Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 55(1); 2021 > Article
-
Review
Standardized pathology report for breast cancer -
Soo Youn Cho1
 , So Yeon Park2
, So Yeon Park2 , Young Kyung Bae3
, Young Kyung Bae3 , Jee Yeon Kim4
, Jee Yeon Kim4 , Eun Kyung Kim5
, Eun Kyung Kim5 , Woo Gyeong Kim6
, Woo Gyeong Kim6 , Youngmee Kwon7
, Youngmee Kwon7 , Ahwon Lee8
, Ahwon Lee8 , Hee Jin Lee9
, Hee Jin Lee9 , Ji Shin Lee10
, Ji Shin Lee10 , Jee Young Park11
, Jee Young Park11 , Gyungyub Gong9
, Gyungyub Gong9 , Hye Kyoung Yoon12
, Hye Kyoung Yoon12
-
Journal of Pathology and Translational Medicine 2021;55(1):1-15.
DOI: https://doi.org/10.4132/jptm.2020.11.20
Published online: January 11, 2021
1Department of Pathology and Translational Genomics, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
2Department of Pathology, Seoul National University Bundang Hospital, Seoul National University College of Medicine, Seongnam, Korea
3Department of Pathology, Yeungnam University College of Medicine, Daegu, Korea
4Department of Pathology, Pusan National University Yangsan Hospital, Pusan National University College of Medicine, Yangsan, Korea
5Department of Pathology, Eulji University College of Medicine, Seoul, Korea
6Department of Pathology, Inje University Haeundae Paik Hospital, Busan, Korea
7Department of Pathology, National Cancer Center, Goyang, Korea
8Department of Hospital Pathology, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea
9Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
10Department of Pathology, Chonnam National University Medical School, Gwangju, Korea
11Department of Pathology, School of Medicine, Kyungpook National University, Kyungpook National University Chilgok Hospital, Daegu, Korea
12Department of Pathology, Busan Paik Hospital, Inje University, Busan, Korea
- Corresponding Author: Hye Kyoung Yoon, MD, PhD, Department of Pathology, Busan Paik Hospital, Inje University, 75 Bokji-ro, Busanjin-gu, Busan 47392, Korea Tel: +82-51-890-6628, Fax: +82-51-890-5825, E-mail: pathyoon@inje.ac.kr
- This article has been published jointly, with consent, in both Journal of Pathology and Translational Medicine and Journal of Breast Cancer.
© 2021 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
- Given the recent advances in management and understanding of breast cancer, a standardized pathology report reflecting these changes is critical. To meet this need, the Breast Pathology Study Group of the Korean Society of Pathologists has developed a standardized pathology reporting format for breast cancer, consisting of ‘standard data elements,’ ‘conditional data elements,’ and a biomarker report form. The ‘standard data elements’ consist of the basic pathologic features used for prognostication, while other factors related to prognosis or diagnosis are described in the ‘conditional data elements.’ In addition to standard data elements, all recommended issues are also presented. We expect that this standardized pathology report for breast cancer will improve diagnostic concordance and communication between pathologists and clinicians, as well as between pathologists inter-institutionally.
- All essential standard data elements for the report form are summarized in Table 1. In addition, all data elements including recommended issues as well as standard data elements can be found in Supplementary Table S1.
- Breast specimen types
- Breast specimen types can be roughly divided into wide excision and total mastectomy. Wide excision is defined as removal of only part of the breast tissue, with or without axillary contents, and includes specimens designated as excisional biopsy, segmental or partial mastectomy, lumpectomy, or quadrantectomy. Total mastectomy refers to removal of all breast tissue, which may include skin, nipple, or areola, with or without axillary contents, and includes simple mastectomy (total mastectomy without axillary node removal), skin-sparing mastectomy (total mastectomy with removal of the nipple and a narrow surrounding rim of skin), nipple-sparing mastectomy (total mastectomy without removal of skin or nipple), modified radical mastectomy (total mastectomy with axillary node dissection and with occasional removal of a small portion of the pectoralis muscle), and radical mastectomy (total mastectomy with pectoralis muscle removal and axillary node dissection) [6].
- Histological type
- The histopathologic classification of breast tumors in this paper is based on the WHO Classification of Breast Tumors, 5th edition [5] (Supplementary Table S2). The term “invasive breast carcinoma (IBC) of no special type (NST)” defines a large and heterogeneous group of IBCs that cannot be classified morphologically as any of the special histological types. The terms “invasive breast carcinoma, not otherwise specified (NOS),” “invasive ductal carcinoma,” or “infiltrating ductal carcinoma” are also acceptable; however, invasive mammary carcinoma of NST is not recommended.
- IBC-NST encompasses a wide spectrum of histological patterns, including some special morphological patterns. Oncocytic, lipid-rich, glycogen-rich, clear cell, and sebaceous carcinomas; carcinoma with medullary pattern; invasive carcinoma with neuroendocrine differentiation; and carcinomas with pleomorphic and choriocarcinomatous patterns are considered wide morphological patterns of IBC-NST regardless of the extent of differentiation or pattern. Breast carcinomas showing a specialized subtype in ≥90% of the tumor are designated as a pure special tumor type, such as mucinous cystadenocarcinoma or lobular, tubular, cribriform, mucinous, micropapillary, apocrine, or metaplastic carcinoma.
- The phrase “mixed IBC-NST and special subtype carcinoma” can be used when the special subtype comprises 10% to 90% of the carcinoma. For mixed tumors, overall percentage of the special subtype, grade and biomarkers status of both IBC-NST and special type carcinoma components should be reported. Carcinomas in which the special subtype comprises <10% should be classified as IBC-NST, with the optional comment of focal specialized subtype. Tumors lacking such specific features are designated IBC-NST, which accounts for the majority of IBC cases. Currently, estrogen receptor (ER) and human epidermal growth factor receptor type 2 (HER2) biomarker status are used for management purposes instead of histological subtype or pattern.
- The traditionally used classifications of medullary carcinoma, atypical medullary carcinoma, and carcinoma with medullary features found in the 4th edition of the WHO Breast Tumor Classification were removed in the 5th edition. Carcinomas formerly classified as these subtypes are now categorized as “IBC-NST with medullary pattern,” representing one end of the spectrum of tumor-infiltrating lymphocyte (TIL)-rich IBC-NSTs, rather than a distinct morphological subtype. IBC-NST with medullary pattern belongs to triple negative breast carcinomas, characterized by high expression of immune-related genes.
- Metaplastic carcinoma is a heterogeneous group of IBCs characterized by differentiation of the neoplastic epithelium toward squamous cells and/or mesenchymal-looking elements, including but not restricted to spindle, chondroid, and osseous cells. The type of metaplastic elements present may be recorded using a descriptive classification system.
- Tumor focality
- If multiple invasive carcinomas are present, tumor focality should be recorded. Multifocal tumors are associated with increased risk of lymph node involvement compared to similar unifocal disease [7-9], which reflects increased tumor load [10]. Counting the number of invasive foci is not essential but is recommended. When there is difficulty in determining whether two tumors are separate or not, microscopic examination of the tissue between the two masses should be performed. There are several occasions when multiple foci of invasion are present: extensive carcinoma in situ with multifocal invasion, invasive carcinoma with satellite foci, extensive lymphovascular invasion (LVI), multiple separate invasive carcinomas, invasive carcinomas after neoadjuvant chemotherapy, and transection of a single carcinoma [3].
- Except for cases presenting multiple separate invasive carcinomas, most multifocal tumors have similar appearance and immunophenotype to the largest tumor. When multifocal tumors have similar histology, only the largest tumor is tested for ER, progesterone receptor (PR), and HER2. If multifocal tumors have different histological subtypes and grades, it is recommended to evaluate ER, PR, and HER2 status of each component, separately [3,11].
- Tumor size
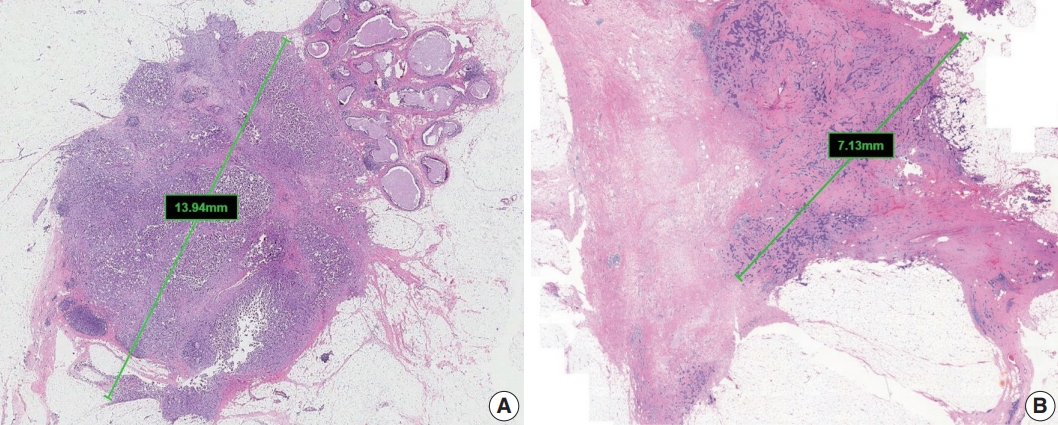
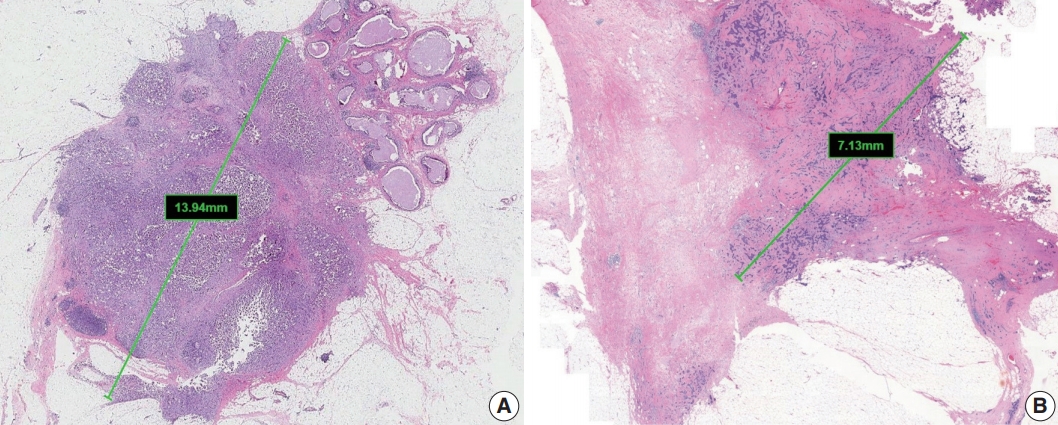
- The single greatest dimension of the largest invasive tumor is used to ascertain the pathologic tumor (pT) category, regardless of extent of accompanying in situ carcinoma [4] (Fig. 1A). Three-dimensional measurement of tumor size is essential. In cases in which it is difficult to determine the tumor size, information from imaging, gross findings, and microscopic evaluation should be used. For multifocal tumors, measurement of each tumor is recommended.
- The post-treatment pT category (ypT) is based on the largest contiguous focus of residual invasive carcinoma. Treatmentassociated fibrosis adjacent to residual tumor or between foci of residual invasive carcinoma is not included in the ypT category (Fig. 1B).
- Histological grade
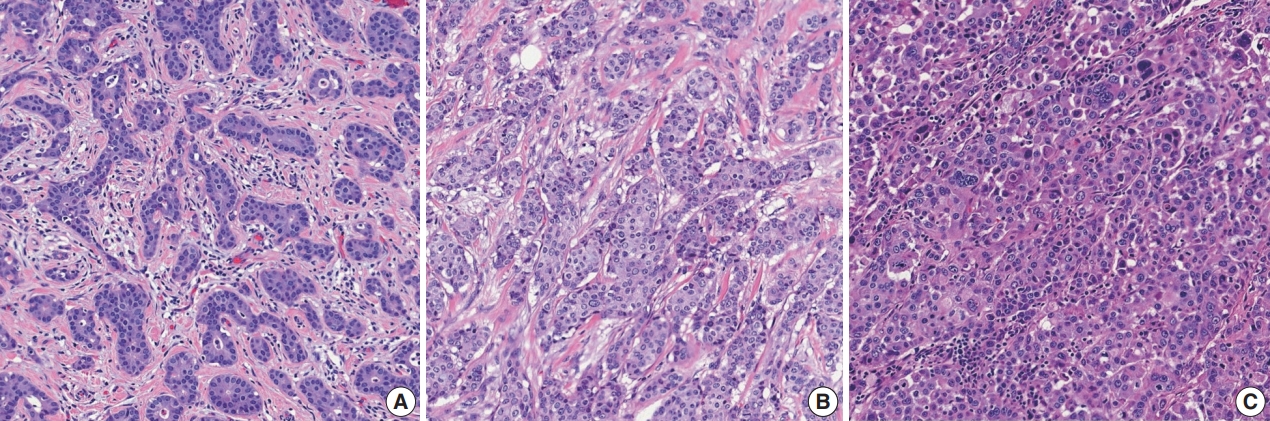
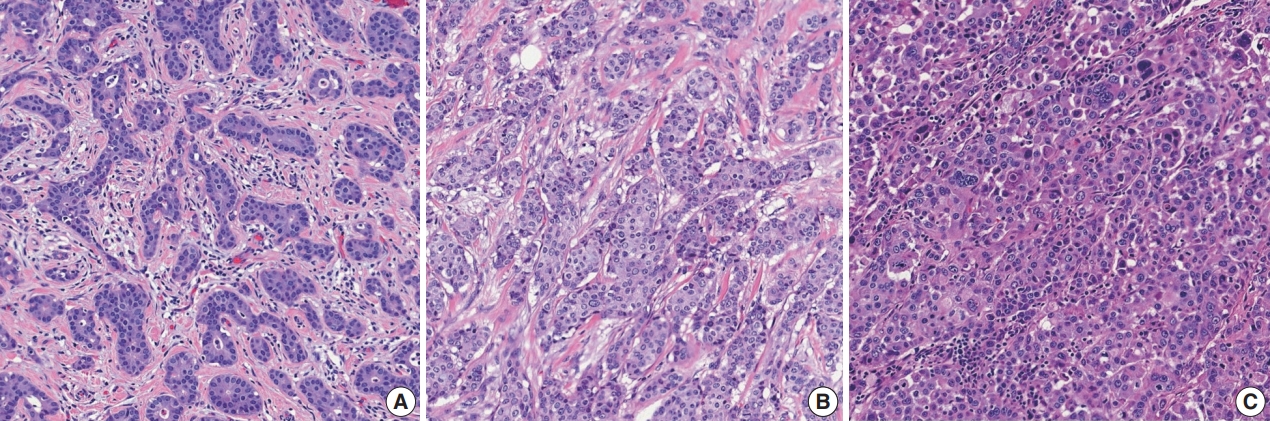
- Histological grading should be performed according to the Elston-Ellis modification of Bloom-Richardson grading [12]. Histological grading of IBCs is determined by three components: tubule formation, nuclear pleomorphism, and mitotic count (Supplementary Table S3). Tubule formation is assessed under low-power magnification. Scoring is performed according to the proportion of tumor cells forming tubules: more than 75% (score 1), 10%–75% (score 2), and less than 10% (score 3). Nuclear pleomorphism should be assessed in the area showing the highest degree of pleomorphism. A score of 1 is given to small (less than 1.5 times the size of benign epithelial cell nuclei) and uniform nuclei with finely dispersed chromatin. A score of 3 is given to large (more than two times the size of benign epithelial cell nuclei), vesicular, and pleomorphic nuclei with prominent nucleoli and irregular chromatin. A score of 2 is given to nuclei with characteristics that lie between those two categories. Mitotic count is the number of mitotic figures present in 10 high-power fields (HPFs). Counting should be performed in the hot spot (area with the most frequent mitotic figures), which is usually at the peripheral, leading edge of the tumor. Care should be taken not to count hyperchromatic and apoptotic nuclei. The cutoff points for mitotic count scores differ according to the field diameter of the 40× objective lens. The 5th edition WHO Breast Tumor Classification system recommends the use of number of mitoses per mm2 instead of number of mitoses per 10 HPFs for standardization [5].
- These three scores are summed, and the total score of 3–9 is used for overall tumor grade: score 3–5=grade 1, well differentiated; score 6–7=grade 2, moderately differentiated; score 8–9=grade 3, poorly differentiated (Fig. 2). The histological grade of IBC shows a strong correlation with prognosis [12,13].
- Ductal carcinoma in situ
- Ductal carcinoma in situ (DCIS) is concomitantly present in as many as 80% of IBC cases and is associated with increased risk of local recurrence after breast-conserving surgery [5].
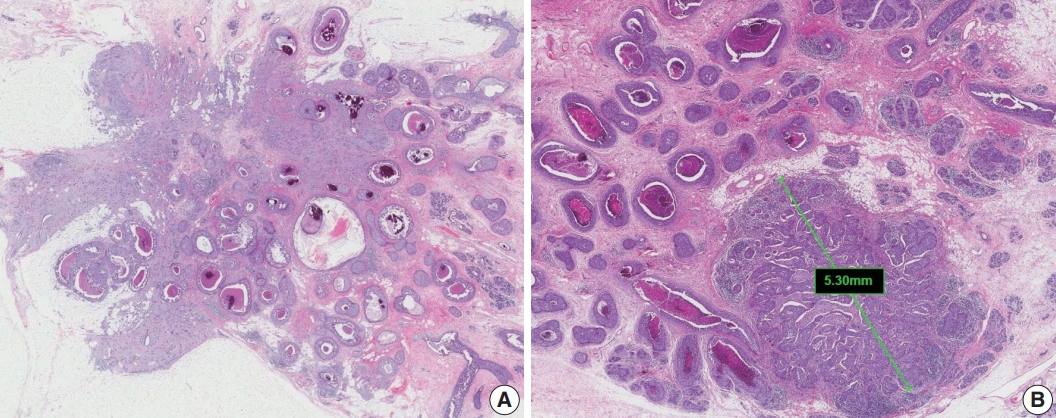
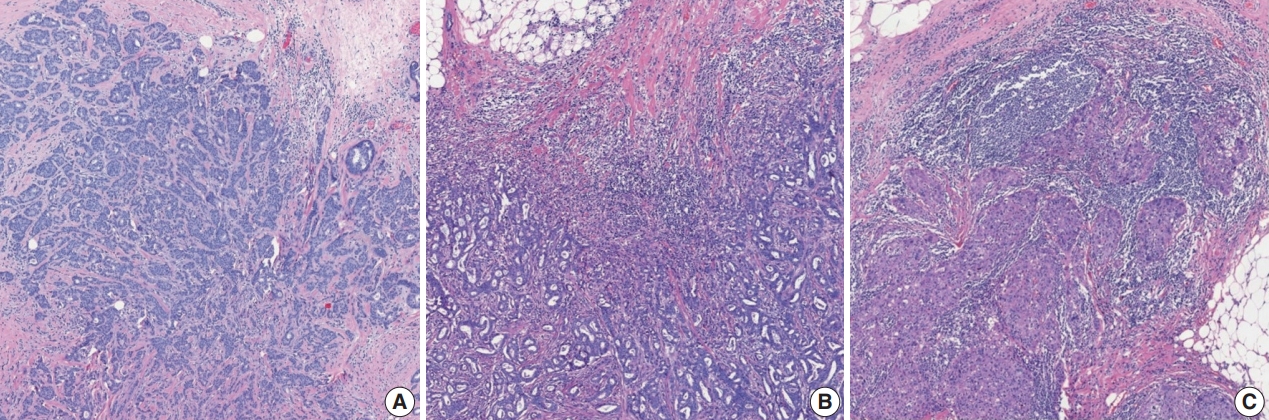
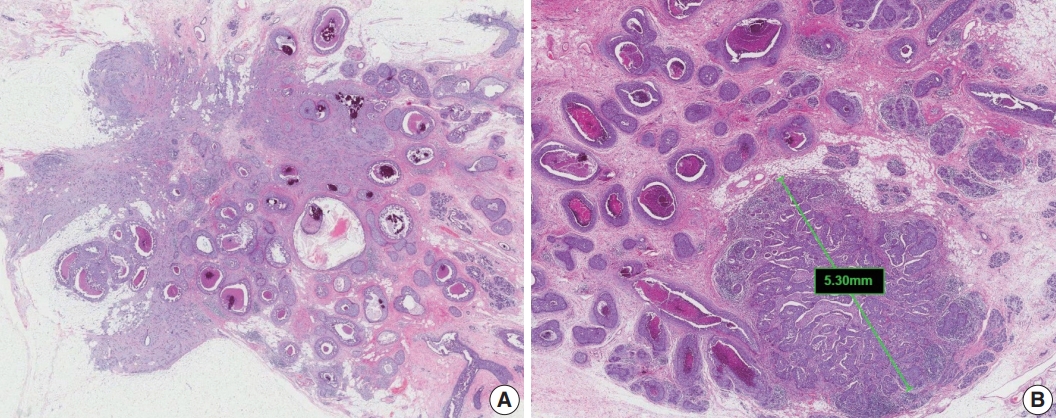
- Extensive intraductal component (EIC)-positive carcinoma is present when (1) DCIS is a major component (≥ 25%) of the area of invasive carcinoma and also is present outside the area of invasive carcinoma (Fig. 3A) or (2) there is extensive DCIS associated with a small (≤10 mm) invasive carcinoma (Fig. 3B) [3].
- The histological features of DCIS associated with increased risk of recurrence are large lesion size, high nuclear grade, certain architectural patterns, central necrosis, and positive surgical margin [5]. It is essential to report the features of DCIS, including nuclear grade, presence of necrosis, and extent of DCIS, in cases of EIC-positive carcinoma [3].
- Nuclear grade is determined according to pleomorphism, nuclear size, chromatin, nucleoli, mitoses, and orientation (Supplementary Table S4) and is predictive of clinical outcome (recurrence) [14]. Central (comedo) necrosis is easily detected at low magnification within the central portion of ducts affected by DCIS. Focal necrosis means necrosis in small foci or single-cell necrosis and is indistinct at low magnification.
- Reporting the architectural pattern of DCIS is not essential but is recommended. Comedo DCIS is characterized by high nuclear grade associated with central necrosis, often with calcification. Solid DCIS shows compact proliferation of tumor cells with low-to-intermediate nuclear grade that fills the entire duct. Small necrotic foci may be present. Cribriform DCIS is characterized by intraductal proliferation with a sieve-like or fenestrated pattern. The secondary lumens are round, rigid, and surrounded by low-to-intermediate grade nuclei or occasional high-grade nuclei. Micropapillary DCIS has papillary fronds that lack fibrovascular cores and that protrude into the ductal lumen in a regular distribution. Micropapillary DCIS tends to be extensive in distribution (multifocal and multicentric). Papillary DCIS contains arborizing papillae with thin fibrovascular cores. Although it may be seen only microscopically, papillary DCIS more commonly presents as a large mass [14]. Encapsulated papillary carcinoma without invasion and solid papillary carcinoma without invasion are unusual patterns of DCIS.
- Reporting the extent of DCIS is essential in cases of EIC-positive carcinoma. However, a precise measurement of the extent of DCIS may be difficult or, at times, impossible. There are several methods for estimating the extent of DCIS. If DCIS is confined to a single tissue block, it is possible to estimate the extent of DCIS by direct measurement of the histological slides. If the entire specimen is blocked sequentially, the extent of DCIS can be calculated by multiplying the number of slices involved by average slice thickness. If the specimen is sampled, rather than sequentially blocked in its entirety, the extent of DCIS can be estimated by counting the number of blocks with DCIS [3,15].
- Tumor extension
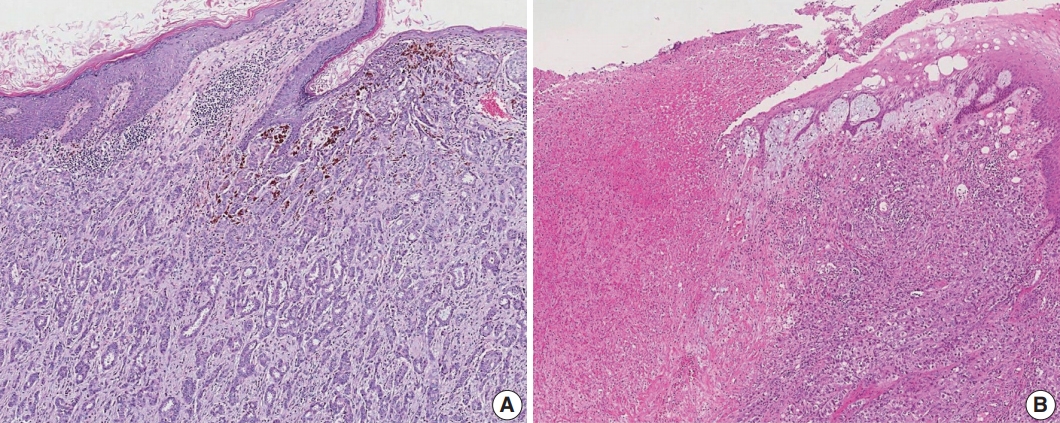
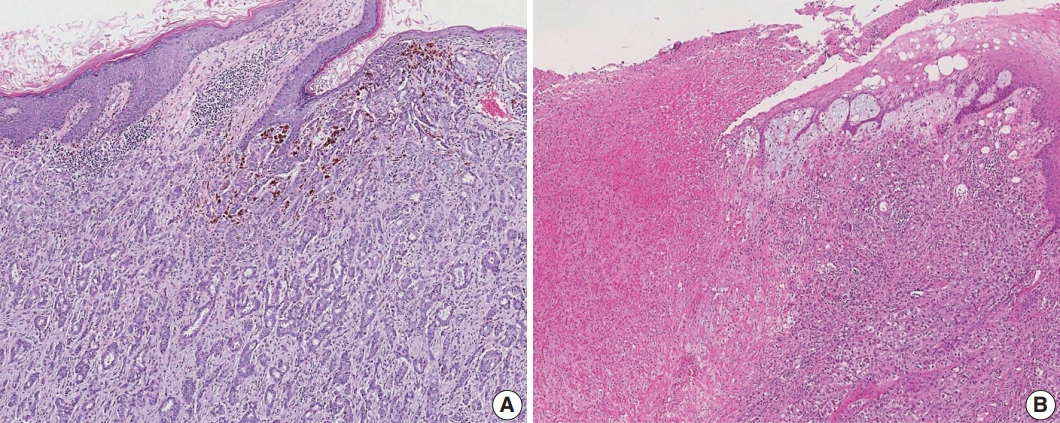
- Satellite tumor nodules in the skin are separate from the primary tumor and macroscopically identifiable. Skin and dermal satellite nodules identified only on microscopic examination and skin involvement without epidermal ulceration or skin edema (clinically peau d’orange) do not qualify as pT4b category (Fig. 4A, B). Such tumors should be categorized based on tumor size. Inflammatory carcinoma is categorized only when there are clinical symptoms of erythema and edema in more than one-third of the entire breast skin and not by the pathologic findings of tumor emboli in the dermal lymphatics.
- The chest wall includes ribs, intercostal muscles, and serratus anterior muscle but not the pectoralis muscles. Therefore, involvement of the pectoralis muscles in the absence of invasion of these chest wall elements does not constitute chest wall invasion, and cancers with such involvement are categorized based on tumor size.
- Resection margin
- Whenever possible, specimens should be oriented to identify specific margins for the pathologist. All identifiable margins should be evaluated for carcinoma involvement both grossly and microscopically [16].
- Orientation may be conducted using sutures or clips placed on the specimen surface or by other means of communication between the surgeon and pathologist and should be documented in the pathology report. Margins can be identified in several ways, including using multiple colored inks, submitting the margins in specific cassettes, or submitting each margin as a separately excised specimen.
- Margin status is listed as “positive” if there is ink on the cancer cells during pathologic margin evaluation. If the specimen is oriented, the specific site(s) of involvement should be reported. The approximate extent of margin involvement can be reported as follows: unifocal, 1 focal area of carcinoma (<4 mm) at the margin; multifocal, 2 or more carcinoma foci at the margin; extensive, carcinoma present at the margin over a broad front (>5 mm).
- In lobular carcinoma in situ (LCIS), assessment of resection margin is optional. However, for pleomorphic type LCIS, evaluation of resection margin is recommended.
- Regional lymph node metastasis
- Most patients with invasive carcinoma will have lymph nodes sampled for pathologic regional lymph node (pN) categorization [4]. All lymph nodes must be examined histologically [4]. The nodes commonly examined include sentinel nodes, nonsentinel nodes, nodes from axillary dissections, and intramammary nodes. When the total number of sentinel and nonsentinel nodes removed is less than 6, the AJCC “sn” modifier is used.
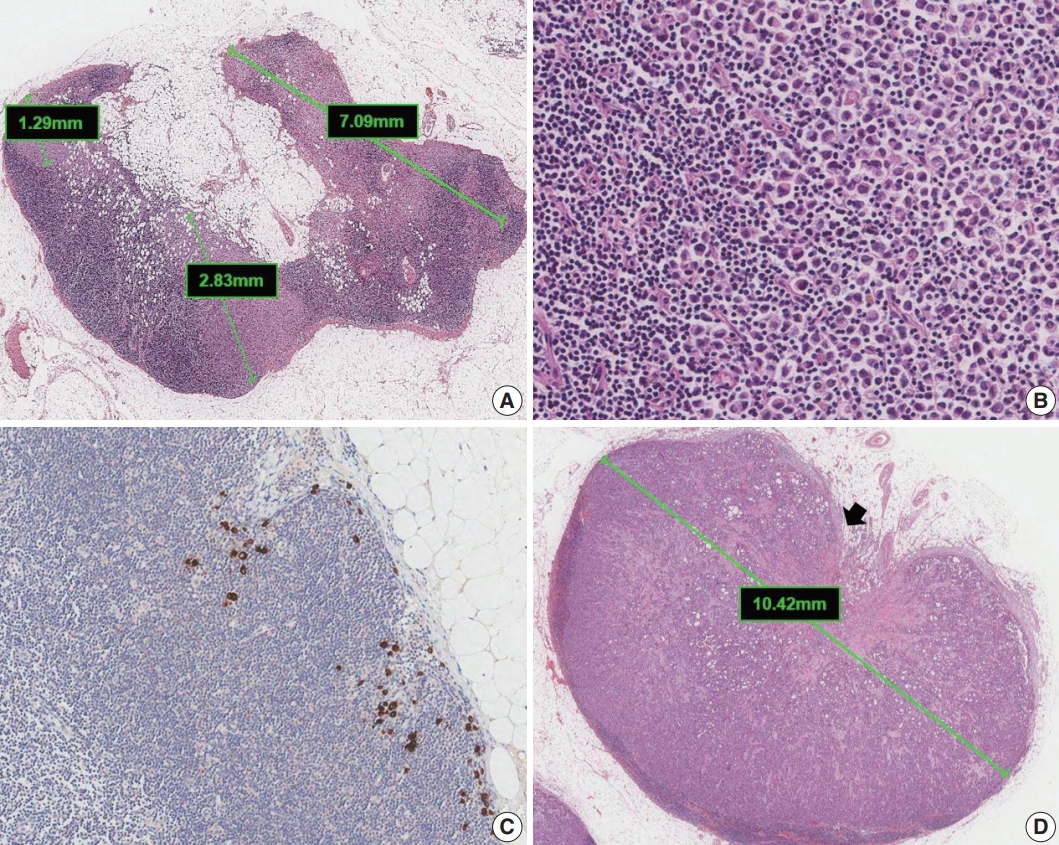
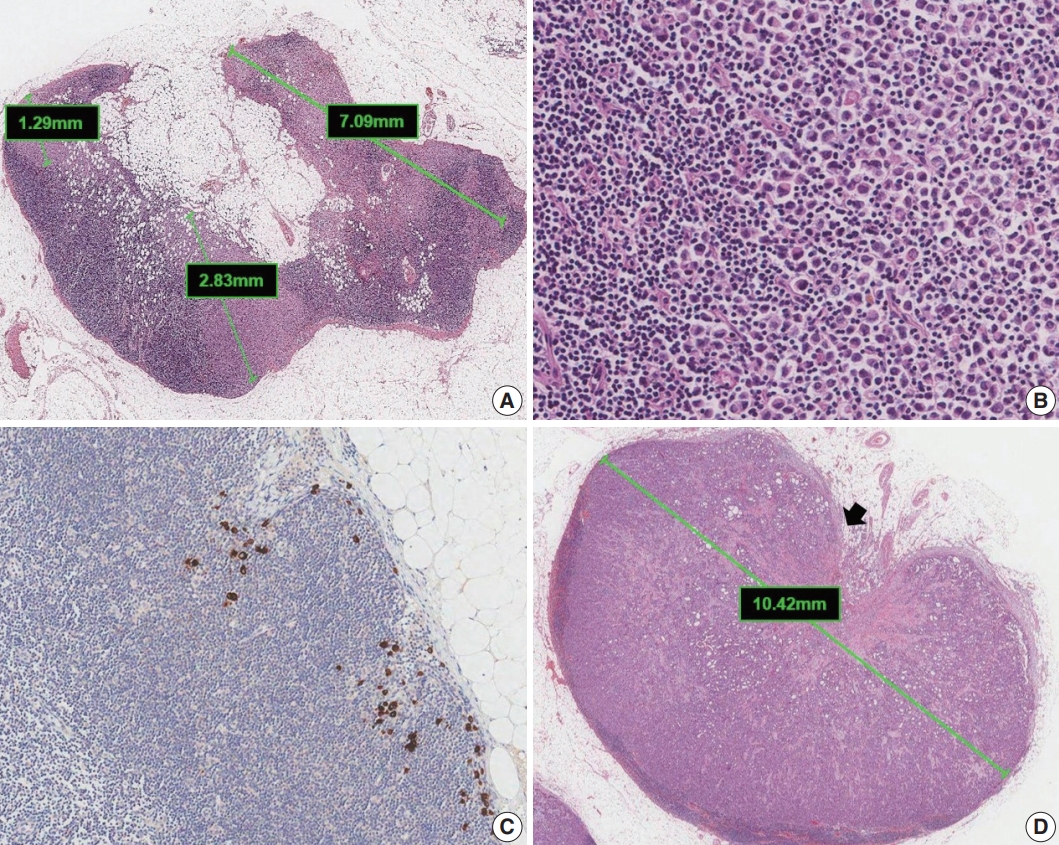
- Metastases are classified into three groups based on size: isolated tumor cells (ITCs), micrometastases, and macrometastases [4]. ITCs are defined as single cells, small clusters of cells no larger than 0.2 mm, or no more than 200 cells in a single cross section. The AJCC states that a cluster is a group of cells in contact with each other (confluent or contiguous). Cells that are not touching each other should be considered independent and measured independently. In cases of multiple clusters of tumor cells within a lymph node, only the largest should be considered when determining N category (Fig. 5A). The AJCC states that the size of the tumor should include both the tumor cells and the surrounding desmoplastic reaction. Some carcinomas, particularly lobular carcinomas, may metastasize as individual single cells and not as clusters and present as a dispersed pattern of nodal metastases (Fig. 5B). In such cases, single cells are measured separately. If fewer than 200 tumor cells are present in a node cross section, then classification of ITCs is recommended (Fig. 5C). Nodes containing only ITCs are not included in the total number of positive nodes when determining N category, so cases with only ITCs are classified as node negative (pN0 (i+)). Micrometastases measure greater than 0.2 mm but not greater than 2 mm and/or comprise more than 200 cells in a single cross section. If only micrometastases are present on lymph node examination, the N category is pN1mi. If at least 1 macrometastasis is present, nodes with micrometastases are included in the total number of positive nodes. Any lesion where the largest cluster is greater than 2 mm represents a macrometastasis (Fig. 5D).
- Extranodal extension is defined as the presence of full-thickness (i.e., into and through) lymph node capsular invasion, as seen with metastatic tumor invasion of extranodal fat with or without an associated desmoplastic stromal response (Fig. 5D). The area of extranodal extension is included when measuring the overall size of the lymph node metastasis. Extranodal extension is a marker of poor prognosis in breast cancer patients, and the status of extranodal extension should be reported [17]. Reporting of extranodal extension size based on a 2-mm cutoff is incorporated into the CAP reporting guidelines, but more evidence is needed for this practice to become widely accepted [3,17].
- When cancerous nodules that are not associated with residual lymph node tissue are present in the axillary fat, the AJCC states that these nodules should be classified as positive lymph nodes [4]. However, if there is surrounding normal breast parenchyma or DCIS, then cancerous nodules in the axillary fat should be classified as invasive carcinoma and not as a nodal metastasis.
- The post-treatment pN (ypN) classification system is the same as that for pre-treatment lymph nodes. Only the largest contiguous focus of residual tumor in the node evaluation is used for determining N category; any treatment-associated fibrosis is not included [3,4].
- Lymphovascular invasion
- LVI is associated with local recurrence and reduced survival [18]. Strict criteria or immunohistochemistry (IHC) stains have been proposed to differentiate LVI from DCIS and retraction artifacts [3]. If a limited area is involved in LVI, a measurement in millimeters can be given. Alternatively, LVI can be quantified as focal or extensive, with ‘extensive’ defined as one or more foci in more than one block [19].
- The presence of pure LVI without stromal invasion after neoadjuvant therapy may be called ypTX and should not be classified as pathologic complete response (pCR) [20].
- Pathologic stage classification [4]
- Pathologic stage classification according to the AJCC 8th edition should be reported as a standard data element [4]. Classification of primary tumor (T), regional lymph nodes (N), and distant metastasis (M) by pathologic examination is denoted by the prefix “p” (pT, pN, and pM). The descriptor “m” is used when invasive cancer is observed in multiple foci, and the prefix “r” is used for recurrent cancer. If the patient has undergone neoadjuvant chemotherapy, hormonal therapy, immunotherapy, or radiation therapy before surgery, the prefix “y” is used. pM0 is not a valid category. When distant metastases cannot be confirmed by pathologic examination, staging can be performed by combining pT, pN, and the clinical evaluation of metastases (cM).
- Criteria for each pT category are summarized in Supplementary Table S5. For multiple invasive cancers, use the tumor with the highest T category for classification and staging, and use the descriptor “m” or number of invasive cancers in parentheses (e.g., T2(m) or T2(3)). For simultaneous bilateral breast cancers, staging should be conducted separately because they are considered independent tumors in different organs.
- Criteria for each pN category are summarized in Supplementary Table S6. If no lymph nodes were submitted for evaluation, record pNX. It is not the pathologist’s obligation to record the pN status by integrating the previous pathologic results. pN1a, pN2a, and pN3a refer to metastases in 1 to 3, 4 to 9, and 10 or more axillary lymph nodes, respectively, with at least one macrometastasis. If the specimen contains internal mammary lymph nodes, infraclavicular lymph nodes, or supraclavicular lymph nodes with metastases, or if clinically metastatic internal mammary lymph node(s) are identified, refer to the AJCC staging manual for accurate lymph node categorization. A regional lymph node with direct extension of the primary tumor or a tumor nodule in a regional lymph node area should be considered as a positive node.
- When nodal metastasis is confirmed by fine-needle aspiration cytology or core needle biopsy without further resection of nodes, use the “f” modifier (e.g., pN(f)).
- The pM category is assigned only if metastasis larger than 0.2 mm (pM1) is histologically confirmed. When staging after neoadjuvant therapy, the classification should remain M1 regardless of responsiveness to therapy, if the case was confirmed to be M1 prior to therapy.
STANDARD DATA ELEMENTS
pT category
pN category
pM category
- All conditional data elements for this report form are summarized in Table 2.
- Perineural invasion
- Perineural invasion (PNI) is infrequently observed in IBC, occurring in approximately 1% of cases, perhaps in part because nerves of notable size are not numerous in mammary tissues [21].
- PNI may occur more frequently in IBC-NST than in the other histological subtypes. It tends to occur in high-grade tumors, where it is frequently associated with LVI, but it has not been proven to have independent prognostic significance [21-24]. PNI can also be observed in some benign lesions, such as sclerosing adenosis, as well as in DCIS.
- Tumor border
- The tumor margins of IBC can be grossly described as ill-demarcated, well-demarcated (circumscribed), or mixed [21].
- Approximately one-third of tumors have grossly circumscribed margins. However, some carcinomas that appear to have circumscribed margins grossly exhibit an invasive growth pattern microscopically [21]. Grossly ill-demarcated tumors tend to be larger, and they are more likely to have axillary metastases than those with circumscribed margins [21].
- Microcalcification
- DCIS/invasive carcinoma found in biopsies performed for microcalcifications will almost always be at the site of the microcalcifications or in close proximity [25,26]. The presence of targeted microcalcifications in the specimen can be confirmed by radiography.
- The pathologist needs to confirm that the specimen has been sampled from the lesion responsible for the microcalcifications. Microcalcifications are commonly present in secretions and/or in necrotic materials [5]. The radiological and pathologic correlation of all microcalcifications, including information about the presence and site of microcalcifications (e.g., invasive carcinoma, DCIS, benign lesion, or mixed), should be indicated [5]. Information about the microcalcifications can be an important consideration when correlating imaging findings with the pathologic diagnosis, when guiding further management of the disease, and when identifying recurrent carcinoma in the breast or metastatic diseases [21].
- Tumor-infiltrating lymphocytes
- The prognostic and predictive value of TILs in breast cancer has been studied extensively [27,28]. TILs are lymphocytes present in the stroma of a tumor or inside tumor cell nests. Assessment of TIL level in hematoxylin and eosin (H&E) sections can be easily performed. The International Immuno-Oncology Biomarker Working Group on Breast Cancer published guidelines for evaluation of TIL level in H&E sections of invasive breast cancer in 2014, and they later extended this method to DCIS, metastatic tumor deposits, and specimens obtained after neoadjuvant chemotherapy [7,29,30]. In brief, TIL level is determined by measuring the percentage of the total stromal area, excluding tumor necrosis and crush artifacts, occupied by mononuclear inflammatory cells, including plasma cells, within the borders of the invasive carcinoma. TILs are usually evaluated in increments of 10% (e.g., <10%, 10%–19%, 20%–29%) (Fig. 6). Since distribution of TILs is usually not even throughout the tumor tissue, assessment of the average number of TILs, without focusing on hot spots, is recommended. The International Immuno-Oncology Biomarker Working Group showed that proper training of pathologists could achieve more consistent results with regard to evaluation of TILs in ring studies and suggested potential pitfalls in assessment of TILs [31,32].
- Effect of treatment
- Many classifications have been proposed to evaluate the pathologic status of breast cancer after treatment, including those of Chevallier [33], Sataloff [34], the National Surgical Adjuvant Breast and Bowel Project (NSABP) B-18 [35], MillerPayne [36], the Residual Cancer Burden (RCB) system [37], the Clinical-Pathologic Stage-Estrogen/Grade (CPS-EG) system [38], the Residual Disease in Breast and Nodes (RDBN) system [39], and the AJCC [4]. Among these classifications, the AJCC and the RCB calculator are the most widely used systems to measure residual disease [40].
- The RCB index is calculated from the following five variables [37]: (1) primary tumor bed area (mm2), (2) overall cancer cellularity (%), (3) percentage of carcinoma in the tumor bed that is in situ disease (%), (4) number of positive lymph nodes, and (5) diameter of the largest lymph node metastasis (mm).
- Primary tumor bed area is the two largest dimensions between invasive tumor cells, even if these are widely scattered and separated by treatment-induced fibrosis. Overall cancer cellularity is the overall percentage of the residual tumor bed area that is occupied by carcinoma (invasive and in situ). It is assessed in each slide, and the average is calculated using all fields that fall within the perimeter of the largest cross-sectional area of residual tumor bed, even those with very low cellularity or no disease [37]. The same method can be used for the in situ component to assess the percentage of cancer that is in situ disease [37]. Unlike the AJCC ypN category, the number of positive lymph nodes includes the number of lymph nodes with ITCs. The diameter of the largest lymph node metastasis used in the RCB system may be different from that used for AJCC staging because the former includes intervening treatment-related fibrosis [40,41].
- A mathematical formula combines these variables into a continuous index to define four RCB classes: RCB-0 for pCR and RCB 1 to 3, representing progressively greater extent of residual cancer [37]. A web-based calculator and detailed instructions for calculating RCB indices are publicly available (http://www3.mdanderson.org/app/medcalc/index.cfm?pagename=jsconvert3).
- As the presence of positive lymph nodes after treatment represents a worse prognosis even when there is no residual invasive carcinoma in the breast [42,43], several classification systems and the CAP cancer protocol have recommended that effect of treatment be evaluated in both the breast and lymph nodes [3,4,34,37,39,44,45].
CONDITIONAL DATA ELEMENTS
- Determination of biomarker status, including ER, PR, and HER2 status, is essential for newly diagnosed IBC. ER, PR, and HER2 should be evaluated according to the current American Society of Clinical Oncology (ASCO)/CAP guidelines [46,47]. Currently, there are no established guidelines regarding re-evaluation of biomarkers in post-treatment specimens. However, it is recommended that ER, PR, and HER2 testing be repeated on post-treatment invasive carcinomas, such as when there was insufficient invasive tumor tissue or negative or equivocal results on pre-treatment core biopsy; when biopsies were performed and biomarkers assessed at other institutions; or when post-treatment tumors display heterogeneous morphology or no response to therapy.
- All essential data elements for reporting biomarker status are summarized in Table 3. In addition, all data elements for biomarkers including all recommended issues as well as essential data elements can be found in Supplementary Table S7.
- ER and PR status
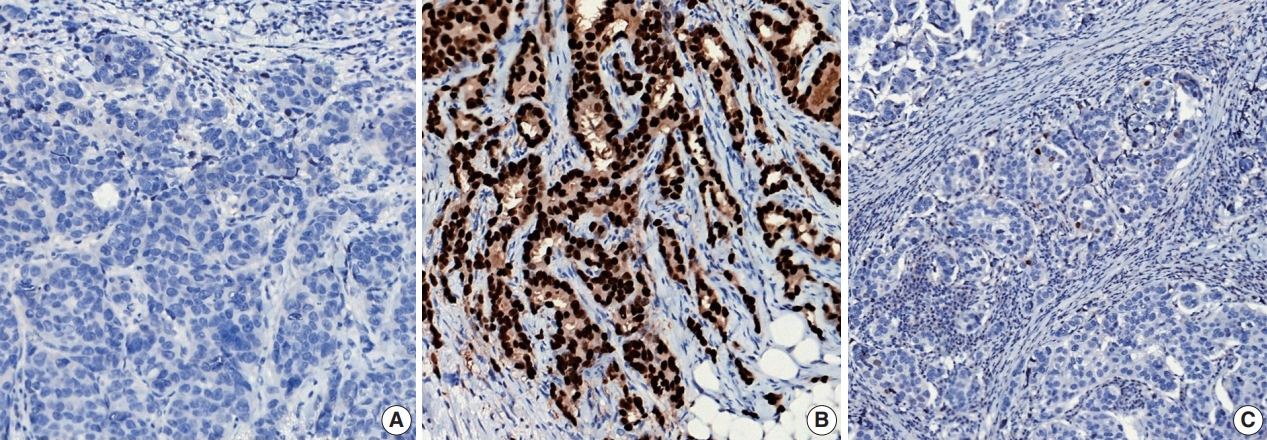
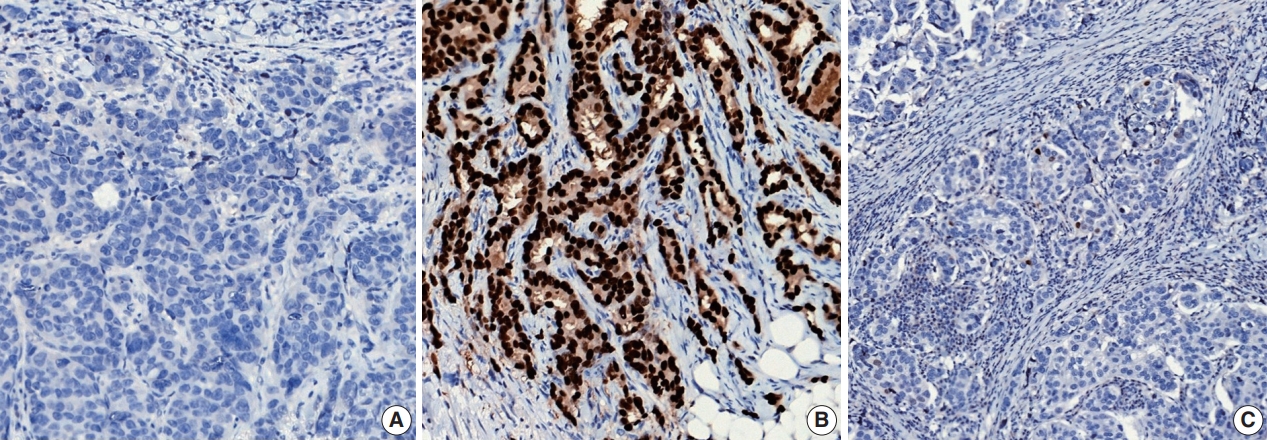
- ER or PR positivity is defined when more than 1% of tumor cells demonstrate nuclear positivity [46] (Fig. 7A, B). The proportion of positive cells can be reported as a specific percentage or as a discrete range. The intensity is defined as the degree of nuclear positivity (weak, moderate, strong) and is recommended for reporting.
- Alternatively, the Allred scoring system can be used to evaluate ER and PR status semi-quantitatively. The Allred score combines the percentage of stained nuclei (0, <1%, 1%–10%, 11%–33%, 34%–66%, and >67%) and the average intensity of the immunoreactivity (0, 1, 2, and 3) for a final score out of 8. Scores between 0 and 2 are classified as a negative result, while scores 3 and above are considered positive. Very rarely, carcinomas with <1% positive cells and intensity scores of 2 or 3 are classified as positive, as the total score would be 3 or 4. However, responses to hormonal therapy in these tumors have not been specifically proven [48].
- Recently, it has been recommended that invasive carcinomas with 1%–10% positive ER staining be reported as ER low positive [46] (Fig. 7C). There are data that suggest that invasive cancers with these results are heterogeneous in both behavior and biology and often have gene expression profiles similar to those of ER-negative cancers [46].
- The status of controls should be reported in cases with ER/PR negative or ER low positive tumors [46]. On-slide controls are ideal and, wherever possible, routine evaluation of internal normal epithelial elements or inclusion of normal breast sections (or other appropriate control) on each tested slide is recommended.
- Inadequate samples and technical issues, such as prolonged cold ischemia time, insufficient sample amount, severe processing artifacts, inappropriate external/internal controls, and unavailable information on pre-analytical variables associated with fixation, may cause difficulties in interpretation of results, including production of false-negative results. These cases are uninterpretable, and repeat staining of another block or specimen is recommended [46].
- HER2 status
- IHC and in situ hybridization (ISH) are regarded as standard methods to assess HER2 status in breast cancer. The ASCO/CAP have jointly released guidelines and recommendations on HER2 testing in breast cancer since 2007, and, recently, have updated these guidelines to provide clear instructions for HER2 testing and accurate determination of HER2 status in breast cancer [49].
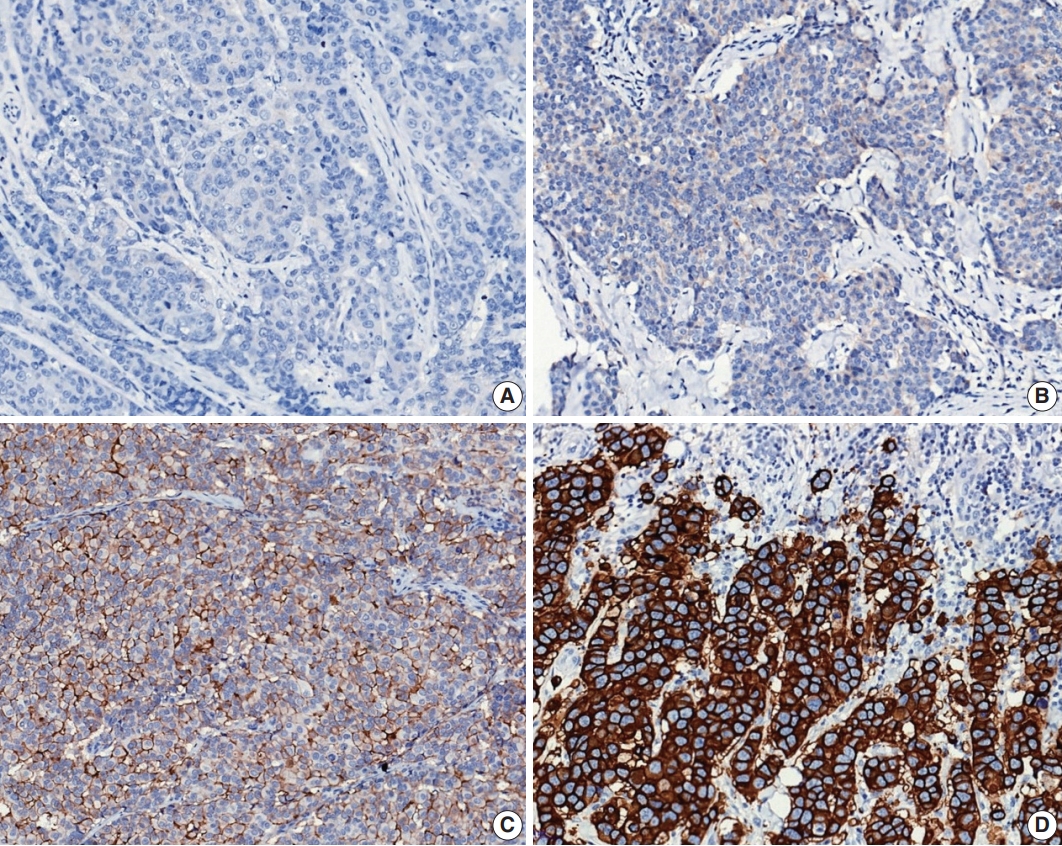
- Currently, HER2 status determined by IHC and ISH should be interpreted based on the 2018 updated ASCO/CAP guidelines [47]. HER2 IHC status should be assessed using a semiquantitative scoring system (Fig. 8A–D). For HER2 IHC equivocal (2+) cases, HER2 status should be confirmed by ISH.
- Interpretation of HER2 ISH is performed by counting at least 20 cells in the invasive tumor area. Scanning of entire slides or use of IHC slides prior to counting is mandatory to define the areas of potential HER2 amplification [50]. Please refer to a previous article for interpretation of HER2 heterogeneity [49].
- For a diagnostic approach using HER2 ISH, concomitant IHC review for dual-probe ISH groups 2 to 4 is required in the updated guidelines [47]. In laboratories using single-probe ISH assays, concomitant IHC review is included as part of the interpretation of all ISH assay results [47]. By this approach, the HER2 ISH status is designated as positive or negative, with no equivocal category. Determining HER2 ISH status using dual-probe ISH is summarized in Supplementary Table S8. In reporting the results of HER2 ISH, final HER2 ISH status (negative or positive), number of counted cells, HER2/centromere on chromosome 17 (CEP17) ratio, average HER2 copy number per cell, average CEP17 copy number per cell, and designation of ISH group (optional) should be included.
- Ki-67 index
- The Ki-67 index is defined by the percentage of tumor cells with positive Ki-67 nuclear staining out of all tumor cells counted in a given field. A high Ki-67 index is regarded as a prognostic marker associated with high risk of recurrence and as a predictive marker for treatment response in breast cancer [51-54]. However, there remain controversies in its use as a standard prognostic or predictive biomarker owing to high inter-observer variability and lack of a standardized measurement method. Currently, assessment of Ki-67 can be performed applying recommendations from the International Ki-67 in Breast Cancer Working Group [55] in clinical practice. Briefly, at least three high-power (40× objective) fields should be selected to represent the spectrum of staining on initial overview of the whole section. If there are clear hot spots, data from these should be included in the overall average score. Scoring should involve the counting of at least 500 malignant invasive cells (and preferably at least 1,000 cells). Currently, computational imaging analysis methods are widely used for Ki-67 quantification, but their superiority over the direct counting method is unclear. When morphological analysis is used, the number of tumor cells counted and the number of tumor cells with positive Ki-67 nuclear staining should be included in the report form.
BIOMARKERS
- In accordance with recent advances in collective understanding of breast cancer biology and in treatment of breast cancer patients, a committee of the Breast Pathology Study Group of the Korean Society of Pathologists presents in this publication a ‘Standardized Pathology Report for Breast Cancer.’ This report form is composed of ‘standard data elements,’ ‘conditional data elements,’ and a biomarker report form to guide diagnosis, prognosis, and treatment of breast cancer patients. It is our hope that this report will lead to standardization of the pathologic diagnosis of breast cancer and to improvement in communication between clinicians and pathologists, as well as between pathologists.
CONCLUSION
Supplementary Materials
Ethics Statement
Not applicable.
Author contributions
Conceptualization: HKY. Project administration: SYP, YKB. Supervision: HKY. Writing—original draft: SYC, SYP, YKB, JYK, EKK, WGK, YK, AL, HJL, JSL, JYP. Writing—review & editing: SYC, SYP, YKB, GG, HKY. Approval of final manuscript: all authors.
Conflicts of Interest
SYP, the editor-in-chief and YKB and GG, contributing editors of the Journal of Pathology and Translational Medicine, were not involved in the editorial evaluation or decision to publish this article. All remaining authors have declared no conflicts of interest.
Funding Statement
No funding to declare.


- 1. Kang SY, Kim YS, Kim Z, et al. Breast cancer statistics in Korea in 2017: data from a breast cancer registry. J Breast Cancer 2020; 23: 115-28. ArticlePubMedPMCPDF
- 2. Li X, Oprea-Ilies GM, Krishnamurti U. New developments in breast cancer and their impact on daily practice in pathology. Arch Pathol Lab Med 2017; 141: 490-8. ArticlePubMedPDF
- 3. Fitzgibbons PL, Connolly JL, Bose S, et al. Protocol for the examination of resection specimens from patients with invasive carcinoma of the breast [Internet]. Northfield: College of American Pathologists, c2020 [cited 2020 Sep 10]. Available from: https://documents.cap.org/protocols/cp-breast-invasive-resection-20-4400.pdf.
- 4. Hortobagyi GN, Connolly JL, D'Orsi CJ, et al. Breast. In: Amin MB, Edge S, Greene F, eds. AJCC cancer staging manual. 8th ed. New York: Springer, 2017; 589-636.
- 5. WHO Classification of Tumours Editorial Board. WHO classification of tumours: breast tumours. 5th ed. Lyon: International Agency for Research on Cancer, 2019.
- 6. Lester SC, Bose S, Chen YY, et al. Protocol for the examination of specimens from patients with invasive carcinoma of the breast. Arch Pathol Lab Med 2009; 133: 1515-38. ArticlePubMedPDF
- 7. Salgado R, Aftimos P, Sotiriou C, Desmedt C. Evolving paradigms in multifocal breast cancer. Semin Cancer Biol 2015; 31: 111-8. ArticlePubMed
- 8. Andea AA, Wallis T, Newman LA, Bouwman D, Dey J, Visscher DW. Pathologic analysis of tumor size and lymph node status in multifocal/multicentric breast carcinoma. Cancer 2002; 94: 1383-90. ArticlePubMed
- 9. Weissenbacher TM, Zschage M, Janni W, et al. Multicentric and multifocal versus unifocal breast cancer: is the tumor-node-metastasis classification justified? Breast Cancer Res Treat 2010; 122: 27-34. ArticlePubMedPDF
- 10. Coombs NJ, Boyages J. Multifocal and multicentric breast cancer: does each focus matter? J Clin Oncol 2005; 23: 7497-502. ArticlePubMed
- 11. Choi Y, Kim EJ, Seol H, et al. The hormone receptor, human epidermal growth factor receptor 2, and molecular subtype status of individual tumor foci in multifocal/multicentric invasive ductal carcinoma of breast. Hum Pathol 2012; 43: 48-55. ArticlePubMed
- 12. Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology 1991; 19: 403-10. ArticlePubMed
- 13. Rakha EA, El-Sayed ME, Lee AH, et al. Prognostic significance of Nottingham histologic grade in invasive breast carcinoma. J Clin Oncol 2008; 26: 3153-8. ArticlePubMed
- 14. Jaffer S, Bleiweiss IJ. Histologic classification of ductal carcinoma in situ. Microsc Res Tech 2002; 59: 92-101. ArticlePubMed
- 15. Grin A, Horne G, Ennis M, O'Malley FP. Measuring extent of ductal carcinoma in situ in breast excision specimens: a comparison of 4 methods. Arch Pathol Lab Med 2009; 133: 31-7. ArticlePubMedPDF
- 16. Morrow M, Van Zee KJ, Solin LJ, et al. Society of Surgical Oncology-American Society for Radiation Oncology-American Society of Clinical Oncology consensus guideline on margins for breast-conserving surgery with whole-breast irradiation in ductal carcinoma in situ. Pract Radiat Oncol 2016; 6: 287-95. ArticlePubMedPMC
- 17. Choi AH, Blount S, Perez MN, et al. Size of extranodal extension on sentinel lymph node dissection in the American College of Surgeons Oncology Group Z0011 trial era. JAMA Surg 2015; 150: 1141-8. ArticlePubMed
- 18. Mohammed RA, Martin SG, Mahmmod AM, et al. Objective assessment of lymphatic and blood vascular invasion in lymph nodenegative breast carcinoma: findings from a large case series with long-term follow-up. J Pathol 2011; 223: 358-65. ArticlePubMed
- 19. Provenzano E, Bossuyt V, Viale G, et al. Standardization of pathologic evaluation and reporting of postneoadjuvant specimens in clinical trials of breast cancer: recommendations from an international working group. Mod Pathol 2015; 28: 1185-201. ArticlePubMedPDF
- 20. Baker GM, King TA, Schnitt SJ. Evaluation of breast and axillary lymph node specimens in breast cancer patients treated with neoadjuvant systemic therapy. Adv Anat Pathol 2019; 26: 221-34. ArticlePubMed
- 21. Hoda SA, Brogi E, Koerner FC, Rosen PP. Rosen's breast pathology. Philadelphia: Wolters Kluwer, 2014.
- 22. Duraker N, Caynak ZC, Turkoz K. Perineural invasion has no prognostic value in patients with invasive breast carcinoma. Breast 2006; 15: 629-34. ArticlePubMed
- 23. McCready DR, Chapman JA, Hanna WM, et al. Factors affecting distant disease-free survival for primary invasive breast cancer: use of a log-normal survival model. Ann Surg Oncol 2000; 7: 416-26. ArticlePubMedPDF
- 24. McCready DR, Chapman JA, Hanna WM, et al. Factors associated with local breast cancer recurrence after lumpectomy alone: postmenopausal patients. Ann Surg Oncol 2000; 7: 562-7. ArticlePubMedPDF
- 25. Owings DV, Hann L, Schnitt SJ. How thoroughly should needle localization breast biopsies be sampled for microscopic examination? A prospective mammographic/pathologic correlative study. Am J Surg Pathol 1990; 14: 578-83. PubMed
- 26. Silverstein MJ, Lagios MD, Recht A, et al. Image-detected breast cancer: state of the art diagnosis and treatment. J Am Coll Surg 2005; 201: 586-97. PubMed
- 27. Aaltomaa S, Lipponen P, Eskelinen M, et al. Lymphocyte infiltrates as a prognostic variable in female breast cancer. Eur J Cancer 1992; 28a: 859-64. ArticlePubMed
- 28. Wein L, Savas P, Luen SJ, Virassamy B, Salgado R, Loi S. Clinical validity and utility of tumor-infiltrating lymphocytes in routine clinical practice for breast cancer patients: current and future directions. Front Oncol 2017; 7: 156.ArticlePubMedPMC
- 29. Hendry S, Salgado R, Gevaert T, et al. Assessing tumor-infiltrating lymphocytes in solid tumors: a practical review for pathologists and proposal for a standardized method from the International Immuno-Oncology Biomarkers Working Group: part 2: TILs in melanoma, gastrointestinal tract carcinomas, non-small cell lung carcinoma and mesothelioma, endometrial and ovarian carcinomas, squamous cell carcinoma of the head and neck, genitourinary carcinomas, and primary brain tumors. Adv Anat Pathol 2017; 24: 311-35. ArticlePubMedPMC
- 30. Dieci MV, Radosevic-Robin N, Fineberg S, et al. Update on tumorinfiltrating lymphocytes (TILs) in breast cancer, including recommendations to assess TILs in residual disease after neoadjuvant therapy and in carcinoma in situ: a report of the International Immuno-Oncology Biomarker Working Group on Breast Cancer. Semin Cancer Biol 2018; 52: 16-25. ArticlePubMed
- 31. Denkert C, Wienert S, Poterie A, et al. Standardized evaluation of tumor-infiltrating lymphocytes in breast cancer: results of the ring studies of the international immuno-oncology biomarker working group. Mod Pathol 2016; 29: 1155-64. PubMed
- 32. Kos Z, Roblin E, Kim RS, et al. Pitfalls in assessing stromal tumor infiltrating lymphocytes (sTILs) in breast cancer. NPJ Breast Cancer 2020; 6: 17.PubMedPMC
- 33. Chevallier B, Roche H, Olivier JP, Chollet P, Hurteloup P. Inflammatory breast cancer. Pilot study of intensive induction chemotherapy (FEC-HD) results in a high histologic response rate. Am J Clin Oncol 1993; 16: 223-8. PubMed
- 34. Sataloff DM, Mason BA, Prestipino AJ, Seinige UL, Lieber CP, Baloch Z. Pathologic response to induction chemotherapy in locally advanced carcinoma of the breast: a determinant of outcome. J Am Coll Surg 1995; 180: 297-306. PubMed
- 35. Fisher ER, Wang J, Bryant J, Fisher B, Mamounas E, Wolmark N. Pathobiology of preoperative chemotherapy: findings from the National Surgical Adjuvant Breast and Bowel (NSABP) protocol B-18. Cancer 2002; 95: 681-95. ArticlePubMed
- 36. Ogston KN, Miller ID, Payne S, et al. A new histological grading system to assess response of breast cancers to primary chemotherapy: prognostic significance and survival. Breast 2003; 12: 320-7. ArticlePubMed
- 37. Symmans WF, Peintinger F, Hatzis C, et al. Measurement of residual breast cancer burden to predict survival after neoadjuvant chemotherapy. J Clin Oncol 2007; 25: 4414-22. ArticlePubMed
- 38. Mittendorf EA, Jeruss JS, Tucker SL, et al. Validation of a novel staging system for disease-specific survival in patients with breast cancer treated with neoadjuvant chemotherapy. J Clin Oncol 2011; 29: 1956-62. ArticlePubMedPMC
- 39. Chollet P, Abrial C, Durando X, et al. A new prognostic classification after primary chemotherapy for breast cancer: residual disease in breast and nodes (RDBN). Cancer J 2008; 14: 128-32. ArticlePubMed
- 40. Mrkonjic M, Berman HK, Done SJ, Youngson B, Mulligan AM. Breast specimen handling and reporting in the post-neoadjuvant setting: challenges and advances. J Clin Pathol 2019; 72: 120-32. ArticlePubMed
- 41. Bossuyt V. Processing and reporting of breast specimens in the neoadjuvant setting. Surg Pathol Clin 2018; 11: 213-30. ArticlePubMed
- 42. Hennessy BT, Hortobagyi GN, Rouzier R, et al. Outcome after pathologic complete eradication of cytologically proven breast cancer axillary node metastases following primary chemotherapy. J Clin Oncol 2005; 23: 9304-11. ArticlePubMed
- 43. von Minckwitz G, Untch M, Blohmer JU, et al. Definition and impact of pathologic complete response on prognosis after neoadjuvant chemotherapy in various intrinsic breast cancer subtypes. J Clin Oncol 2012; 30: 1796-804. ArticlePubMed
- 44. Pinder SE, Provenzano E, Earl H, Ellis IO. Laboratory handling and histology reporting of breast specimens from patients who have received neoadjuvant chemotherapy. Histopathology 2007; 50: 409-17. ArticlePubMed
- 45. Smith IC, Heys SD, Hutcheon AW, et al. Neoadjuvant chemotherapy in breast cancer: significantly enhanced response with docetaxel. J Clin Oncol 2002; 20: 1456-66. ArticlePubMed
- 46. Allison KH, Hammond ME, Dowsett M, et al. Estrogen and progesterone receptor testing in breast cancer: ASCO/CAP guideline update. J Clin Oncol 2020; 38: 1346-66. ArticlePubMed
- 47. Wolff AC, Hammond MEH, Allison KH, et al. Human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline focused update. J Clin Oncol 2018; 36: 2105-22. PubMed
- 48. Harvey JM, Clark GM, Osborne CK, Allred DC. Estrogen receptor status by immunohistochemistry is superior to the ligand-binding assay for predicting response to adjuvant endocrine therapy in breast cancer. J Clin Oncol 1999; 17: 1474-81. ArticlePubMed
- 49. Ahn S, Woo JW, Lee K, Park SY. HER2 status in breast cancer: changes in guidelines and complicating factors for interpretation. J Pathol Transl Med 2020; 54: 34-44. ArticlePubMedPDF
- 50. Wolff AC, Hammond ME, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol 2013; 31: 3997-4013. PubMed
- 51. Luporsi E, Andre F, Spyratos F, et al. Ki-67: level of evidence and methodological considerations for its role in the clinical management of breast cancer: analytical and critical review. Breast Cancer Res Treat 2012; 132: 895-915. ArticlePubMedPDF
- 52. Yerushalmi R, Woods R, Ravdin PM, Hayes MM, Gelmon KA. Ki67 in breast cancer: prognostic and predictive potential. Lancet Oncol 2010; 11: 174-83. ArticlePubMed
- 53. Viale G, Giobbie-Hurder A, Regan MM, et al. Prognostic and predictive value of centrally reviewed Ki-67 labeling index in postmenopausal women with endocrine-responsive breast cancer: results from Breast International Group Trial 1-98 comparing adjuvant tamoxifen with letrozole. J Clin Oncol 2008; 26: 5569-75. ArticlePubMedPMC
- 54. Penault-Llorca F, André F, Sagan C, et al. Ki67 expression and docetaxel efficacy in patients with estrogen receptor-positive breast cancer. J Clin Oncol 2009; 27: 2809-15. ArticlePubMed
- 55. Dowsett M, Nielsen TO, A'Hern R, et al. Assessment of Ki67 in breast cancer: recommendations from the International Ki67 in Breast Cancer working group. J Natl Cancer Inst 2011; 103: 1656-64. PubMedPMC
REFERENCES
Figure & Data
References
Citations

- Adenoid Cystic Carcinoma of Breast Associated With an Incidental Radial Scar: A Cyto‐Histopathology Correlation
Rallapalli Rajyalakshmi, Valasapalli Rajani, Tanuku Sreedhar, Kollabathula Arpitha
Diagnostic Cytopathology.2026;[Epub] CrossRef - Refined risk stratification in residual triple-negative breast cancer after neoadjuvant therapy using residual cancer burden class and lymphovascular invasion
Tae Hoon Lee, Hyunwoo Lee, Jeong Yun Jang, Won Park, Won Kyung Cho, Eun Yoon Cho, Jin Seok Ahn, Yeon Hee Park, Seok Jin Nam, Seok Won Kim, Jeong Eon Lee, Haeyoung Kim
Breast Cancer Research and Treatment.2026;[Epub] CrossRef - Navigating discrepancies: The assessment of residual lymphovascular invasion in breast carcinoma after neoadjuvant treatment
Anikó Kovács, Åsa Rundgren-Sellei, Gunilla Rask, Annette Bauer, Anna Bodén, Johannes van Brakel, Eugenia Colón-Cervantes, Anna Ehinger, Johan Hartman, Balazs Acs
The Breast.2025; 82: 104519. CrossRef - Residual pure intralymphatic carcinoma component only (lymphovascular tumor emboli without invasive carcinoma) after neoadjuvant chemotherapy is associated with poor outcome: Not pathologic complete response
Hyunwoo Lee, Yunjeong Jang, Yoon Ah Cho, Eun Yoon Cho
Human Pathology.2024; 145: 1. CrossRef - Sentinel lymph node biopsy in patients with ductal carcinomain situ: systematic review and meta-analysis
Matthew G. Davey, Colm O’Flaherty, Eoin F. Cleere, Aoife Nohilly, James Phelan, Evan Ronane, Aoife J. Lowery, Michael J. Kerin
BJS Open.2022;[Epub] CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure
- Related articles








Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
Fig. 6.
Fig. 7.
Fig. 8.
| Breast specimen type | ||
| □ Wide excision (specify) | ||
| □ Total mastectomy (specify) | ||
| Specimen laterality | ||
| □ Right | ||
| □ Left | ||
| □ Unspecified | ||
| Tumor location | ||
| □ UOQ | ||
| □ LOQ | ||
| □ UIQ | ||
| □ LIQ | ||
| □ Central | ||
| □ Unspecified | ||
| Histologic type | ||
| □ Invasive breast carcinoma of no special type (specify for special morphological patterns) | ||
| □ Invasive lobular carcinoma (specify for subtype) | ||
| □ Tubular carcinoma | ||
| □ Cribriform carcinoma | ||
| □ Mucinous carcinoma | ||
| □ Invasive micropapillary carcinoma | ||
| □ Carcinoma with apocrine differentiation | ||
| □ Metaplastic carcinoma (specify for subtype) | ||
| □ Other rare subtype (specify) | ||
| Tumor focality | ||
| □ Unifocal | ||
| □ Multifocal | ||
| Tumor size | ||
| × × cm | ||
| Histologic grade | ||
| □ Grade I (Low) □ Grade II (Intermediate) □ Grade III (High) | ||
| Ductal carcinoma in situ (DCIS) | ||
| □ Not identified | ||
| □ Present (□ EIC-positive, □ EIC-negative) | ||
| Nuclear grade | ||
| □ Grade I (Low) | ||
| □ Grade II (Intermediate) | ||
| □ Grade III (High) | ||
| Necrosis | ||
| □ Not identified | ||
| □ Present (□ focal, □ central) | ||
| Extent of DCIS (for EIC-positive case) | ||
| Estimated size: × cm | ||
| No. of blocks with DCIS/No. of blocks examined: / | ||
| Lobular carcinoma in situ (in case of invasive lobular carcinoma) | ||
| □ Not identified | ||
| □ Present (□ classic type, □ pleomorphic type) | ||
| Tumor extension | ||
| Skin | ||
| □ Not present | ||
| □ Present (□ uninvolved, □ involved, without skin ulceration/ with skin ulceration/with satellite skin nodule) | ||
| Skeletal muscle | ||
| □ Not present | ||
| □ Present (□ uninvolved, □ involved in pectoralis muscle, □ involved in pectoralis muscle and chest wall) | ||
| Resection margin | ||
| □ Cannot be assessed | ||
| □ Positive for carcinoma | ||
| Location (specify): invasive carcinoma/DCIS/invasive carcinoma and DCIS (unifocal, multifocal, extensive) | ||
| □ Uninvolved by invasive carcinoma and/or DCIS | ||
| Distance from closest margin : mm from margin | ||
| Regional lymph node metastasis | ||
| Total number of lymph nodes examined: | ||
| Number of lymph nodes involved with metastases: | ||
| (sentinel node: / , nonsentinel node: / ) | ||
| Size of largest metastasis: mm | ||
| Extranodal extension: Not identified/Present | ||
| Lymphovascular invasion | ||
| □ Not identified | ||
| □ Present | ||
| Pathologic stage classification (pTNM, AJCC 8th edition) | ||
| TNM descriptors: □ m □ r □ y | ||
| Primary tumor (pT): | ||
| Regional lymph nodes (pN): | ||
| Perineural invasion | |
| □ Not identified | |
| □ Present | |
| Tumor border | |
| □ Not applicable | |
| □ Well-demarcated/Circumscribed | |
| □ Ill-demarcated | |
| □ Mixed | |
| Microcalcification | |
| □ Not identified | |
| □ Present in invasive carcinoma | |
| □ Present in DCIS | |
| □ Present in non-neoplastic tissue | |
| Tumor-infiltrating lymphocytes (TILs) | |
| % | |
| Treatment effect (RCB class) | |
| □ RCB class 0 □ RCB class I □ RCB class II □ RCB class III | |
| RCB index: | |
| Additional pathologic findings | |
| Estrogen receptor | |
| □ Positive (≥ 1% of tumor cells with nuclear positivity) | |
| □ Negative (< 1%) | |
| Progesterone receptor | |
| □ Positive (≥ 1% of tumor cells with nuclear positivity) | |
| □ Negative (< 1%) | |
| HER2 status by immunohistochemistry | |
| □ Negative (0) | |
| □ Negative (1+) | |
| □ Equivocal (2+) | |
| □ Positive (3+) | |
| HER2 status by in situ hybridization | |
| □ ISH negative | |
| □ ISH positive | |
| No. of counted cells: | |
| HER2/CEP17 ratio: | |
| Average HER2 copy number per cell: | |
| Average CEP17 copy number per cell: | |
| Ki-67 index | |
| Ki-67 index: % | |
UOQ, upper outer quadrant; LOQ, lower outer quadrant; UIQ, upper inner quadrant; LIQ, lower inner quadrant; EIC, extensive intraductal component; AJCC, American Joint Committee on Cancer.
DCIS, ductal carcinoma in situ; RCB, Residual Cancer Burden.
HER2, human epidermal growth factor receptor type 2; ISH, in situ hybridization; CEP17, centromere on chromosome 17.

 E-submission
E-submission