Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 57(5); 2023 > Article
-
Original Article
Establishing molecular pathology curriculum for pathology trainees and continued medical education: a collaborative work from the Molecular Pathology Study Group of the Korean Society of Pathologists -
Jiwon Koh1
 , Ha Young Park2
, Ha Young Park2 , Jeong Mo Bae1
, Jeong Mo Bae1 , Jun Kang3
, Jun Kang3 , Uiju Cho4
, Uiju Cho4 , Seung Eun Lee5
, Seung Eun Lee5 , Haeyoun Kang6
, Haeyoun Kang6 , Min Eui Hong7
, Min Eui Hong7 , Jae Kyung Won1
, Jae Kyung Won1 , Youn-La Choi8
, Youn-La Choi8 , Wan-Seop Kim5
, Wan-Seop Kim5 , Ahwon Lee3,9
, Ahwon Lee3,9 , The Molecular Pathology Study Group of the Korean Society of Pathologists
, The Molecular Pathology Study Group of the Korean Society of Pathologists -
Journal of Pathology and Translational Medicine 2023;57(5):265-272.
DOI: https://doi.org/10.4132/jptm.2023.08.26
Published online: September 15, 2023
1Department of Pathology, Seoul National University Hospital, Seoul National University College of Medicine, Seoul, Korea
2Department of Pathology, Busan Paik Hospital, Inje University College of Medicine, Busan, Korea
3Department of Hospital Pathology, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea
4Department of Pathology, St. Vincent’s Hospital, College of Medicine, The Catholic University of Korea, Suwon, Korea
5Department of Pathology, Konkuk University Medical Center, Konkuk University School of Medicine, Seoul, Korea
6Department of Pathology, CHA Bundang Medical Center, CHA University, Seongnam, Korea
7Department of Pathology, Chung-Ang University Hospital, Seoul, Korea
8Department of Pathology and Translational Genomics, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
9Cancer Research Institute, College of Medicine, The Catholic University of Korea, Seoul, Korea
- Corresponding Author: Ahwon Lee, MD, Department of Hospital Pathology, College of Medicine, The Catholic University of Korea, Seoul St. Mary’s Hospital, 222 Banpo-daero, Seocho-gu, Seoul 06591, Korea Tel: +82-2-2258-1621, Fax: +82-2-2258-1627, E-mail: klee@catholic.ac.kr
© 2023The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- The importance of molecular pathology tests has increased during the last decade, and there is a great need for efficient training of molecular pathology for pathology trainees and as continued medical education.
-
Methods
- The Molecular Pathology Study Group of the Korean Society of Pathologists appointed a task force composed of experienced molecular pathologists to develop a refined educational curriculum of molecular pathology. A 3-day online educational session was held based on the newly established structure of learning objectives; the audience were asked to score their understanding of 22 selected learning objectives before and after the session to assess the effect of structured education.
-
Results
- The structured objectives and goals of molecular pathology was established and posted as a web-based interface which can serve as a knowledge bank of molecular pathology. A total of 201 pathologists participated in the educational session. For all 22 learning objectives, the scores of self-reported understanding increased after educational session by 9.9 points on average (range, 6.6 to 17.0). The most effectively improved items were objectives from next-generation sequencing (NGS) section: ‘NGS library preparation and quality control’ (score increased from 51.8 to 68.8), ‘NGS interpretation of variants and reference database’ (score increased from 54.1 to 68.0), and ‘whole genome, whole exome, and targeted gene sequencing’ (score increased from 58.2 to 71.2). Qualitative responses regarding the adequacy of refined educational curriculum were collected, where favorable comments dominated.
-
Conclusions
- Approach toward the education of molecular pathology was refined, which would greatly benefit the future trainees.
- The Molecular Pathology Study Group of the KSP appointed a TF composed of experienced molecular pathologists to develop a refined educational curriculum of MP. The TF initially started with literature review of MP education and training approaches in US and established structured key learning objectives of MP training in Korea. Online resource repository composed of key references for each learning objective was organized by the TF.
- Based on the newly established structure of learning objectives, 3-day online MP educational sessions were held on February 2, 12, and 26, 2022. To assess the usefulness of the learning objective-oriented educational sessions, the TF also created survey asking the session audience to score their understanding of selected learning objectives before and after the session by Likert scale.
MATERIALS AND METHODS
- Lessons from US experiences
- In US, the need for adequate MP education started around late 1990s [9], owing to the advances in the application of molecular biology technology. In the year 1999, the Training and Education Committee of Association for Molecular Pathology (AMP) provides an outline of important elements of resident MP education [7]. The general goals provided by the AMP include basic knowledge in human genetics, basics of molecular biology, and specific skills relevant to microbiology, molecular oncology, genetics, histocompatibility, and identity determination [7]. In addition, the AMP also provided the list of sentinel papers highlighting the important concepts of infectious disease, molecular oncology, inherited disorders, histocompatibility, and identity determination [7].
- Along with the advances in NGS technologies and increase in the importance of genomic medicine in the clinical practice, the need for the refinement of educational topics was increased over time. Therefore, the Stanford group pioneered to establish a genomic pathology curriculum–so-called The Stanford Open Curriculum–to help pathology residents build a foundation for the understanding of genomic medicine and the implications for clinical practice [10]. The 10 lectures encompassed the overview of the fundamental principles of molecular biology, clinical genomics, and personalized medicine, and the curriculum also provided 7 additional topics as the elective course of advanced genomic medicine [11].
- However, the real-world data suggested that there existed a significant discrepancy between the literal description of curriculum and the subjective perception among the trainees. In a survey of 42 pathology residency programs, only 31% reported that genomic medicine training was included in their program [12]. The serious educational gap prompted the pathologists and medical education specialists in US to organize Training Residents in Genomics (TRIG) Working Group. The TRIG Working Group conducted a nationwide survey for the in-serviced pathology residents in 2013 [13]. 42% and 7% of residents reported that they had no training of genomic medicine and MP during their residency, respectively [13]. When asked whether they were able to discuss MP or genomic medicine test results with a provider, only 13% and 28% of the responders scored that their ability as “very good/excellent” [13].
- On observing the obvious educational gap, the Training and Educational Committee of AMP appointed the Molecular Curriculum Task Force; the TF developed an organized curriculum in MP for residents [14]. The major subjects of curriculum included basic MP goals/laboratory management, basic concepts in molecular biology and genetics, technology, inherited disorders, oncology, infectious diseases, pharmacogenetics, histocompatibility and identity, genomics, and information management [14]. Besides, TRIG Working Group provided online genomic pathology modules to improve genomics knowledge and the ability to utilize online genomics tools (https://www.pathologylearning.org/trig). The first version was introduced in the year 2016, and the most recent updates were added in the year 2020 [15,16]. To date, the TRIG has expanded their work toward the education of the undergraduate medical students [17].
- Learning objectives and online repository
- For adaptation of US experiences into real-world education in South Korea, 13 major topics of MP education was selected by the TF, and detailed description of objectives and goals were generated under each topic, comprising a total of 75 structured items as shown in Table 1. The structured objectives included basic items such as ‘1-2. Definition of genome, exome, proteome, transcriptome, and metabolome’, and further encompassed timely topics such as ‘2-10. Able to explain the definitions of companion diagnostics and complementary diagnostics.’
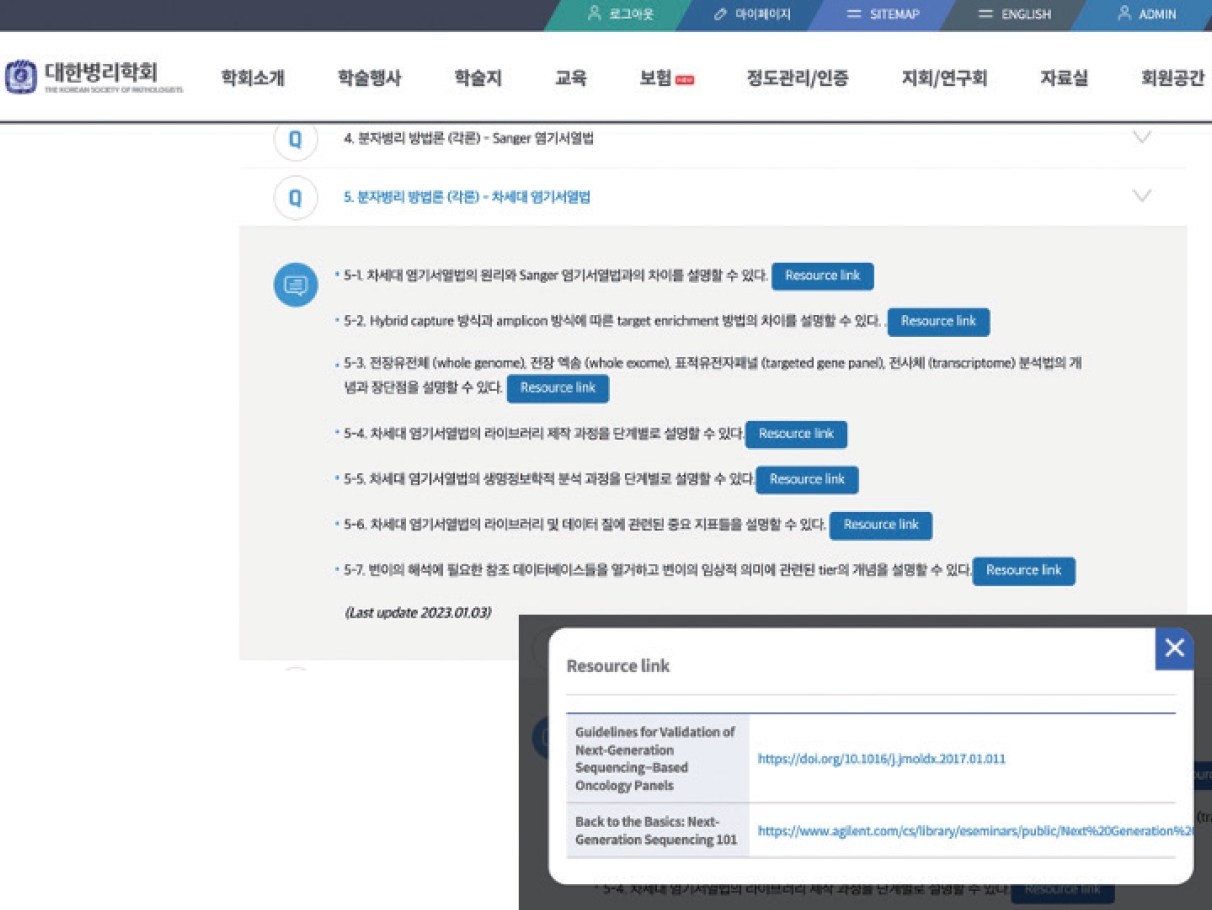
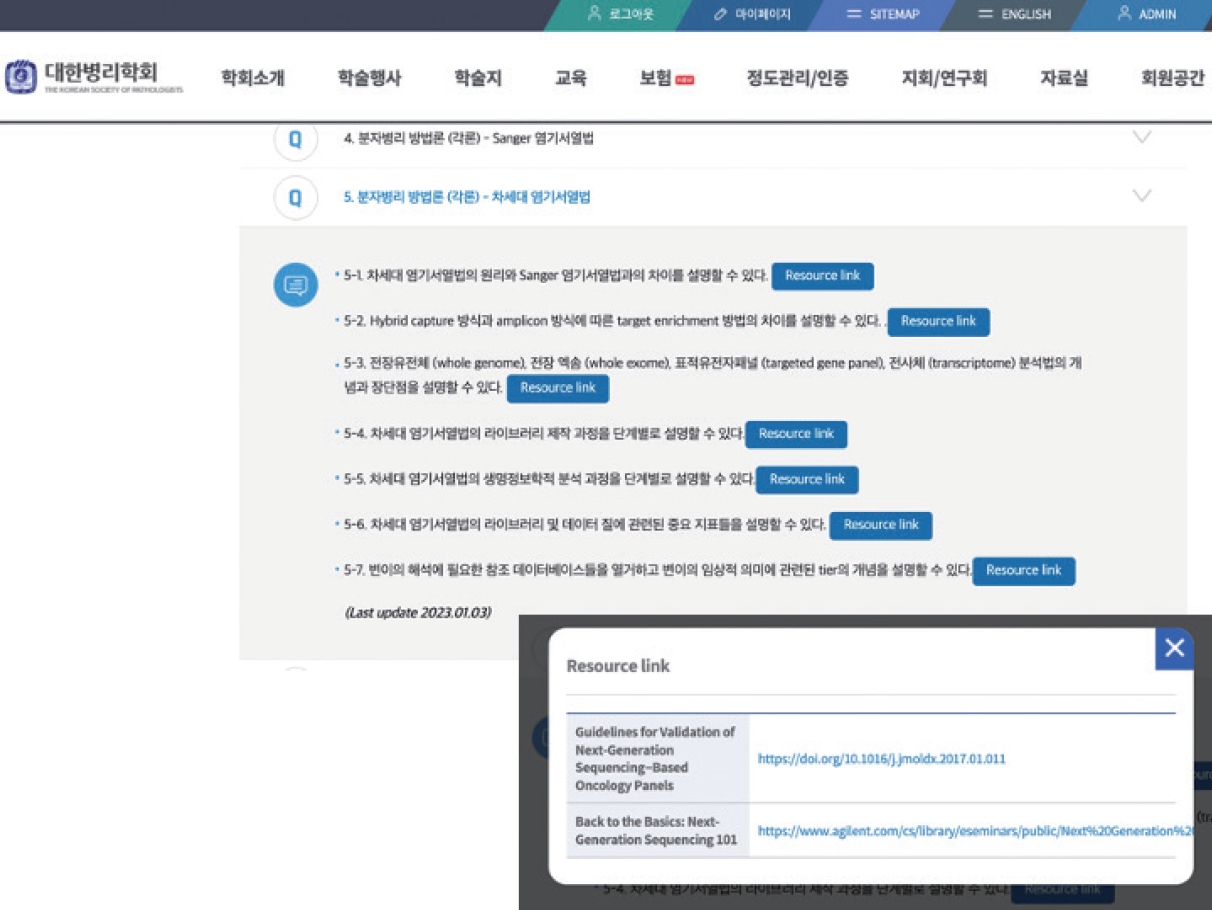
- To provide high-quality resources for trainees, TF members were assigned with each major topic and reviewed the relevant online resources and assessed the quality of contents. After the thorough review of contents, the most adequate learning sources were selected and gathered to form the online repository of MP learning objectives. The online repository became available for all the members of KSP via the official webpage (https://www.pathology.or.kr/html/?pmode=boardlist&MMC_pid=306&cate=study) (Fig. 1).
- 3-Day online structured educational session and survey
- Based on the structural objectives, the Molecular Pathology Study Group of the KSP planned and held 3-day online structured educational session in February 2022 (Table 2). A total of 201 pathologists participated in the educational session.
- To immediately assess the effectiveness of refined educational session, the TF selected 22 key learning objectives out of total 75 elements and asked the session audience to grade their understanding of each topic before and after the lecture, respectively. The survey was performed at day 1 and day 2 with response rates as follows: day 1 pre-lecture, 93/201 (46.3%); day 1 post-lecture 83/201 (41.3%); day 2 pre-lecture 50/201 (24.9%); day 2 post-lecture 34/201 (16.9%).
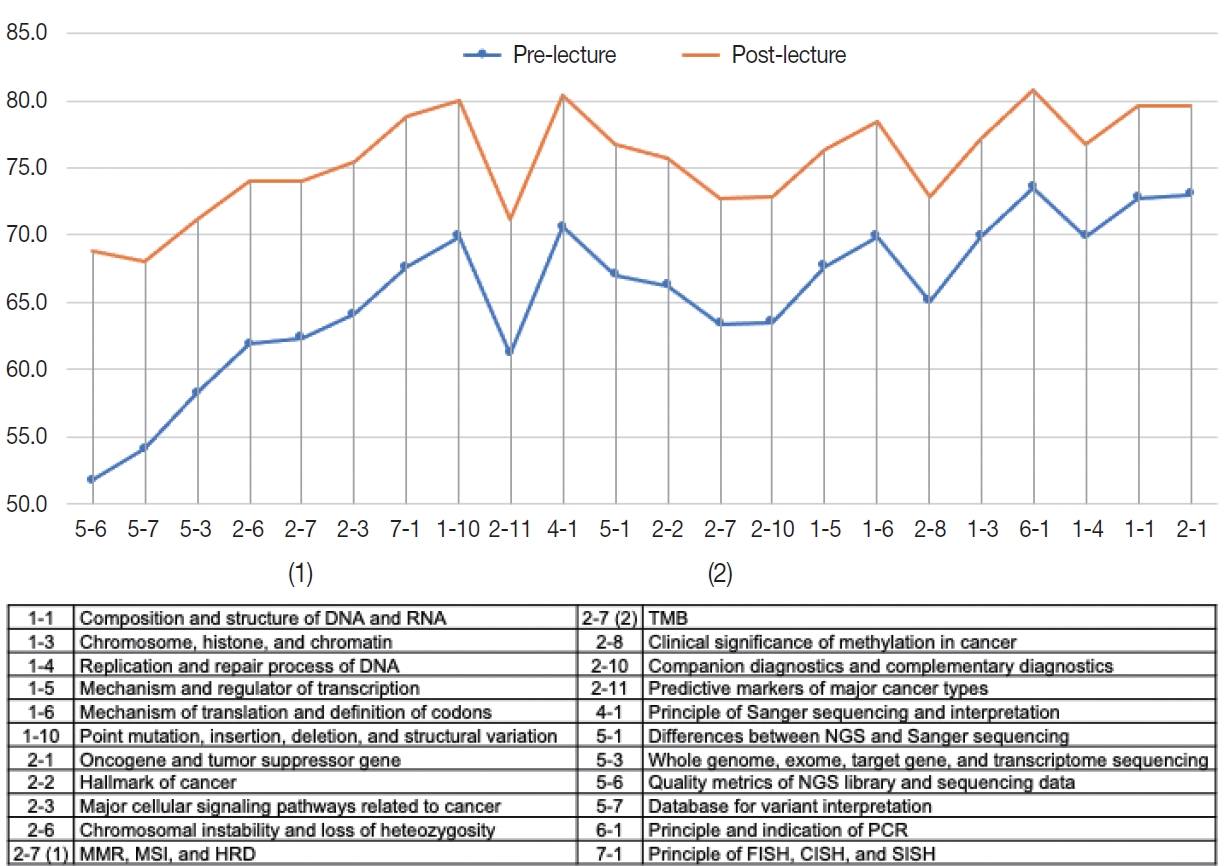
- Among 22 learning objectives (Fig. 2), the audience reported higher scores regarding their prior understanding on the basic concepts of MP including ‘1-1. Composition and structure of DNA and RNA’ (mean score 72.8), ‘2-1. Oncogene and tumor suppressor gene’ (mean score 73.0), and ‘6-1. Principle and indication of PCR’ (mean score 73.5). In contrast, self-reported understanding prior to the session was lowest in the topics related to NGS: ‘5-6. Quality metrics of NGS library and sequencing data’ (mean score 51.8); ‘5-7. Database for variant interpretation’ (mean score 54.1); ‘5-3. Whole genome, whole exome, target gene, and transcriptome sequencing’ (mean score 58.2).
- The scores of self-reported understandings increased after educational session by 9.9 points on average (range, 6.6 to 17.0) (Fig. 2). Of interest, the most effectively improved items were those with lowest pre-lecture scores as follows: ‘5-6. Quality metrics of NGS library and sequencing data’ (score increased from 51.8 to 68.8); ‘5-7. Database for variant interpretation’ (score increased from 54.1 to 68.0); ‘5-3. Whole genome, whole exome, target gene, and transcriptome sequencing’ (score increased from 58.2 to 71.2).
- Additionally qualitative responses regarding the adequacy of refined educational curriculum were collected, where favorable comments dominated. The audience provided responses regarding the further suggestions regarding the topic for future educational sessions, and some of the examples are as follows: real-world clinical NGS cases, NGS raw data analysis using R software, wet-lab techniques of NGS, and cutting-edge novel technologies including proteomics and proteogenomics.
RESULTS
- In line with the recent increase in need for the high-quality education of MP for pathology trainees, the Molecular Pathology Study Group of the KSP initiated this project for the development of MP curriculum.
- The project first started with the review of US experiences on the MP education during the last decade [7,9-12,14-18]. Next, the project focused on the establishment of the structured objectives and goals of MP, which was later online resource repository within KSP official website to enable continuous update and better accessibility. Also, 3-day structured course was held according to the newly established learning objectives, which received favorable responses from the audience.
- The TF appointed by the Molecular Pathology Study Group of the KSP specifically aimed to list up learning objectives and goals, which can readily be introduced into practical educational sessions, like the successful experience by TRIG Working Group [12,16,17]. These efforts resulted in the production of online resource repository and 3-day online MP course. The favorable responses from the KSP members suggest that the purpose of this project was relatively well achieved. Still, we noticed some limitations and further challenges.
- First, unlike the survey performed by TRIG [13], the overall response rate of the survey was low, which implies that the data may not represent the opinions of all the pathology residents in Korea. In addition, our survey did not collect the information with respect to the training levels of audience; since the subjective need for certain items among the learning objectives may differ according to the training level or current job description, further collection of feedbacks from variable groups among KSP members should follow. Moreover, besides the subjective assessment of participants’ understandings, well-prepared questionnaires and self-assessment programs for objective measurement of participants’ academic accomplishment should be provided in the near future.
- The most important challenge that need to be addressed would be the sustainability and timely adaptability of the education curriculum. The area of MP and genomic medicine is one of the most rapidly evolving sectors among the field of pathology [3,4,19]. The efforts from small group of eager pathologists would not be able to maintain the long-lasting high-quality educational platform. Continuous feedback from all the KSP members along with the leadership from Molecular Pathology Study Group of KSP could surely make the quality of MP education better, which would benefit the whole medical society in South Korea. In addition, an in-depth pool of high-quality assessment contents should be further developed for proper evaluation of the educational effect of this novel approach.
- In conclusion, for better education of MP within KSP, structured learning objectives and goals of MP was refined and listed, which resulted in the production of online reference repository and 3-day MP lecture course. Approach toward the education of MP was refined, and this big step can further greatly benefit the future trainees.
DISCUSSION
Ethics Statement
Not applicable.
Availability of Data and Material
The datasets used and/or analysed during the current study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
Author contributions
Conceptualization: AL. Data curation: JK (Jiwon Koh), HYP, UC, JKW, AL. Formal analysis: JK (Jiwon Koh). Funding acquisition: AL. Investigation: JK (Jiwon Koh), HYP, JKW, AL. Methodology: JK (Jiwon Koh), JKW, AL. Resources: JK (Jiwon Koh), HYP, JMB, JK (Jun Kang), UC, SEL, HK, MEH, JKW, YLC, WSK, AL. Writing—original draft: JK (Jiwon Koh). Writing—review & editing: all authors. Approval of final manuscript: all authors.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding Statement
This research was supported by The Korean Society of Pathologists Grant No. KSPG2021-02.

MMR, mismatch repair; MSI, microsatellite instability; HRD, Homologous recombination deficiency; TMB, tumor mutation burden; NGS, next-generation sequencing; PCR, polymerase chain reaction; FISH, fluorescent in situ hybridization; CISH, chromogenic in situ hybridization; SISH, silver in situ hybridization.
- 1. Harris TJ, McCormick F. The molecular pathology of cancer. Nat Rev Clin Oncol 2010; 7: 251-65. ArticlePubMedPDF
- 2. Tobin NP, Foukakis T, De Petris L, Bergh J. The importance of molecular markers for diagnosis and selection of targeted treatments in patients with cancer. J Intern Med 2015; 278: 545-70. ArticlePubMedPDF
- 3. Yoon S, Kim M, Hong YS, et al. Recommendations for the use of next-generation sequencing and the molecular tumor board for patients with advanced cancer: a report from KSMO and KCSG Precision Medicine Networking Group. Cancer Res Treat 2022; 54: 1-9. ArticlePubMedPDF
- 4. Lee SH, Lee B, Shim JH, et al. Landscape of actionable genetic alterations profiled from 1,071 tumor samples in Korean cancer patients. Cancer Res Treat 2019; 51: 211-22. ArticlePubMedPDF
- 5. Kim J, Park WY, Kim NK, et al. Good laboratory standards for clinical next-generation sequencing cancer panel tests. J Pathol Transl Med 2017; 51: 191-204. ArticlePubMedPMCPDF
- 6. Big Data Hub of Korean Health Insurance Review & Assessment Service [Internet]. Wonju: Health Insurance Review & Assessment Service, 2021 [cited 2022 Nov 3]. Available from: https://opendata.hira.or.kr/.
- 7. The Association for Molecular Pathology Training Education Committee. Goals and objectives for molecular pathology education in residency programs. The Association for Molecular Pathology Training and Education Committee. J Mol Diagn 1999; 1: 5-15. PubMedPMC
- 8. Ministry of Health and Welfare. Residency competency training project [Internet]. Sejong: Introduction of Korean Law Information Center, 2022 [cited 2023 Jun 18]. Available from: https://www.law.go.kr/.
- 9. Arber DA, Battifora H, Brynes RK, Chang KL, Weiss LM, Wilczynski SP. Certification and training in molecular pathology. Hum Pathol 1996; 27: 430-1. ArticlePubMed
- 10. Schrijver I, Natkunam Y, Galli S, Boyd SD. Integration of genomic medicine into pathology residency training: the stanford open curriculum. J Mol Diagn 2013; 15: 141-8. PubMed
- 11. Haspel RL, Genzen JR, Wagner J, Lockwood CM, Fong K; Training Residents in Genomics (TRIG) Working Group. Integration of genomic medicine in pathology resident training. Am J Clin Pathol 2020; 154: 784-91. PubMedPMCPDF
- 12. Haspel RL, Atkinson JB, Barr FG, et al. TRIG on TRACK: educating pathology residents in genomic medicine. Per Med 2012; 9: 287-93. ArticlePubMed
- 13. Haspel RL, Rinder HM, Frank KM, et al. The current state of resident training in genomic pathology: a comprehensive analysis using the resident in-service examination. Am J Clin Pathol 2014; 142: 445-51. PubMed
- 14. Aisner DL, Berry A, Dawson DB, Hayden RT, Joseph L, Hill CE. A suggested molecular pathology curriculum for residents: a report of the association for molecular pathology. J Mol Diagn 2016; 18: 153-62. PubMed
- 15. Haspel RL, Ali AM, Huang GC, et al. Teaching genomic pathology: translating team-based learning to a virtual environment using computer-based simulation. Arch Pathol Lab Med 2019; 143: 513-7. ArticlePubMedPDF
- 16. Haspel RL, Ali AM, Huang GC. Using a team-based learning approach at national meetings to teach residents genomic pathology. J Grad Med Educ 2016; 8: 80-4. ArticlePubMedPMCPDF
- 17. Wilcox RL, Adem PV, Afshinnekoo E, et al. The Undergraduate Training in Genomics (UTRIG) Initiative: early & active training for physicians in the genomic medicine era. Per Med 2018; 15: 199-208. ArticlePubMedPMC
- 18. Clay MR, Fisher KE. Bioinformatics education in pathology training: current scope and future direction. Cancer Inform 2017; 16: 1176935117703389.ArticlePubMedPMCPDF
- 19. Zehir A, Benayed R, Shah RH, et al. Mutational landscape of metastatic cancer revealed from prospective clinical sequencing of 10,000 patients. Nat Med 2017; 23: 703-13. PubMedPMC
REFERENCES
Figure & Data
References
Citations

- Presence of RB1 or Absence of LRP1B Mutation Predicts Poor Overall Survival in Patients with Gastric Neuroendocrine Carcinoma and Mixed Adenoneuroendocrine Carcinoma
In Hye Song, Bokyung Ahn, Young Soo Park, Deok Hoon Kim, Seung-Mo Hong
Cancer Research and Treatment.2025; 57(2): 492. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure


Fig. 1.
Fig. 2.
| 1 | Gene and genome | |
| 1-1 | Able to explain the composition and structure of DNA and RNA | |
| 1-2 | Able to explain the definition of genome, exome, proteome, transcriptome, and metabolome | |
| 1-3 | Able to explain the chromosome, histone, and chromatin | |
| 1-4 | Able to explain the replication and repair process of DNA | |
| 1-5 | Able to explain the mechanism and regulator of transcription | |
| 1-6 | Able to explain the mechanism of translation and definition of codons | |
| 1-7 | Able to explain the definition of promoter, enhancer, cis- and trans-regulation | |
| 1-8 | Able to explain the types and roles of epigenetic gene regulation | |
| 1-9 | Able to explain the definition of genetic polymorphism | |
| 1-10 | Able to explain the definition of point mutation, insertion, deletion, and structural variation including translocation | |
| 1-11 | Able to explain the definition of missense, synonymous, nonsense, null, and frameshift mutations | |
| 1-12 | Able to explain the association between mutations and genomic instability and RNA splicing | |
| 2 | Molecular oncology (introduction) | |
| 2-1 | Able to explain the definition of oncogene and tumor suppressor gene | |
| 2-2 | Able to explain the hallmark of cancer | |
| 2-3 | Able to explain the major cellular signaling pathways related to oncogenesis and cancer progression | |
| 2-4 | Able to explain the mechanism through which proto-oncogenes are activated and transformed into oncogenes | |
| 2-5 | Able to explain the mechanisms of familial and hereditary tumor development | |
| 2-6 | Able to explain the definition of chromosomal instability and loss of heterozygosity | |
| 2-7 | Able to explain the definition and clinical significance of mismatch repair deficiency, microsatellite instability, homologous recombination deficiency, tumor mutational burden | |
| 2-8 | Able to explain the definition and clinical significance of methylation in cancer | |
| 2-9 | Able to list the indications of molecular pathology testings in cancer (i.e., screening, diagnostic, prognostic, predictive, treatment monitoring) | |
| 2-10 | Able to explain the definitions of companion diagnostics and complementary diagnostics | |
| 2-11 | Able to list the predictive markers of major cancer types | |
| 3 | Techniques of molecular pathology | |
| 3-1 | Able to explain the DNA/RNA extraction process according to the sample types | |
| 3-2 | Able to explain the ways of assessing quality and quantity of DNA/RNA and list the pros and cons of each method | |
| 3-3 | Able to list the ways of assessing genetic mutations and list the pros and cons of each method | |
| 3-4 | Able to explain why the minimum tumor requirement differs between assays | |
| 3-5 | Able to explain the factors influencing the quality of tissue samples and nucleic acid | |
| 3-6 | Able to properly name the genetic mutations according to the Human Genome Variation Society (HGVS) nomenclature | |
| 3-7 | Able to explain the assessment process and criteria of the clinical performance evaluation | |
| 3-8 | Able to explain the assessment process and criteria of the analytical performance evaluation | |
| 4 | Sanger sequencing | |
| 4-1 | Able to explain the principle of Sanger sequencing and interpretation | |
| 4-2 | Able to interpretate the electropherogram of each type of mutation | |
| 4-3 | Able to identify the causes of false positive or false negative results | |
| 4-4 | Able to interpretate the mutation analysis results and generate proper clinical reports | |
| 4-5 | Able to list the specific disease-related genetic mutations | |
| 5 | Next-generation sequencing (NGS) | |
| 5-1 | Able to explain the differences between NGS and Sanger sequencing | |
| 5-2 | Able to explain the differences between hybrid capture and amplicon-based target enrichment | |
| 5-3 | Able to explain the definitions of whole genome, whole exome, targeted gene panel, and transcriptome sequencing and pros/cons of each method | |
| 5-4 | Able to explain each step of library preparation for NGS | |
| 5-5 | Able to explain each step of bioinformatic analysis for NGS data | |
| 5-6 | Able to explain the major quality metrics related to the qualities of library and sequencing data | |
| 5-7 | Able to list major reference database for use in variant interpretation and definition of tier-based classification system | |
| 6 | Polymerase chain reaction (PCR) | |
| 6-1 | Able to explain the principle and indication of PCR | |
| 6-2 | Able to list the technical considerations of performing PCR | |
| 6-3 | Able to interpretate the PCR analysis results and generate proper clinical reports | |
| 6-4 | Able to explain the principle and indication of real-time PCR (RT-PCR) | |
| 6-5 | Able to list the technical considerations of performing RT-PCR | |
| 6-6 | Able to interpretate the RT-PCR analysis results and generate proper clinical reports | |
| 7 | In situ hybridization (ISH) | |
| 7-1 | Able to explain the principle and process of fluorescent in situ hybridization (FISH), chromogenic in situ hybridization (CISH), and silver in situ hybridization (SISH) | |
| 7-2 | Able to list the indications of FISH, CISH, and SISH | |
| 7-3 | Able to explain the types of FISH probes | |
| 7-4 | Able to explain the clinical significance of FISH results | |
| 7-5 | Able to interpretate the results of FISH, CISH, SISH and generate proper clinical reports | |
| 8 | Microsatellite instability (MSI) test | |
| 8-1 | Able to explain the definition of microsatellite | |
| 8-2 | Able to explain the principles and techniques of MSI tests | |
| 8-3 | Able to explain the clinicopathological significance of MSI | |
| 9 | Methylation analysis | |
| 9-1 | Able to explain the principles and techniques of methylation analysis | |
| 9-2 | Able to explain the definition of CpG island methylator phenotype | |
| 9-3 | Able to list the technical considerations of performing methylation analysis | |
| 9-4 | Able to interpretate the results of methylation analysis and generate proper clinical reports | |
| 10 | Gene rearrangement test | |
| 10-1 | Able to explain the definition of gene rearrangement and utilize the results in the diagnosis and treatment planning | |
| 10-2 | Able to list the methods of gene rearrangement tests and explain the pros/cons of each method | |
| 10-3 | Able to interpretate the results of gene rearrangement analysis and generate proper clinical reports | |
| 11 | Human papillomavirus genotyping by DNA microarray | |
| 11-1 | Able to explain the principle of DNA microarray and utilize the results in the diagnosis and treatment planning | |
| 11-2 | Able to explain the process of DNA microarray test | |
| 11-3 | Able to list the methods of DNA microarray tests and explain the pros/cons of each method | |
| 11-4 | Able to interpretate the results of DNA microarray analysis and generate proper clinical reports | |
| 12 | Chromosome analysis | |
| 12-1 | Able to explain the principle of chromosome analysis | |
| 12-2 | Able to explain the result and clinical significance of chromosome analysis | |
| 13 | Molecular pathology laboratory management | |
| 13-1 | Able to list precautions for preventing cross-contamination during each step of molecular pathology testings | |
| 13-2 | Able to list the reference database for genetic testings | |
| 13-3 | Able to list the credential criteria and quality metrics for laboratory certificate | |
| 13-4 | Able to guide the clinicians for proper molecular pathology testings | |
| 13-5 | Able to explain the definitions of research use only, investigational use only, in vitro diagnostics, laboratory developed test, and analyte specific reagents |
| Day 1 | ||
| 1 | Introduction to molecular pathology | |
| 2 | DNA/RNA | |
| 3 | Gene expression | |
| 4 | Epigenetics | |
| 5 | Molecular oncology (I) | |
| 6 | Molecular oncology (II) | |
| 7 | Hallmark of cancer | |
| Day 2 | ||
| 1 | Molecular technology (1) – Principle, application, and practice | |
| 2 | Molecular technology (2) – Principle, application, and practice | |
| 3 | Molecular technology (3) – Principle, application, and practice | |
| 4 | NGS system: Technology & wet-lab | |
| 5 | NGS-bioinformatics and knowledge base | |
| 6 | NGS interpretation and reporting: in the view of pathologists | |
| 7 | Pathologists role in immuno-oncology | |
| 8 | Companion diagnosis/liquid biopsy | |
| Day 3 | ||
| 1 | Molecular diagnostics & personalized medicine in breast | |
| 2 | Molecular diagnostics & personalized medicine in female genital tract | |
| 3 | Molecular diagnostics & personalized medicine in gastrointestinal tract | |
| 4 | Molecular diagnostics & personalized medicine in primary brain tumor | |
| 5 | Molecular diagnostics & personalized medicine in endocrine organ | |
| 6 | Molecular diagnostics & personalized medicine in urogenital tract | |
| 7 | Molecular diagnostics & personalized medicine in lung and mediastinum | |
| 8 | Molecular diagnostics & personalized medicine in skin | |
| 9 | Molecular diagnostics & personalized medicine in hematopathology | |
| 10 | Molecular diagnostics & personalized medicine in bone and soft tissue | |
MMR, mismatch repair; MSI, microsatellite instability; HRD, Homologous recombination deficiency; TMB, tumor mutation burden; NGS, next-generation sequencing; PCR, polymerase chain reaction; FISH, fluorescent in situ hybridization; CISH, chromogenic in situ hybridization; SISH, silver in situ hybridization.

 E-submission
E-submission