ki67 labeling index in surgical specimens than in CNB has been reported in several studies. The authors explained that it was due to tumor heterogeneity [15,27]. Another study reported that the CR for Ki67 between CNB and surgical specimen improved slightly with increased number of core passes to account for tumor heterogeneity; however, after more than six core passes, a plateau was reached [34]. In the current study, the Ki67 labeling index was obtained by digital image analysis or manual scoring (Table 4). Digital image analysis of Ki67 in CNB showed better concordance with surgical specimen Ki67 (CR, 82.3% vs. 76.2; kappa, 0.639 vs. 0.534, respectively).
Finally, 21.0% of cases switched molecular subtype between CNB and surgical specimen, mostly because Ki67 changed from low to high; for example, 75 luminal A cases as assessed by CNB were moved to luminal B after evaluating surgical specimens (Table 5). There were also a few cases where hormone status changed from positive to negative, 16 cases of luminal B changed to triple negative, and six cases changed to HER2. The reason for these discrepancies could be fixation conditions such as delayed, under-, or over-fixation with formalin; crush artifacts by needle sampling; or tumor heterogeneity.
In conclusion, CNB with adequate core passes can be reliably used to access molecular subtypes for systemic treatment in invasive breast cancer. Digital image analysis of Ki67 should be used to achieve better, more accurate molecular classifications from CNBs.
Notes
Authors’ contribution
Conceptualization: AL.
Data curation: YSJ, TKY.
Formal analysis: YSJ, SHK, AL.
Methodology: YSJ, SHK, JK, AL.
Project administration: YSJ.
Resources: AL.
Supervision: AL.
Validation: YSJ, JL, AL.
Visualization: YSJ.
Writing—original draft: YSJ.
Writing—review & editing: YSJ, SHK, JK, AL.
Conflicts of Interest: The authors declare that they have no potential conflicts of interest.
Funding
No funding to declare.