Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 55(6); 2021 > Article
-
Letter to the Editor
Fusobacterium nucleatum: caution with interpreting historical patient sample cohort -
Kate L. F. Johnstone1
 , Sinead Toomey1
, Sinead Toomey1 , Stephen Madden2
, Stephen Madden2 , Brian D. P. O’Neill3, Bryan T Hennessy1
, Brian D. P. O’Neill3, Bryan T Hennessy1
-
Journal of Pathology and Translational Medicine 2021;55(6):415-418.
DOI: https://doi.org/10.4132/jptm.2021.08.27
Published online: September 27, 2021
1Medical Oncology Laboratory, Department of Molecular Medicine, Royal College of Surgeons in Ireland, Beaumont Hospital, Dublin, Ireland
2Data Science Centre, Royal College of Surgeons in Ireland, Dublin, Ireland
3St Luke’s Radiation Oncology Network, Dublin, Ireland
- Corresponding Author: Kate L. F. Johnstone, MBChB Medical Oncology Laboratory, Department of Molecular Medicine, Royal College of Surgeons in Ireland, Smurfit Building, Beaumont Hospital, Dublin 9, Ireland Tel: +353-1-809-3823, Fax: +353-1-809-3778, E-mail: katejohnstone@rcsi.com
© 2021 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (https://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
- 4,724 Views
- 104 Download
Ethics Statement
Tumor samples were obtained from patients enrolled on the TRI-LARC clinical trial (NCT02151019). Historical tumor samples were obtained from tumor banks under the auspices of Institutional Review Board-approved protocols. Informed consent for tumor biobanking and research studies was obtained from all patients.
Availability of Data and Material
The datasets generated or analyzed during the study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
Author Contributions
Conceptualization: KLFJ, ST, BTH. Data curation: KLFJ, ST. Formal analysis: KLFJ, SM. Funding acquisition: ST, BTH, BDPO. Investigation: KLFJ, ST. Methodology: KLFJ, ST. Supervision: ST, BTH. Validation: KLFJ, ST. Writing—original draft: KLFJ. Writing—review & editing: KLFJ, ST, SM, BTH. Approval of final manuscript: all authors.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding Statement
No funding to declare.
| Author (year published) | Rate of Fn detection in CRC tumors (%) | Detection method of Fn | Sample form | Sample age |
|---|---|---|---|---|
| Nosho et al. (2016) [6] | 8.6 | qPCR ΔCT | FFPE | Unclear |
| Author’s work | 10.4 | qPCR ΔCT | FFPE | 0–20 yr |
| Lee et al. (2018) [7] |
12 in FFPE 98 in methacarn 100 in frozen tissues |
qPCR ΔCT | FFPE, methacarn or frozen | 6–13 yr |
| Mima et al. (2015, 2016) [8–10] | 13 | qPCR ΔCT | FFPE | 7–20 yr |
| Borowsky et al. (2021) [11] | 13.7 | qPCR ΔCT | FFPE |
8 to > 26 years NB updated same cohort used by Mima et al., may have used FN data from previous analysis in 2015 for older samples |
| Shariati et al. (2021) [12] | 23 | qPCR ΔCT | Fresh frozen | 0–2 yr |
| Kunzmann et al. (2019) [13] | 31 ‘high’ for FN but unclear % positive | qPCR ΔCT | Fresh frozen | 6–10 yr |
| Yan et al. (2017) [14] | 38 | qPCR ΔCT | Frozen | 4–5 yr |
| Okita et al. (2020) [15] | 38 | qPCR ΔCT (different human genome control gene) | FFPE | Age unclear |
| Yamamoto et al. (2021) [16] | 45 | qPCR ΔCT | FFPE | 3–4 yr |
| Tahara et al. (2014) [17] |
52 for FN 73.8 for pan-fusobacterium |
qPCR ΔCT | Unclear | Unclear |
| Lee et al. (2020) [18] | 54 | qPCR ΔCT | FFPE | Age unclear |
| Chen et al. (2019) [19] | 55 | qPCR ΔCT | FFPE | <2 yr |
| Ito et al. (2015) [20] | 56 | qPCR ΔCT | FFPE | 1–13 yr |
| Serna et al. (2020) [21] | 57 pre-treatment (rectal cancers only) | FISH for Fn RNA | FFPE | 2–13 yr |
| Tunsjo et al. (2019) [22] | 60 | qPCR ΔCT (used B-globin as control gene) | Allprotect tissue reagent, unclear time to DNA extraction or storage before analysis | 2–5 yr |
| Yu et al. (2016) [23] | 61.3 overall | FISH for Fn RNA | FFPE | 1–2 yr |
| Kashani et al. (2020) [24] | 68 | PCR of Fn and electrophoresis without control gene | Fresh samples |
1–3 years but may have extracted DNA immediately |
| Oh et al. (2019) [2] | 68.8 | qPCR ΔCT | FFPE |
5–13 yr Excluded 20% due to poor quality |
| Yamaoka et al. (2018) [25] | 75 | ddPCR | Frozen | 10–18 yr |
| Kim et al. (2020) [26] | 82 | qPCR relative to other bacterial genes | Fresh samples with immediate DNA extraction | 4–5 yr |
| Li et al. (2016) [27] | 87.1 | PCR ΔCT | Frozen | 3–4 yr |
Comparison of Fn detection rates in publications analyzing CRC tumors by detection method, sample storage format and sample age. There is some tendency for older sample cohorts to have lower rates of detection, for frozen or fresh samples to have higher detection rates compared to FFPE, and ddPCR and FISH may be more sensitive methods than the ubiquitous quantitative polymerase chain reaction for Fn relative to control gene (qPCR ΔCT).
Fn, Fusobacterium nucleatum; CRC, colorectal cancer; qPCR ΔCT, ubiquitous quantitative polymerase chain reaction for Fn relative to control gene; FFPE, formalin-fixed paraffin-embedded; ddPCR, digital droplet polymerase chain reaction; FISH, fluorescent in situ hybridization.
- 1. Brennan CA, Garrett WS. Fusobacterium nucleatum: symbiont, opportunist and oncobacterium. Nat Rev Microbiol 2019; 17: 156-66. ArticlePubMedPMCPDF
- 2. Oh HJ, Kim JH, Bae JM, Kim HJ, Cho NY, Kang GH. Prognostic impact of Fusobacterium nucleatum depends on combined tumor location and microsatellite instability status in stage II/III colorectal cancers treated with adjuvant chemotherapy. J Pathol Transl Med 2019; 53: 40-9. ArticlePubMedPDF
- 3. Groelz D, Viertler C, Pabst D, Dettmann N, Zatloukal K. Impact of storage conditions on the quality of nucleic acids in paraffin embedded tissues. PLoS One 2018; 13: e0203608.ArticlePubMedPMC
- 4. Flores Bueso Y, Walker SP, Tangney M. Characterization of FFPE-induced bacterial DNA damage and development of a repair method. Biol Methods Protoc 2020; 5: bpaa015.
- 5. Walker SP, Tangney M, Claesson MJ. Sequence-based characterization of intratumoral bacteria: a guide to best practice. Front Oncol 2020; 10: 179.ArticlePubMedPMC
- 6. Nosho K, Sukawa Y, Adachi Y, et al. Association of Fusobacterium nucleatum with immunity and molecular alterations in colorectal cancer. World J Gastroenterol 2016; 22: 557-66. ArticlePubMedPMC
- 7. Lee DW, Han SW, Kang JK, et al. Association between Fusobacterium nucleatum, pathway mutation, and patient prognosis in colorectal cancer. Ann Surg Oncol 2018; 25: 3389-95. ArticlePubMedPDF
- 8. Mima K, Sukawa Y, Nishihara R, et al. Fusobacterium nucleatum and T cells in colorectal carcinoma. JAMA Oncol 2015; 1: 653-61. ArticlePubMedPMC
- 9. Mima K, Cao Y, Chan AT, et al. Fusobacterium nucleatum in colorectal carcinoma tissue according to tumor location. Clin Transl Gastroenterol 2016; 7: e200.ArticlePubMedPMCPDF
- 10. Mima K, Nishihara R, Qian ZR, et al. Fusobacterium nucleatum in colorectal carcinoma tissue and patient prognosis. Gut 2016; 65: 1973-80. ArticlePubMed
- 11. Borowsky J, Haruki K, Lau MC, et al. Association of Fusobacterium nucleatum with specific T-cell subsets in the colorectal carcinoma microenvironment. Clin Cancer Res 2021; 27: 2816-26. PubMedPMC
- 12. Shariati A, Razavi S, Ghaznavi-Rad E, et al. Association between colorectal cancer and Fusobacterium nucleatum and Bacteroides fragilis bacteria in Iranian patients: a preliminary study. Infect Agent Cancer 2021; 16: 41.ArticlePubMedPMCPDF
- 13. Kunzmann AT, Proenca MA, Jordao HW, et al. Fusobacterium nucleatum tumor DNA levels are associated with survival in colorectal cancer patients. Eur J Clin Microbiol Infect Dis 2019; 38: 1891-9. ArticlePubMedPMCPDF
- 14. Yan X, Liu L, Li H, Qin H, Sun Z. Clinical significance of Fusobacterium nucleatum, epithelial-mesenchymal transition, and cancer stem cell markers in stage III/IV colorectal cancer patients. Onco Targets Ther 2017; 10: 5031-46. PubMedPMC
- 15. Okita Y, Koi M, Takeda K, et al. Fusobacterium nucleatum infection correlates with two types of microsatellite alterations in colorectal cancer and triggers DNA damage. Gut Pathog 2020; 12: 46.ArticlePubMedPMCPDF
- 16. Yamamoto S, Kinugasa H, Hirai M, et al. Heterogeneous distribution of Fusobacterium nucleatum in the progression of colorectal cancer. J Gastroenterol Hepatol 2021; 36: 1869-76. ArticlePubMedPDF
- 17. Tahara T, Yamamoto E, Suzuki H, et al. Fusobacterium in colonic flora and molecular features of colorectal carcinoma. Cancer Res 2014; 74: 1311-8. ArticlePubMedPMCPDF
- 18. Lee MS, Keku TO, McCoy A, et al. Association of Fusobacterium nucleatum (F. nucleatum) with progression-free survival (PFS) and overall survival (OS) with 2nd-line FOLFIRI+/-regorafenib in metastatic colorectal cancer (mCRC). Cancer Res 2020; 80(8 Suppl): A03.
- 19. Chen Y, Lu Y, Ke Y, Li Y. Prognostic impact of the Fusobacterium nucleatum status in colorectal cancers. Medicine (Baltimore) 2019; 98: e17221.ArticlePubMedPMC
- 20. Ito M, Kanno S, Nosho K, et al. Association of Fusobacterium nucleatum with clinical and molecular features in colorectal serrated pathway. Int J Cancer 2015; 137: 1258-68. ArticlePubMed
- 21. Serna G, Ruiz-Pace F, Hernando J, et al. Fusobacterium nucleatum persistence and risk of recurrence after preoperative treatment in locally advanced rectal cancer. Ann Oncol 2020; 31: 1366-75. ArticlePubMed
- 22. Tunsjo HS, Gundersen G, Rangnes F, Noone JC, Endres A, Bemanian V. Detection of Fusobacterium nucleatum in stool and colonic tissues from Norwegian colorectal cancer patients. Eur J Clin Microbiol Infect Dis 2019; 38: 1367-76. ArticlePubMedPDF
- 23. Yu J, Chen Y, Fu X, et al. Invasive Fusobacterium nucleatum may play a role in the carcinogenesis of proximal colon cancer through the serrated neoplasia pathway. Int J Cancer 2016; 139: 1318-26. ArticlePubMed
- 24. Kashani N, Bezmin Abadi AT, Rahimi F, Forootan M. FadA-positive Fusobacterium nucleatum is prevalent in biopsy specimens of Iranian patients with colorectal cancer. New Microbes New Infect 2020; 34: 100651.ArticlePubMedPMC
- 25. Yamaoka Y, Suehiro Y, Hashimoto S, et al. Fusobacterium nucleatum as a prognostic marker of colorectal cancer in a Japanese population. J Gastroenterol 2018; 53: 517-24. ArticlePubMedPDF
- 26. Kim M, Lee ST, Choi S, et al. Fusobacterium nucleatum in biopsied tissues from colorectal cancer patients and alcohol consumption in Korea. Sci Rep 2020; 10: 19915.ArticlePubMedPMCPDF
- 27. Li YY, Ge QX, Cao J, et al. Association of Fusobacterium nucleatum infection with colorectal cancer in Chinese patients. World J Gastroenterol 2016; 22: 3227-33. ArticlePubMedPMC
REFERENCES
Figure & Data
References
Citations

 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure

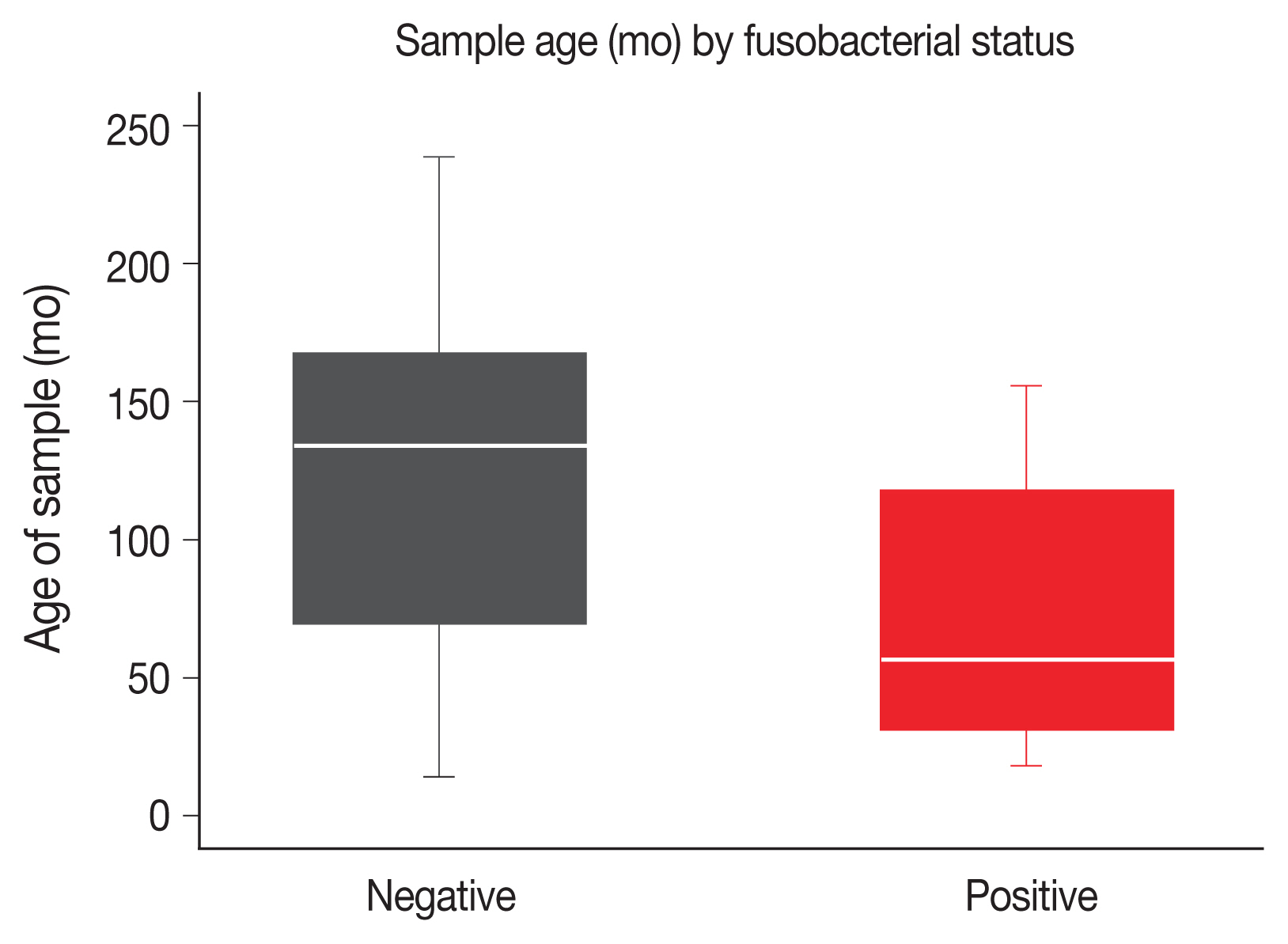
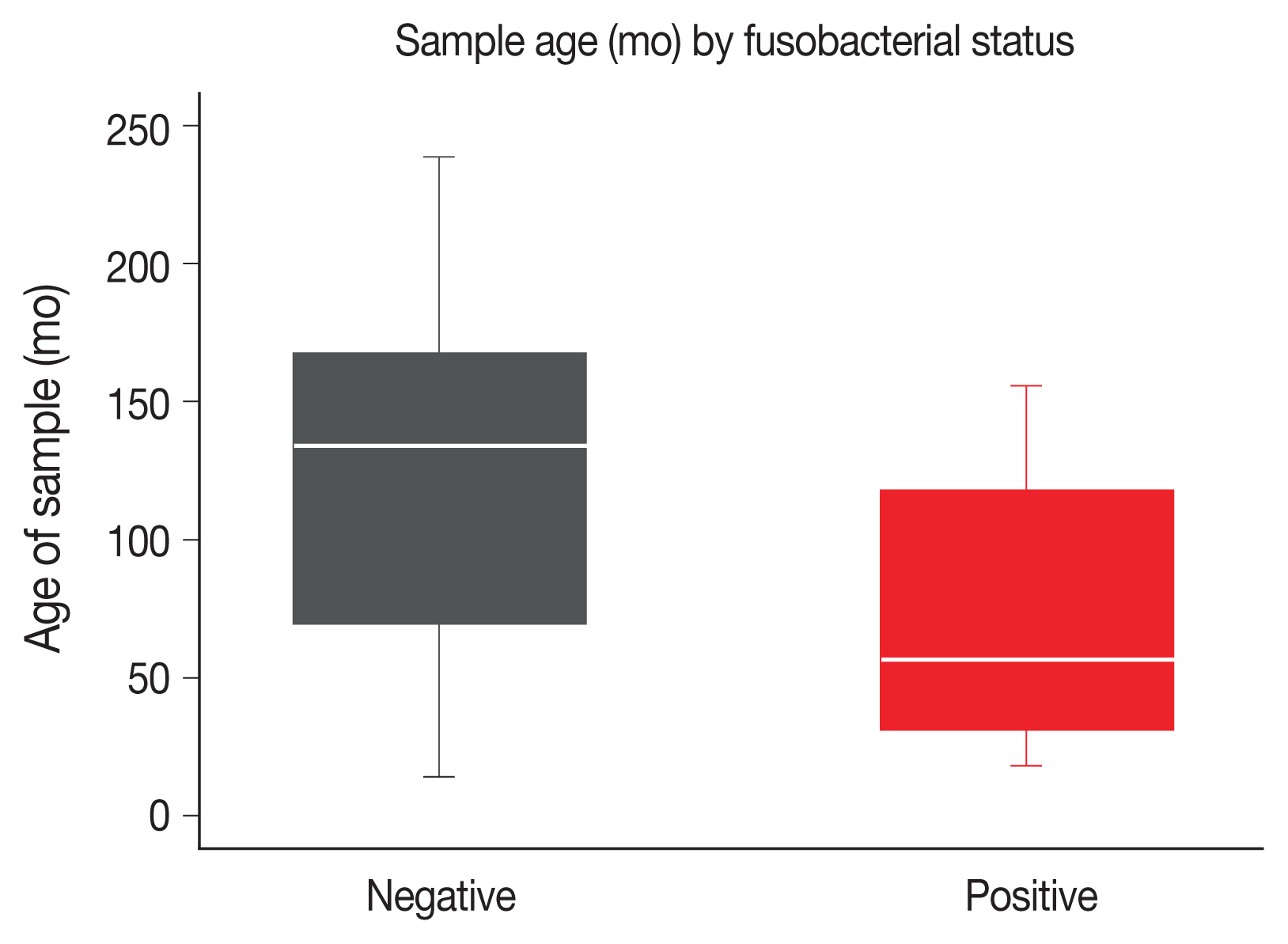
Fig. 1
| Author (year published) | Rate of Fn detection in CRC tumors (%) | Detection method of Fn | Sample form | Sample age |
|---|---|---|---|---|
| Nosho et al. (2016) [ |
8.6 | qPCR ΔCT | FFPE | Unclear |
| Author’s work | 10.4 | qPCR ΔCT | FFPE | 0–20 yr |
| Lee et al. (2018) [ |
12 in FFPE 98 in methacarn 100 in frozen tissues |
qPCR ΔCT | FFPE, methacarn or frozen | 6–13 yr |
| Mima et al. (2015, 2016) [ |
13 | qPCR ΔCT | FFPE | 7–20 yr |
| Borowsky et al. (2021) [ |
13.7 | qPCR ΔCT | FFPE | 8 to > 26 years NB updated same cohort used by Mima et al., may have used FN data from previous analysis in 2015 for older samples |
| Shariati et al. (2021) [ |
23 | qPCR ΔCT | Fresh frozen | 0–2 yr |
| Kunzmann et al. (2019) [ |
31 ‘high’ for FN but unclear % positive | qPCR ΔCT | Fresh frozen | 6–10 yr |
| Yan et al. (2017) [ |
38 | qPCR ΔCT | Frozen | 4–5 yr |
| Okita et al. (2020) [ |
38 | qPCR ΔCT (different human genome control gene) | FFPE | Age unclear |
| Yamamoto et al. (2021) [ |
45 | qPCR ΔCT | FFPE | 3–4 yr |
| Tahara et al. (2014) [ |
52 for FN 73.8 for pan-fusobacterium |
qPCR ΔCT | Unclear | Unclear |
| Lee et al. (2020) [ |
54 | qPCR ΔCT | FFPE | Age unclear |
| Chen et al. (2019) [ |
55 | qPCR ΔCT | FFPE | <2 yr |
| Ito et al. (2015) [ |
56 | qPCR ΔCT | FFPE | 1–13 yr |
| Serna et al. (2020) [ |
57 pre-treatment (rectal cancers only) | FISH for Fn RNA | FFPE | 2–13 yr |
| Tunsjo et al. (2019) [ |
60 | qPCR ΔCT (used B-globin as control gene) | Allprotect tissue reagent, unclear time to DNA extraction or storage before analysis | 2–5 yr |
| Yu et al. (2016) [ |
61.3 overall | FISH for Fn RNA | FFPE | 1–2 yr |
| Kashani et al. (2020) [ |
68 | PCR of Fn and electrophoresis without control gene | Fresh samples | 1–3 years but may have extracted DNA immediately |
| Oh et al. (2019) [ |
68.8 | qPCR ΔCT | FFPE | 5–13 yr Excluded 20% due to poor quality |
| Yamaoka et al. (2018) [ |
75 | ddPCR | Frozen | 10–18 yr |
| Kim et al. (2020) [ |
82 | qPCR relative to other bacterial genes | Fresh samples with immediate DNA extraction | 4–5 yr |
| Li et al. (2016) [ |
87.1 | PCR ΔCT | Frozen | 3–4 yr |
Comparison of Fn detection rates in publications analyzing CRC tumors by detection method, sample storage format and sample age. There is some tendency for older sample cohorts to have lower rates of detection, for frozen or fresh samples to have higher detection rates compared to FFPE, and ddPCR and FISH may be more sensitive methods than the ubiquitous quantitative polymerase chain reaction for Fn relative to control gene (qPCR ΔCT). Fn,

 E-submission
E-submission