Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 52(3); 2018 > Article
-
Original Article
Utility of BRAF VE1 Immunohistochemistry as a Screening Tool for Colorectal Cancer Harboring BRAF V600E Mutation - Jeong-Hwa Kwon, Byung-Kwan Jeong, Yong Sik Yoon1, Chang Sik Yu1, Jihun Kim
-
Journal of Pathology and Translational Medicine 2018;52(3):157-163.
DOI: https://doi.org/10.4132/jptm.2018.03.28
Published online: March 29, 2018
Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
1Department of Surgery, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
- Corresponding Author Jihun Kim, MD, PhD Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, 88 Olympic-ro 43-gil, Songpa-gu, Seoul 05505, Korea Tel: +82-2-3010-4556 Fax: +82-2-472-7898 E-mail: jihunkim@amc.seoul.kr
© 2018 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- BRAF mutation has been recognized as an important biomarker of colorectal cancer (CRC) for targeted therapy and prognosis prediction. However, sequencing for every CRC case is not cost-effective. An antibody specific for BRAF V600E mutant protein has been introduced, and we thus examined the utility of BRAF VE1 immunohistochemistry for evaluating BRAF mutations in CRC.
-
Methods
- Fifty-one BRAF-mutated CRCs and 100 age and sexmatched BRAF wild-type CRCs between 2005 and 2015 were selected from the archives of Asan Medical Center. Tissue microarrays were constructed and stained with BRAF VE1 antibody.
-
Results
- Forty-nine of the 51 BRAF-mutant CRCs (96.1%) showed more than moderate cytoplasmic staining, except for two weakly stained cases. Six of 100 BRAF wild-type cases also stained positive with BRAF VE1 antibody; four stained weakly and two stained moderately. Normal colonic crypts showed nonspecific weak staining, and a few CRC cases exhibited moderate nuclear reactivity (3 BRAF-mutant and 10 BRAF wild-type cases). BRAF-mutated CRC patients had higher pathologic stages and worse survival than BRAF wild-type patients.
-
Conclusions
- BRAF VE1 immunohistochemistry showed high sensitivity and specificity, but occasional nonspecific staining in tumor cell nuclei and normal colonic crypts may limit their routine clinical use. Thus, BRAF VE1 immunohistochemistry may be a useful screening tool for BRAF V600E mutation in CRCs, provided that additional sequencing studies can be done to confirm the mutation in BRAF VE1 antibody-positive cases.
- Patients and samples
- The study group consisted of 51 surgically resected primary or metastatic CRC cases harboring BRAF V600E mutation (colonoscopic biopsy [n = 17], primary tumor resection [n = 31], and metastasectomy [n = 3]) and 100 age and sex-matched BRAF wild-type CRCs (colonoscopic biopsy [n = 14], primary tumor resection [n = 81], and metastasectomy [n = 5]). They were selected from the surgical pathology files between 2005 and 2015 at the Department of Pathology, Asan Medical Center, University of Ulsan Collage of Medicine, Seoul, Korea. The BRAF V600E mutation status was confirmed by Sanger sequencing (n = 75), quantitative allele-specific PCR (n = 16), and mass spectrometry-based genotyping (n = 60). All cases were KRAS wild-type. Histopathological features of the 151 CRCs were reviewed by two pathologists (J.H.K. and J.K.) and clinical information including age, gender, tumor location, histology, lymphovascular invasion, perineural invasion, serosal involvement, nodal status, and follow-up results was obtained from the medical records. This study was approved by the Institutional Review Board (IRB) (2015-1393) of Asan Medical Center, and patient innformed consent was waived by the IRB.
- Tissue microarray construction and IHC
- Tissue microarrays (TMAs) were constructed from 34 surgically resected BRAF-mutated samples and 86 BRAF wild-type samples.
- The TMA was constructed using a hollow needle to remove a tissue core (0.2 cm in diameter) from tumors on paraffin-embedded tissue blocks. These cores were then inserted into recipient blocks. Sections of the TMA blocks were cut using a microtome, mounted on a microscope slide, and then stained. TMA and biopsy samples were subjected to IHC analysis using anti-BRAF antibody (mouse monoclonal, clone VE1, catalog number: 790-4855, Ventana Medical Systems, Tucson, AZ, USA) and a BenchMark XT automatic immunostaining device (Ventana Medical Systems) with an OptiView DAB IHC Detection Kit (Ventana Medical Systems) according to the manufacturer’s instructions with slight modifications: we diluted primary antibody with recommended dilution buffer to 1:4 and increased primary antibody incubation time from 16 to 32 minutes, in order to prevent nonspecific background signals.
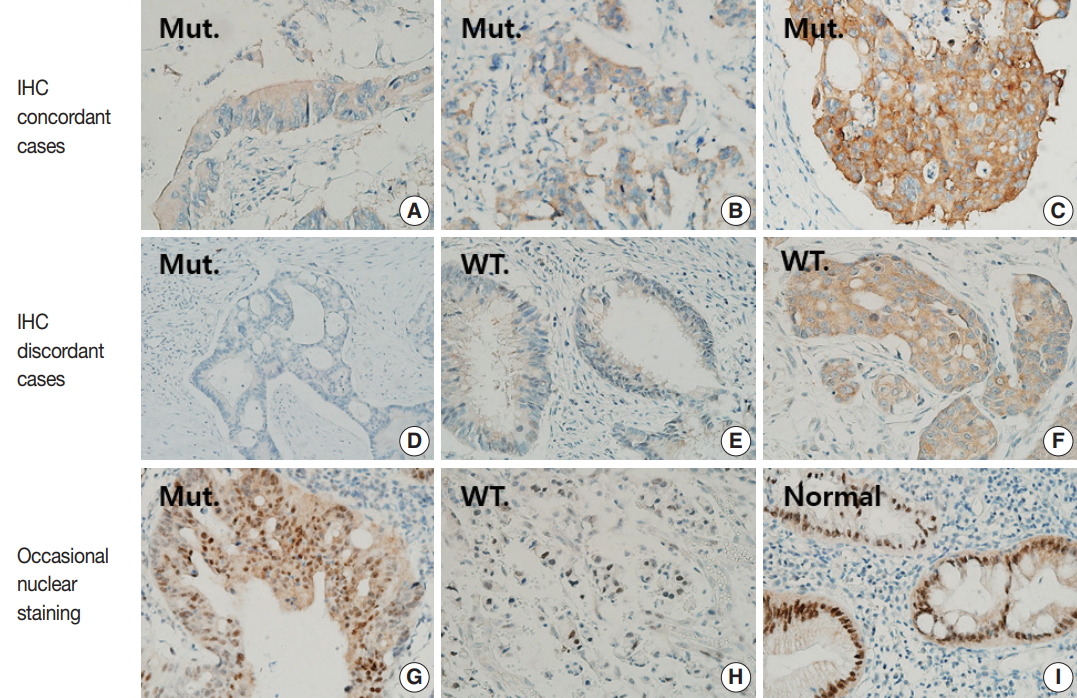
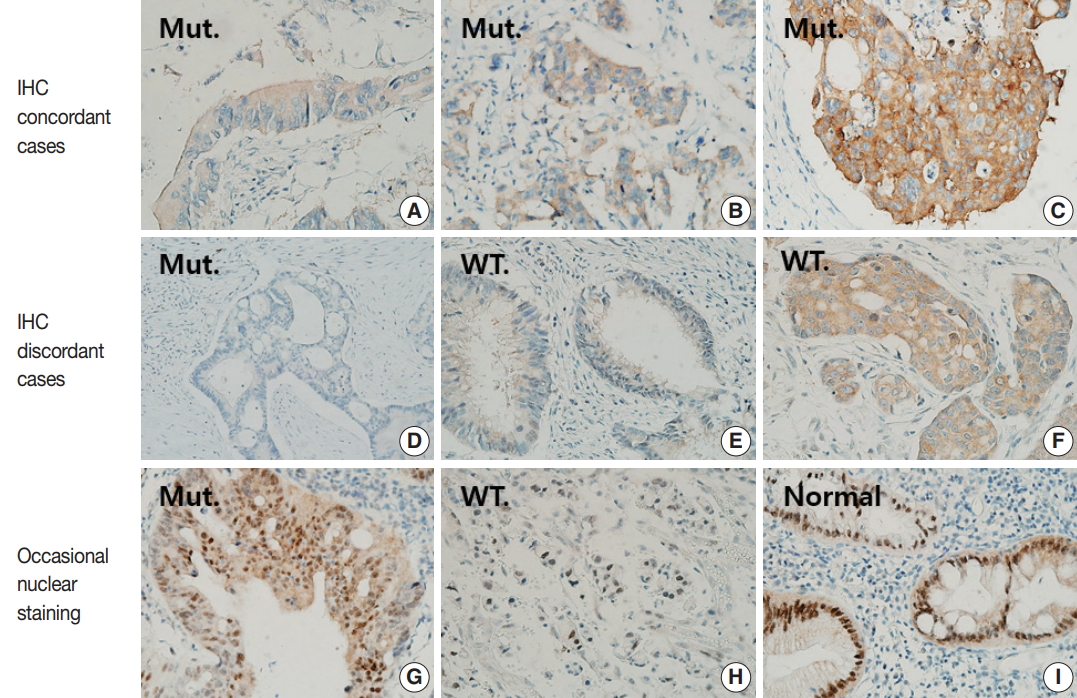
- IHC staining results were graded using a 4-tier grading system according to the staining intensity as follows: 0 (no staining), 1+ (faint), 2+ (moderate), and 3+ (strong) (Fig. 1A–C). Only cytoplasmic staining was considered positive. As a positive control, we selected a case of papillary thyroid carcinoma harboring BRAF V600E mutation and strong BRAF VE1 staining. As a negative control, we used normal tonsil tissue stained in the same manner with and without primary antibody. When the results of BRAF VE1 IHC differed from those of BRAF sequencing, we repeated BRAF VE1 IHC using whole tumor sections that were cut from the same paraffin block from which DNA had been extracted for BRAF sequencing.
- Determination of BRAF mutation status
- BRAF V600E mutation status was confirmed by Sanger sequencing (n = 75), quantitative allele-specific PCR (n = 16), or mass spectrometry-based genotyping (n = 60) as described previously [9-11]. All tumor tissue sections were macrodissected to increase tumor purity. When tumor purity in the macrodissected area was low (< 40%) and Sanger sequencing did not detect BRAF mutations, the BRAF mutation status was confirmed by a more sensitive method such as quantitative allele-specific PCR or mass spectrometric genotyping.
- Statistical analysis
- To compare clinicopathologic variables, statistical analyses were performed using SPSS ver. 20.0 statistical software (IBM Corp., Armonk, NY, USA) and differences between the two groups were compared by either chi-square test or Fisher’s exact test. The Kaplan-Meier method with log-rank test and multivariate Cox proportional hazards regression models were applied for survival analyses. Two-sided p-values of < .05 were considered statistically significant.
MATERIALS AND METHODS
- Diagnostic performance of BRAF VE1 IHC
- Forty-nine of 51 CRCs (96.1%) harboring BRAF V600E mutation showed cytoplasmic staining for BRAF VE1 antibody with variable intensities (Table 1, Fig. 1A–D): 3+ in 23 cases (45.1%), 2+ in 24 cases (47.1%), and 1+ in two cases (3.9%). In two BRAF-mutant cases (3.9%), no signal was detected by BRAF VE1 IHC. In 100 BRAF wild-type controls, 94 (94%) cases showed no staining, while six cases (6%) showed cytoplasmic staining with moderate (2 cases, 2%) or weak (4 cases, 4%) intensities (Table 1, Fig. 1E, F). Thus, the sensitivity, specificity, positive predictive value, and negative predictive value of BRAF VE1 IHC were 96.1%, 94%, 89.1%, and 97.9%, respectively. The cutoff for a positive staining was set to 1+ or bigger score because the area under curve was maximal at this cutoff in receiver operating characteristic (ROC) curve analysis (Supplementary Fig. S1). BRAF V600E mutant tumors with negative BRAF VE1 staining or BRAF wild-type tumors with positive BRAF VE1 staining did not exhibit any distinct clinicopathologic features (Supplementary Tables S1, S2).
- Analysis of cases with discrepant results between BRAF mutation status and BRAF VE1 IHC results
- For cases with discrepant results between BRAF sequencing and BRAF VE1 IHC, we repeated BRAF VE1 IHC on the same paraffin block from which DNA had been extracted for sequencing analyses. However, BRAF VE1 IHC on the whole tumor section showed the same results as those on TMA. As for BRAF-mutant cases that showed negative BRAF VE1 IHC results, IHC was repeated using matched biopsy tissues to exclude the possibility of false negative results due to poor fixation. However, the matched biopsy tissues showed the same results. Conversely, for BRAF wild-type cases that showed positive IHC results, we first investigated whether the discrepancies were due to false negative sequencing results associated with low tumor purity. All BRAF wild-type cases with positive immunostaining were examined for tumor purity on hematoxylin and eosin–stained slides; in most cases, tumor purity was more than 30%, and BRAF wild-type status of those cases were confirmed by allele-specific PCR study. One BRAF wild-type CRC with BRAF IHC staining intensity of 2+ had tumor purity of about 5%, but repeated allele-specific PCR study failed to reveal BRAF V600E mutation.
- Atypical patterns of BRAF VE1 IHC and nonspecific staining in normal colonic mucosa
- Three BRAF V600E mutated cases (5.9%) showed moderate nuclear staining together with moderate cytoplasmic staining, and one BRAF V600E wild type case showed only moderate nuclear staining (Fig. 1G, H, Supplementary Table S3). In addition, normal colonic mucosa was also stained, especially along the crypt surface (Fig. 1I).
- Clinicopathologic characteristics of BRAF mutant CRC
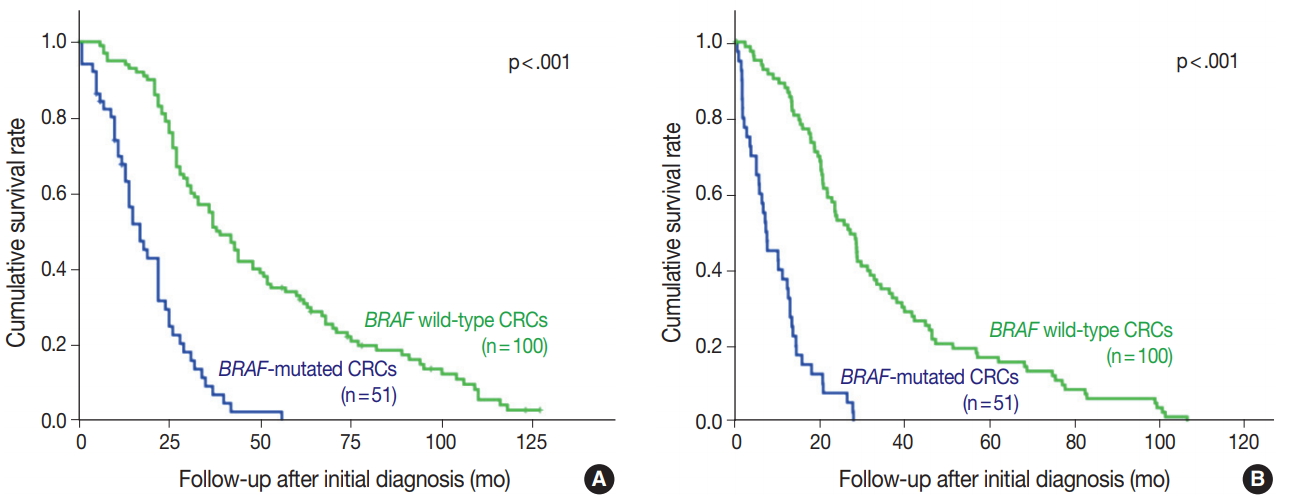
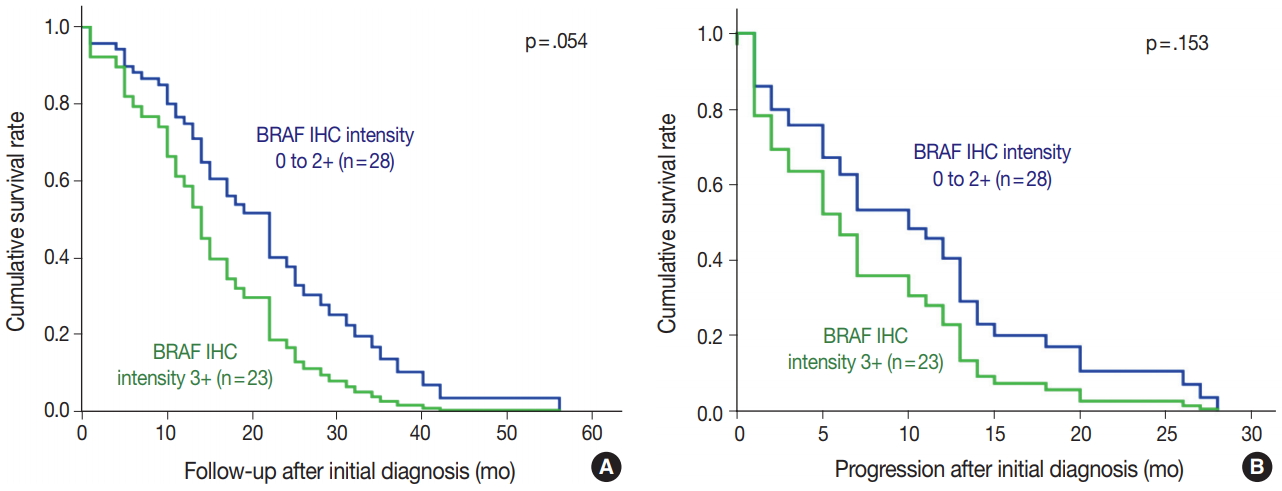
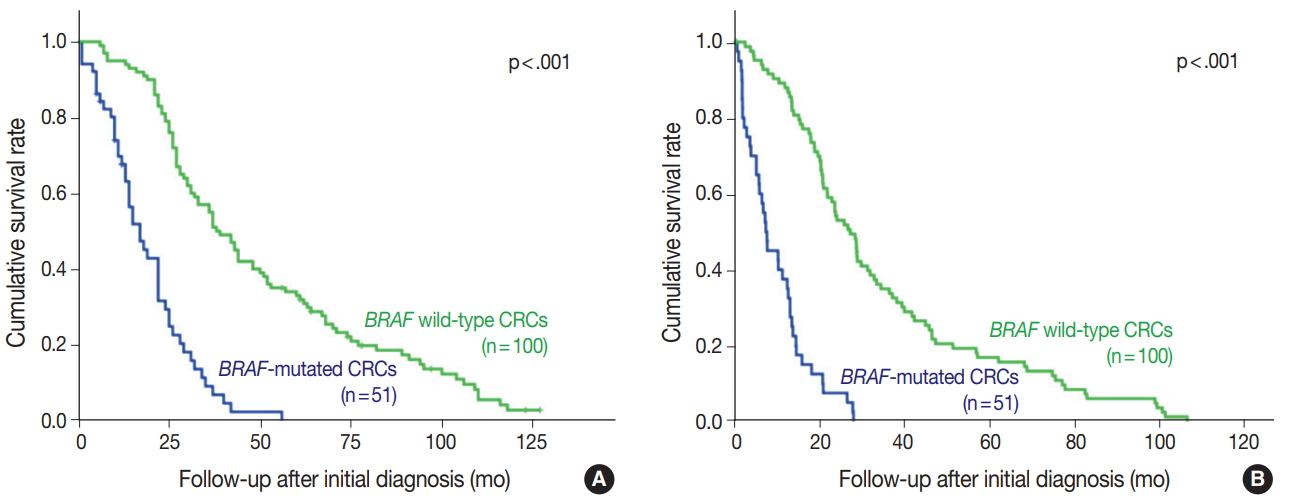
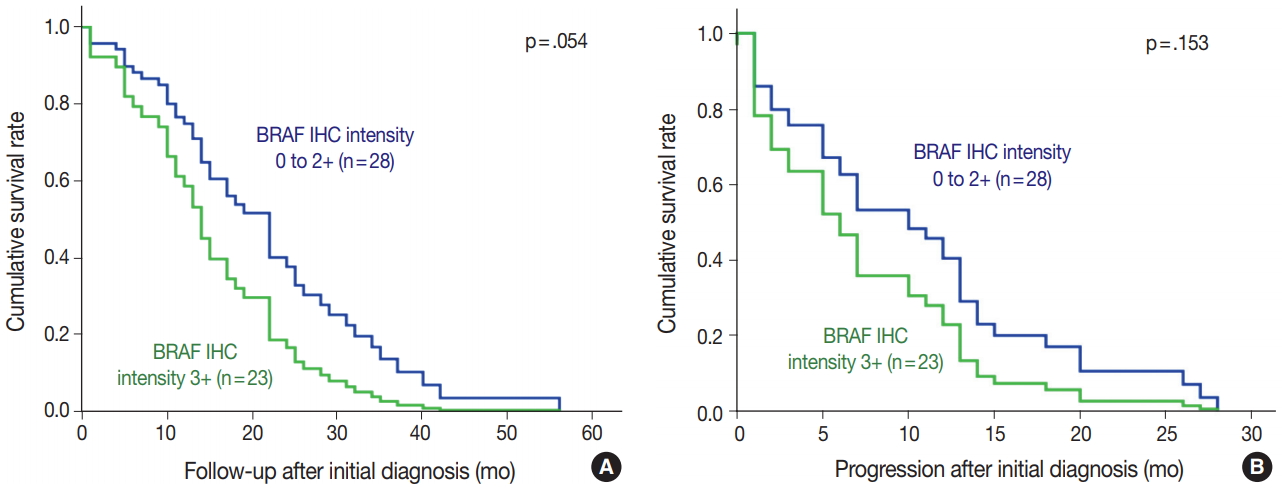
- BRAF V600E mutated CRC cases showed significantly worse overall and progression-free survival (Fig. 2). Patients with BRAF V600E mutant CRC more frequently showed right-sided location, lymphovascular invasion, larger tumor size, and higher TNM stage at diagnosis than did patients with BRAF wildtype CRC. Particularly, BRAF V600E mutant CRCs showed more frequent serosal penetration and peritoneal seeding (Table 2). Because the intensities of BRAF VE1 immunostaining varied within the BRAF V600E mutant CRC group, we speculated that BRAF mutant CRCs with higher mutant BRAF protein expression might show worse prognosis if mutant BRAF protein actually plays a role in the aggressive biologic behavior. Indeed, BRAF mutant CRC cases with higher mutant BRAF protein expression tended to show shorter overall and progression-free survival than those with lower mutant BRAF protein expression, although the differences were not statistically significant (Table 3, Fig. 3).
RESULTS
- In the present study, we showed that the diagnostic performance of BRAF VE1 antibody was relatively good (sensitivity, specificity, and positive predictive values of 96.1%, 94%, and 89.1%, respectively). However, several BRAF V600E mutant CRCs showed no or weak BRAF VE1 staining (n = 4) or BRAF wild-type CRCs showed unequivocal cytoplasmic BRAF VE1 staining (n = 6). In addition, four CRC cases showed nonspecific nuclear BRAF VE1 staining as did normal colonic mucosa. Thus, the usefulness of BRAF VE1 IHC may be limited; it may be difficult to use BRAF VE1 IHC as a routine clinical test, although it may be useful as a screening tool when used in conjunction with subsequent confirmatory sequencing.
- BRAF-mutant CRCs, which were all microsatellite stable, were in advanced stages at diagnosis (p < .001) and showed worse overall and recurrence-free survival than BRAF wild-type CRCs. These results are consistent with those of most previous studies [12-14]. Moreover, BRAF-mutant CRCs were associated with the right colon, larger primary tumor size, and presence of lymphovascular invasion, all of which are consistent with the results of most previous studies [2].
- Recently, BRAF VE1 antibody has been used as a biomarker of CRC in IHC studies of BRAF. The clinical usefulness of BRAF VE1 antibody in colon cancer is controversial, but most studies showed that BRAF VE1 IHC has an excellent sensitivity [5,6]. Interpretation of BRAF VE1 IHC may be difficult due to technical problems such as poor fixation or staining failure [15,16]. Thus, we compared BRAF VE1 IHC and fixation quality between surgically resected tissues and matched colonoscopic biopsy tissues of two BRAF mutant CRC cases that showed negative staining results. There was no difference between the biopsied tissue and surgically resected tissue.
- For BRAF wild-type CRCs showing positive immunostaining results, the discrepancies might have resulted from false-negative sequencing results if the tumor purity is very low. For example, tumors with signet rings or tumors with high mucin content have low tumor purity [17]. Therefore, we examined the tumor purity of all BRAF wild-type CRCs that stained positive in IHC. In most cases, false-negative sequencing results were excluded by repeating BRAF mutation analyses using more sensitive methods, but in one BRAF wild-type CRC with a tumor purity of approximately 5% and BRAF VE1 2+, we could not conduct more sensitive mutation analysis because tissue material was unavailable. Therefore, in this case, the possibility of a falsenegative sequencing result could not be excluded.
- Interestingly, CRC cases with more intense BRAF VE1 immunostaining had a tendency to the shorter overall and progression-free survival. Although this result is difficult to interpret, the expression of BRAF V600E mutant protein may play a biological role in tumor aggressiveness rather than being a simple surrogate marker for prognosis. However, our study has a few limitations. Since we performed BRAF VE1 IHC in CRCs with known BRAF mutational status in a retrospective manner, the strength of evidence may be limited compared to that of a prospective design. In addition, a relatively small number of BRAF V600E mutant CRCs may limit the statistical power. Finally, the evaluation of prognostic value of BRAF mutations might be limited because the study population had a selection bias; it had not been selected in a consecutive manner.
- Based on our results, the diagnostic performance of BRAF VE1 IHC showed relatively good but sometimes ambiguous staining, which may limit its routine clinical use; thus, BRAF VE1 IHC cannot replace BRAF sequencing studies. Despite these limitations, BRAF VE1 IHC may be carefully used as a screening tool for BRAF V600E mutation detection in a research basis, as BRAF VE1 IHC is more cost-effective and less time-consuming than BRAF sequencing studies.
DISCUSSION
Electronic Supplementary Material
| BRAF sequencing |
BRAF VE1 immunostaining |
Total | |||
|---|---|---|---|---|---|
| 1+ | 2+ | 3+ | Negative | ||
| V600E mutation | 2 (3.9) | 24 (47.1)a | 23 (45.1) | 2 (3.9) | 51 |
| Wild-type | 4 (4) | 2 (2) | 0 | 94b (94) | 100 |
| Total | 6 | 26 | 23 | 96 | 151 |
- 1. Barras D. BRAF mutation in colorectal cancer: an update. Biomark Cancer 2015; 7(Suppl 1): 9-12. ArticlePubMedPMCPDF
- 2. Boulagnon C, Dudez O, Beaudoux O, et al. BRAF V600E gene mutation in colonic adenocarcinomas: immunohistochemical detection using tissue microarray and clinicopathologic characteristics: an 86 case series. Appl Immunohistochem Mol Morphol 2016; 24: 88-96. PubMed
- 3. Ritterhouse LL, Barletta JA. BRAF V600E mutation-specific antibody: a review. Semin Diagn Pathol 2015; 32: 400-8. ArticlePubMed
- 4. Safaee Ardekani G, Jafarnejad SM, Tan L, Saeedi A, Li G. The prognostic value of BRAF mutation in colorectal cancer and melanoma: a systematic review and meta-analysis. PLoS One 2012; 7: e47054. ArticlePubMedPMC
- 5. Bledsoe JR, Kamionek M, Mino-Kenudson M. BRAF V600E immunohistochemistry is reliable in primary and metastatic colorectal carcinoma regardless of treatment status and shows high intratumoral homogeneity. Am J Surg Pathol 2014; 38: 1418-28. ArticlePubMedPMC
- 6. Mesteri I, Bayer G, Meyer J, et al. Improved molecular classification of serrated lesions of the colon by immunohistochemical detection of BRAF V600E. Mod Pathol 2014; 27: 135-44. ArticlePubMedPDF
- 7. Loes IM, Immervoll H, Angelsen JH, et al. Performance comparison of three BRAF V600E detection methods in malignant melanoma and colorectal cancer specimens. Tumour Biol 2015; 36: 1003-13. ArticlePubMedPDF
- 8. Schafroth C, Galvan JA, Centeno I, et al. VE1 immunohistochemistry predicts BRAF V600E mutation status and clinical outcome in colorectal cancer. Oncotarget 2015; 6: 41453-63. ArticlePubMedPMC
- 9. Gonzalez de Castro D, Angulo B, Gomez B, et al. A comparison of three methods for detecting KRAS mutations in formalin-fixed colorectal cancer specimens. Br J Cancer 2012; 107: 345-51. ArticlePubMedPMCPDF
- 10. Lang AH, Drexel H, Geller-Rhomberg S, et al. Optimized allele-specific real-time PCR assays for the detection of common mutations in KRAS and BRAF. J Mol Diagn 2011; 13: 23-8. ArticlePubMedPMC
- 11. Shin SJ, Chun SM, Kim TI, et al. Feasibility of multiplexed gene mutation detection in plasma samples of colorectal cancer patients by mass spectrometric genotyping. PLoS One 2017; 12: e0176340. ArticlePubMedPMC
- 12. Zlobec I, Bihl MP, Schwarb H, Terracciano L, Lugli A. Clinicopathological and protein characterization of BRAF- and K-RAS-mutated colorectal cancer and implications for prognosis. Int J Cancer 2010; 127: 367-80. ArticlePubMed
- 13. Lochhead P, Kuchiba A, Imamura Y, et al. Microsatellite instability and BRAF mutation testing in colorectal cancer prognostication. J Natl Cancer Inst 2013; 105: 1151-6. ArticlePubMedPMCPDF
- 14. Ogino S, Nosho K, Kirkner GJ, et al. CpG island methylator phenotype, microsatellite instability, BRAF mutation and clinical outcome in colon cancer. Gut 2009; 58: 90-6. ArticlePubMed
- 15. Adackapara CA, Sholl LM, Barletta JA, Hornick JL. Immunohistochemistry using the BRAF V600E mutation-specific monoclonal antibody VE1 is not a useful surrogate for genotyping in colorectal adenocarcinoma. Histopathology 2013; 63: 187-93. ArticlePubMed
- 16. Qiu T, Lu H, Guo L, et al. Detection of BRAF mutation in Chinese tumor patients using a highly sensitive antibody immunohistochemistry assay. Sci Rep 2015; 5: 9211.ArticlePubMedPMCPDF
- 17. Ogino S, Brahmandam M, Cantor M, et al. Distinct molecular features of colorectal carcinoma with signet ring cell component and colorectal carcinoma with mucinous component. Mod Pathol 2006; 19: 59-68. ArticlePubMedPDF
REFERENCES
Figure & Data
References
Citations

- Diagnostic role of BRAF VE1 immunohistochemistry to distinguish benign ciliated epithelium from malignant epithelium in small pulmonary biopsies
Deepali Jain, Udit Naik, Tim D Oury, Chen Zhang
Virchows Archiv.2026;[Epub] CrossRef - Next-Generation Sequencing in Oncology—A Guiding Compass for Targeted Therapy and Emerging Applications
Laurenția Nicoleta Galeș, Mihai-Andrei Păun, Ioana Butnariu, Laurentiu Simion, Loredana Sabina Cornelia Manolescu, Oana Gabriela Trifănescu, Rodica Maricela Anghel
International Journal of Molecular Sciences.2025; 26(7): 3123. CrossRef - A multi-omic analysis reveals a predictive value of tertiary lymphoid structures in improving the prognosis of colorectal cancer patients with BRAF mutation
Chao Qin, Shumin Cheng, Jingyun Ma, Lujing Li, Yun Leng, Lei Zheng, Huiying Chen, Hui Mo, Shi Li, Yuhong Liang, Yi Zhang, Wenxia Li, Jing Liang, Yuxuan Liu, Junxuan Mai, Linlin Hou, Di Wang, Ke Zhu, Bihui Huang
Frontiers in Immunology.2025;[Epub] CrossRef - The evolving landscape of tissue‐agnostic therapies in precision oncology
Vivek Subbiah, Mohamed A. Gouda, Bettina Ryll, Howard A. Burris, Razelle Kurzrock
CA: A Cancer Journal for Clinicians.2024; 74(5): 433. CrossRef - Deciphering the Role of BRAFV600E Immunohistochemistry in Breast Lesions: A Comprehensive Review
Simran Khan, Arvind Bhake, Shakti Sagar
Cureus.2024;[Epub] CrossRef - Immunohistochemistry as a Surrogate Marker of Underlying Molecular Derangements in Sporadic Colorectal Carcinoma in Children – A Series of Three Cases
Priyanka Maity, Aniket Halder, Ranajoy Ghosh, Uttara Chatterjee, Shibsankar Barman, Ruchirendra Sarkar
Fetal and Pediatric Pathology.2022; 41(1): 98. CrossRef - Risk assessment and genetic counseling for Lynch syndrome – Practice resource of the National Society of Genetic Counselors and the Collaborative Group of the Americas on Inherited Gastrointestinal Cancer
Spring Holter, Michael J. Hall, Heather Hampel, Kory Jasperson, Sonia S. Kupfer, Joy Larsen Haidle, Maureen E. Mork, Selvi Palaniapppan, Leigha Senter, Elena M. Stoffel, Scott M. Weissman, Matthew B. Yurgelun
Journal of Genetic Counseling.2022; 31(3): 568. CrossRef - Current concepts in ameloblastoma-targeted therapies in B-raf proto-oncogene serine/threonine kinase V600E mutation: Systematic review
Rogelio González-González, Sandra López-Verdín, Jesús Lavalle-Carrasco, Nelly Molina-Frechero, Mario Isiordia-Espinoza, Ramón G Carreón-Burciaga, Ronell Bologna-Molina
World Journal of Clinical Oncology.2020; 11(1): 31. CrossRef - Genetic and histopathological analysis of a case of primary intraosseous carcinoma, NOS with features of both ameloblastic carcinoma and squamous cell carcinoma
Akane Yukimori, Maiko Tsuchiya, Akane Wada, Yasuyuki Michi, Kou Kayamori, Kei Sakamoto, Tohru Ikeda
World Journal of Surgical Oncology.2020;[Epub] CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure



Fig. 1.
Fig. 2.
Fig. 3.
| BRAF sequencing | BRAF VE1 immunostaining |
Total | |||
|---|---|---|---|---|---|
| 1+ | 2+ | 3+ | Negative | ||
| V600E mutation | 2 (3.9) | 24 (47.1) |
23 (45.1) | 2 (3.9) | 51 |
| Wild-type | 4 (4) | 2 (2) | 0 | 94 |
100 |
| Total | 6 | 26 | 23 | 96 | 151 |
| Clinicopathologic characteristic | BRAF mutant (n = 51) | BRAF wild-type (n = 100) | p-value |
|---|---|---|---|
| Age (yr) | 57 (36–77) | 56 (36–76) | .419 |
| Sex | .189 | ||
| Male | 27 (52.9) | 64 (64.0) | |
| Female | 24 (47.1) | 36 (36.0) | |
| Location | < .001 | ||
| Left side colon | 20 (39.2) | 91 (91.0) | |
| Right side colon | 31 (60.8) | 9 (9.0) | |
| Tumor size (greatest dimension size, cm) | 5.8 (2–18) | 4.5 (0.9–11.2) | .002 |
| T stage | < .001 | ||
| 1–3 | 22 (43.1) | 77 (77.0) | |
| 4a | 12 (23.5) | 5 (5.0) | |
| 4b | 4 (7.8) | 0 | |
| TX | 13 (25.5) | 18 (18.0) | |
| N stage | .001 | ||
| N0 | 2 (3.9) | 19 (19.0) | |
| N1 | 9 (17.6) | 34 (34.0) | |
| N2 | 26 (49.0) | 29 (29.0) | |
| NX | 13 (25.5) | 18 (18.0) | |
| Distant metastasis | < .001 | ||
| No | 0 | 1 (1.0) | |
| Unifocal | 2 (3.9) | 26 (26.0) | |
| Multifocal | 49 (96.1) | 73 (73.0) | |
| Lymphovascular invasion | 31 (60.8) | 35 (35.0) | < .001 |
| Perineural invasion | 23 (45.1) | 33 (33.0) | .065 |
| Resection margin involve | 7 (18.4) | 3 (3.5) | .005 |
| Immunostaining results of BRAF VE1 | < .001 | ||
| Negative | 2 (3.9) | 94 (94.0) | |
| 1+ | 2 (3.9) | 4 (4.0) | |
| 2+ | 24 (47.1) | 2 (2.0) | |
| 3+ | 23 (45.1) | 0 | |
| Peritoneal seeding | 31 (60.8) | 13 (13.0) | < .001 |
| Univariate HR (95% CI) | p-value | Multivariate HR (95% CI) | p-value | |
|---|---|---|---|---|
| Strong BRAF intensity (3+) | 1.84 (0.99–3.42) | .054 | 3.36 (1.29–8.75) | .013 |
| Sex male | 0.98 (0.70–1.39) | .919 | 2.67 (0.99–7.18) | .052 |
| Location | ||||
| Left side colon | Reference | Reference | ||
| Right side colon | 1.01 (0.55–1.84) | .977 | 1.65 (0.67–4.09) | .279 |
| Involved resection margin | 1.43 (0.69–2.94) | .337 | 4.27 (1.20–15.19) | .025 |
| Perineural invasion | 1.58 (1.06–2.37) | .025 | 0.32 (0.10–0.97) | .044 |
| Lymphovascular invasion | 2.14 (1.45–3.18) | < .001 | 10.05 (2.13–47.43) | .004 |
| Lymph node metastasis | .380 | .107 | ||
| N0 | Reference | Reference | ||
| N1 | 4.27 (0.51–36.08) | .182 | 0.11 (0.01–1.98) | .133 |
| N2 | 4.22 (0.55–32.35) | .166 | 0.29 (0.02–4.01) | .289 |
| T category | .001 | .023 | ||
| 1–3 | Reference | Reference | ||
| 4a | 3.13 (1.71–5.71) | < .001 | 4.89 (1.46–16.40) | .010 |
| 4b | 2.36 (0.73–7.59) | .150 | 0.77 (0.19–3.19) | .720 |
| Peritoneal seeding | 1.53 (0.83–2.87) | .188 | 1.31 (0.56–3.03) | .533 |
Values are presented as number (%). Three One
Values are presented as median (range) or number (%). Crosstab analysis for categorical and ordinal variables used chi-square test and for numerical variables used Student t test.
HR, hazard ratio; CI, confidence interval. Univariate and multivariate Cox-regression analyses were used to calculate hazard ratio of clinicopathologic factors on overall survival. Multivariate Cox-regression analysis used the Enter method.

 E-submission
E-submission