Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 57(1); 2023 > Article
-
Review
A standardized pathology report for gastric cancer: 2nd edition -
Young Soo Park1,*
 , Myeong-Cherl Kook2,*
, Myeong-Cherl Kook2,* , Baek-hui Kim3,*
, Baek-hui Kim3,* , Hye Seung Lee4,*
, Hye Seung Lee4,* , Dong-Wook Kang5
, Dong-Wook Kang5 , Mi-Jin Gu6
, Mi-Jin Gu6 , Ok Ran Shin7
, Ok Ran Shin7 , Younghee Choi8
, Younghee Choi8 , Wonae Lee9
, Wonae Lee9 , Hyunki Kim10
, Hyunki Kim10 , In Hye Song1
, In Hye Song1 , Kyoung-Mee Kim11
, Kyoung-Mee Kim11 , Hee Sung Kim12
, Hee Sung Kim12 , Guhyun Kang13
, Guhyun Kang13 , Do Youn Park14
, Do Youn Park14 , So-Young Jin15
, So-Young Jin15 , Joon Mee Kim16
, Joon Mee Kim16 , Yoon Jung Choi17
, Yoon Jung Choi17 , Hee Kyung Chang18
, Hee Kyung Chang18 , Soomin Ahn11
, Soomin Ahn11 , Mee Soo Chang19
, Mee Soo Chang19 , Song-Hee Han20
, Song-Hee Han20 , Yoonjin Kwak4
, Yoonjin Kwak4 , An Na Seo21
, An Na Seo21 , Sung Hak Lee22
, Sung Hak Lee22 , Mee-Yon Cho23
, Mee-Yon Cho23 , The Gastrointestinal Pathology Study Group of the Korean Society of Pathologists
, The Gastrointestinal Pathology Study Group of the Korean Society of Pathologists -
Journal of Pathology and Translational Medicine 2023;57(1):1-27.
DOI: https://doi.org/10.4132/jptm.2022.12.23
Published online: January 15, 2023
1Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
2Center for Gastric Cancer, National Cancer Center, Goyang, Korea
3Department of Pathology, Korea University Guro Hospital, Seoul, Korea
4Department of Pathology, Seoul National University Hospital, Seoul National University College of Medicine, Seoul, Korea
5Department of Pathology, Chungnam National University Sejong Hospital, Chungnam National University School of Medicine, Sejong, Korea
6Department of Pathology, Yeungnam University College of Medicine, Daegu, Korea
7Department of Hospital Pathology, Uijeongbu St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Uijeongbu, Korea
8Department of Pathology, Hallym University Dongtan Sacred Heart Hospital, Hwaseong, Korea
9Department of Pathology, Dankook University College of Medicine, Cheonan, Korea
10Department of Pathology, Yonsei University College of Medicine, Seoul, Korea
11Department of Pathology and Translational Genomics, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
12Department of Pathology, Chung-Ang University Hospital, Chung-Ang University College of Medicine, Seoul, Korea
13LabGenomics Clinical Laboratories, Seongnam, Korea
14St. Maria Pathology Laboratory, Busan, Korea
15Department of Pathology, Soonchunhyang University Seoul Hospital, Seoul, Korea
16Department of Pathology, Inha University School of Medicine, Incheon, Korea
17Department of Pathology, Yongin Severance Hospital, Yonsei University College of Medicine, Yongin, Korea
18Department of Pathology, Kosin University Gospel Hospital, Kosin University College of Medicine, Busan, Korea
19Department of Pathology, Seoul National University Boramae Hospital, Seoul National University College of Medicine, Seoul, Korea
20Department of Pathology, Dong-A University College of Medicine, Busan, Korea
21Department of Pathology, School of Medicine, Kyungpook National University, Kyungpook National University Chilgok Hospital, Daegu, Korea
22Department of Hospital Pathology, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, Seoul, Korea
23Department of Pathology, Wonju Severance Christian Hospital, Yonsei University Wonju College of Medicine, Wonju, Korea
-
Corresponding Author: Sung Hak Lee, MD, PhD, Department of Hospital Pathology, Seoul St. Mary’s Hospital, College of Medicine, The Catholic University of Korea, 222 Banpodaero, Seocho-gu, Seoul 06591, Korea Tel: +82-2-2258-1617, Fax: +82-2-2258-1627, E-mail: hakjjang@catholic.ac.kr
Corresponding Author: Mee-Yon Cho, MD, PhD, Department of Pathology, Wonju Severance Christian Hospital, Yonsei University Wonju College of Medicine, 20 Ilsan-ro, Wonju 26426, Korea Tel: +82-33-741-1553, Fax: +82-33-731-6590, E-mail: meeyon@yonsei.ac.kr - *Young Soo Park, Myeong-Cherl Kook, Baek-hui Kim, and Hye Seung Lee contributed equally to this work.
This article has been published jointly, with consent, in both Journal of Pathology and Translational Medicine and Journal of Gastric Cancer.
© 2023 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
- The first edition of ‘A Standardized Pathology Report for Gastric Cancer’ was initiated by the Gastrointestinal Pathology Study Group of the Korean Society of Pathologists and published 17 years ago. Since then, significant advances have been made in the pathologic diagnosis, molecular genetics, and management of gastric cancer (GC). To reflect those changes, a committee for publishing a second edition of the report was formed within the Gastrointestinal Pathology Study Group of the Korean Society of Pathologists. This second edition consists of two parts: standard data elements and conditional data elements. The standard data elements contain the basic pathologic findings and items necessary to predict the prognosis of GC patients, and they are adequate for routine surgical pathology service. Other diagnostic and prognostic factors relevant to adjuvant therapy, including molecular biomarkers, are classified as conditional data elements to allow each pathologist to selectively choose items appropriate to the environment in their institution. We trust that the standardized pathology report will be helpful for GC diagnosis and facilitate large-scale multidisciplinary collaborative studies.
- This standard pathology report is for use with primary gastric carcinomas. Neuroendocrine tumors, lymphomas, gastrointestinal stromal tumors, and other sarcomas are excluded. Carcinomas involving the esophagogastric junction (EGJ) with a center ≤2 cm into the proximal stomach are considered to be distal esophageal carcinoma and excluded, as defined in the American Joint Committee on Cancer (AJCC), 8th edition [8]. This pathology report is also used for residual (post-chemotherapy or post-endoscopic resection) carcinomas. The report forms for pathologic diagnosis from radical resection and endoscopic resection specimens are shown in Tables 1 and 2, respectively.
- Radical resection specimens
- The type of surgical procedure should be mentioned in the surgical record.
- The gross type of each lesion should be recorded individually. The classification of early gastric cancer (EGC) uses the Japanese guideline (subclassification of type 0) [9], and classification of advanced gastric cancer (AGC) uses the Borrmann classification. The unclassifiable type is Borrmann type 5, according to the Japanese guideline [9]. The gross type is determined by macroscopic examination. If there is discrepancy between the macroscopic and microscopic findings, i.e., EGC on macroscopic examination but tumor invades the proper muscle microscopically (AGC), the macroscopic type should remain as the gross finding and not be corrected according to the microscopic finding. In such cases, the following descriptions are recommended: AGC, mimicking EGC type X or EGC, mimicking Borrmann type X. If the lesion is AGC grossly, at least four representative sections should be submitted for microscopic examination, including the deepest invasion, and ink should be applied at the serosal surface nearest the tumor. If the lesion is EGC grossly, grid mapping should be performed at 4 to 5 mm width.
- Any treatment before surgical resection should be recorded when applicable. If there are residual tumor foci, it should be mentioned that these are residual tumors. In post-chemotherapy gastrectomy situations, representative sections are sufficient if the lesion is large and obvious. However, the entire tumor bed must be microscopically examined when the representative sections contain no residual cancer cells or the residual lesion is small or inconspicuous grossly. For post-endoscopic resection specimens, the entire tumor bed should be submitted for microscopic evaluation.
- Tumor focality should record whether it is a single lesion or multiple lesions. Multiple lesions should be evaluated individually both macroscopically and microscopically in descending order from the tumor with the deepest level of invasion. However, regional lymph node metastasis, associated findings, and separate lesions are listed only for the deepest lesion.
- The description of the tumor location is recorded in two parts: involvement and center. The involvement of the tumor uses up to three portions from the esophagus to duodenum beginning with the most involved area. The delineation of the upper, middle, and lower thirds of the stomach follows the Japanese guideline [9]. The center of the tumor is described using a combination of locations according to the International Classification of Diseases for Oncology classification [10] (cardia, fundus, body, antrum, pylorus, lesser curvature, greater curvature) plus the anterior wall and posterior wall [11]. If none of those options appropriately describes the location of the tumor, other can be used.
- The tumor size is recorded using the largest dimension of the tumor [11]. Secondary or tertiary dimensions can be measured as conditional data elements. However, the tumor size is not used in the current staging of GC [8], and it is sometimes very difficult to measure accurately, such as in Borrmann type 4 cancer. For scattered residual tumor foci following previous treatment, it is recommended to measure the maximum diameter that includes all foci [12].
- Although preoperative chemotherapy has not been established as a standard treatment for patients in Korea [5], studies have shown survival benefits in local AGC in European [13], Asian [14], and Korean patients [15]. Therefore, the need to adequately evaluate the tumor response to chemotherapy is increasing [16]. Various tumor regression grading (TRG) methods are available for gastrointestinal cancers [17,18]. The Becker system [19] is one that has been proposed for GC. The previous edition of “A Standardized Pathology Report for Gastric Cancer” [3] used the Japanese guideline [9]. The Becker and Japanese systems both estimate the proportion of residual tumor and use it as a cutoff value between TRGs. However, because some tumors have more abundant fibrosis than tumor cells (before chemotherapy), estimation of the residual tumor proportion could show low concordance between observers [20,21]. Therefore, we suggest a new TRG system: the modified Ryan system currently recommended in the College of American Pathologists (CAP) guideline [11] and the second edition of the standardized pathology report for colorectal cancer in Korea [22]. It is a descriptive fourtier system that evaluates residual cancer rather than fibrosis as none, single cells or rare small groups, more than single cells but evident tumor response, and extensive residual cancer cells. Acellular mucin pools and necrotic or degenerative cells are not considered to be residual cancer [8]. Only the primary tumor is evaluated in this TRG, but tumor regression of the regional lymph nodes [16,23] can be reported as a conditional data element when there is evidence of partial (viable cancer cells with regressive changes) or complete tumor regression (only fibrosis, mucin pool, or foam cells without viable cells) in the regional lymph nodes. Evidence suggests that the presence of tumor regression in the lymph nodes is associated with better clinical outcomes [24,25].
- The depth of the tumor invasion follows the AJCC 8th edition [8] and Japanese guidelines [9]. Notably, the Japanese guideline does not accept carcinoma in situ (pTis). In the AJCC 8th edition, pTis is defined as an intraepithelial tumor without invasion of the lamina propria, which is equivalent to high-grade dysplasia. pT1b is subdivided into sm1, sm2, and sm3. If cancer cells are present below an imaginary line dividing the submucosa and proper muscle, the case is considered pT2 even if the cancer cells are not actually within the muscle fibers. If there is no proper muscle layer due to ulceration, and the cancer cells are below the imaginary line drawn at the lower border of the proper muscle, the case is considered pT3. Invasion of the omentum and perigastric fat is considered pT3. Ink should be applied at the serosal surface nearest the tumor during gross examination to properly evaluate serosal (visceral peritoneum) invasion. The case is considered pT4a if the cancer cells are adherent to or exposed beyond mesothelial cells. Because the mesocolon and gastric serosa (including the greater and lesser omentum) have different embryological origins, invasion of the mesocolon should be classified as pT4b. However, some areas are tightly fused, such as the posterior wall of the antrum, the gastric serosa, and the anterior side of the transverse mesocolon. Therefore, the Japanese guideline indicates that invasion of the transverse mesocolon is not pT4b unless it extends to the colic vessels or penetrates the posterior surface of the mesocolon [9]. Some cases can be either pT4a or pT4b, depending on the site of the tumor. Invasion of the pancreas capsule is considered pT4b. Direct duodenal or esophageal invasion is not considered pT4b. Any involvement of other organs, such as the liver, pancreas, colon, spleen, diaphragm, or kidney, should be recorded. Cancer cells within lymphatic or vascular spaces are not considered in the determination of invasion depth [8]. The presence of lymphatic or vascular invasion should be recorded separately in parentheses (e.g., tumor invades proper muscle [involvement of subserosa by lymphatic emboli]).
- Resection margin
- The distance from the proximal or distal resection margin is the length from the edge of the carcinoma to the nearest resection margin. It is important to locate the true resection margin in the gross specimen, especially when the stomach is opened along the lesser curvature or obliquely along the anterior or posterior wall. In some cases, cancer cells approach the resection margin much more closely than can be observed grossly (cancer spreading underneath the mucosa). Therefore, the resection margin is finalized in a microscopic evaluation. The circumferential and radial resection margin statuses can be reported as conditional data elements. Determination of the circumferential margin is often required if the tumor is located near the EGJ.
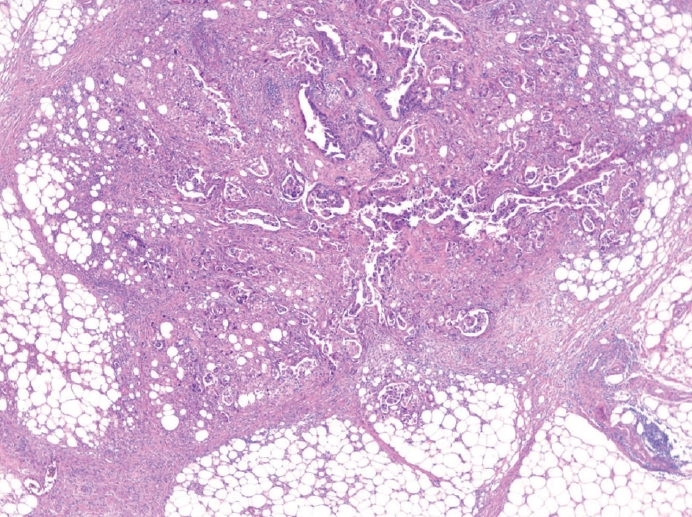
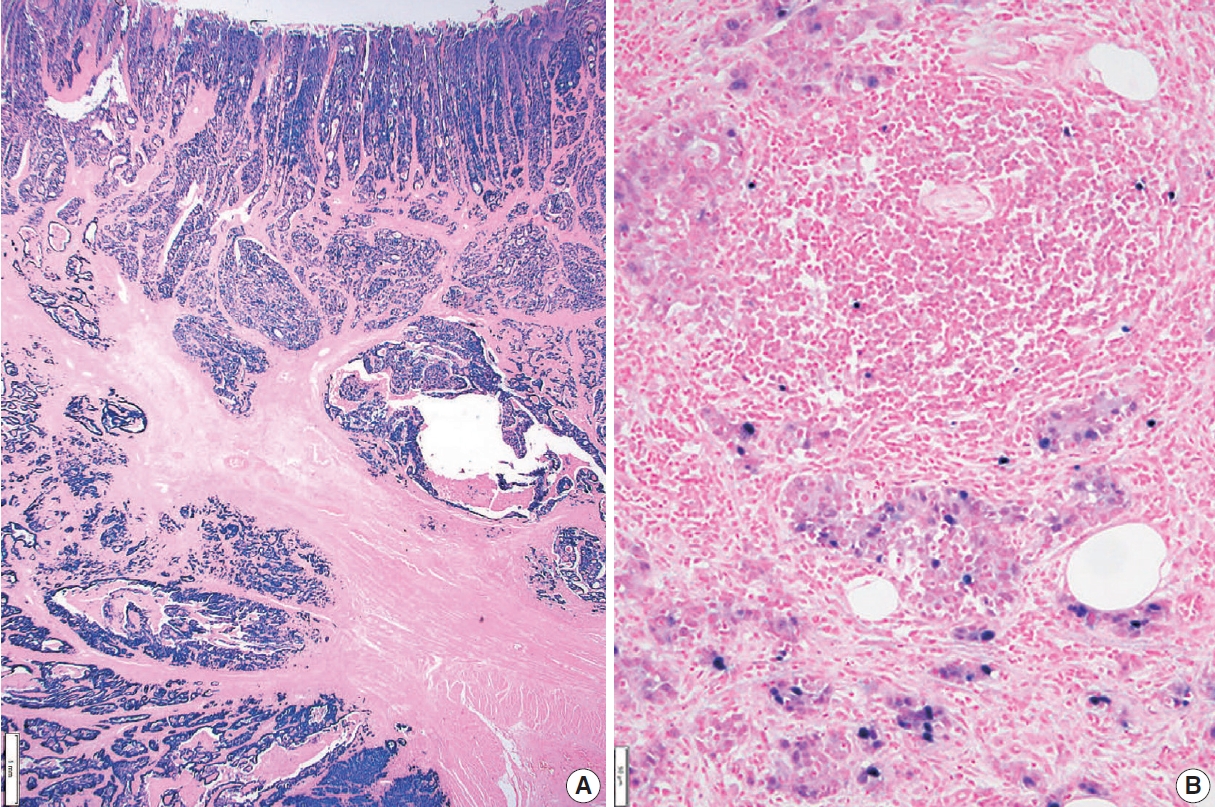
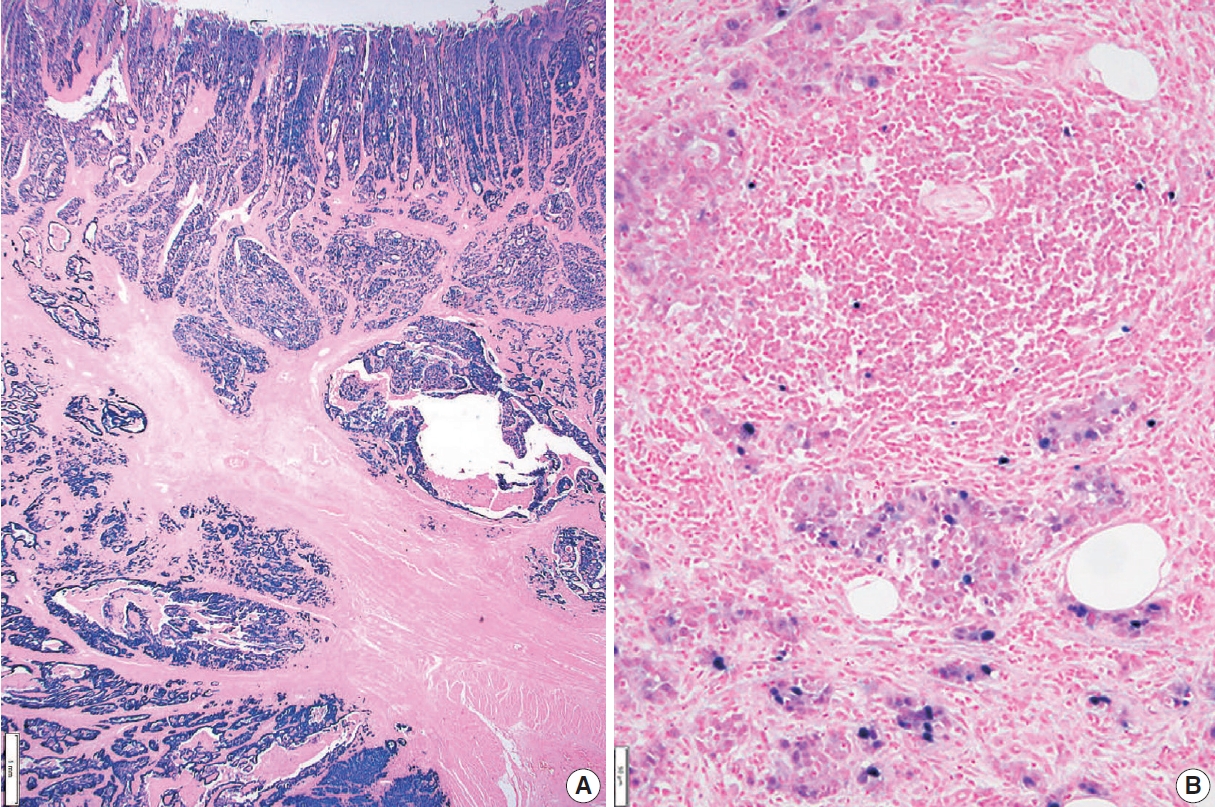
- The presence of lymph node metastasis is one of the most important prognostic factors, even post-chemotherapy [26,27]. Both the total number of evaluated lymph nodes and the number of metastatic lymph nodes are reported. Although pathological evaluation of more than 30 regional lymph nodes is desirable according to the AJCC 8th edition [8], a minimum of 16 regional lymph nodes is acceptable per the CAP guideline [11] because the definition of pN3b is 16 or more metastases. Therefore, if fewer than 16 lymph nodes were initially retrieved for evaluation, additional effort to recover more lymph nodes should be made and reported. This does not apply in cases of previous partial gastrectomy, preoperative chemotherapy, or radiation therapy. Microscopic evaluation should be performed on the largest plane of each lymph node. In general, if the size of the metastasis observed in the lymph node is ≤0.2 mm, the metastasis is called isolated tumor cells (ITCs); if the size is more than 0.2 mm but not greater than 2 mm, it is a micrometastasis. Because micrometastases are not reported separately in GC, they are considered to be positive lymph nodes [8]. According to the AJCC 8th edition, ITCs should not be reflected in the pN stage and should be reported as pN0 (i+) in the absence of another lymph node metastasis. However, it is hard to ignore ITCs, which are readily seen on hematoxylin and eosin (H&E) slides. Therefore, in most practices, all metastatic tumor cell clusters in the lymph nodes are reflected in the pN stage regardless of size, and only ITCs incidentally detected by cytokeratin immunohistochemistry (IHC) are excluded from the pN stage. The stations of the lymph nodes are not reported unless they are separately submitted with corresponding labels. Tumor deposit (TD) is defined as discrete tumor nodules separate from the tumor bed (within the lymphatic drainage of the primary tumor) without identifiable lymph node tissue or vascular or neural structure (Fig. 1) [8]. Unlike colorectal carcinoma, TDs are considered to be metastatic lymph nodes in GC and are thus reflected in the pN stage. TD and serosal (peritoneal) seeding nodules should be distinguished because peritoneal seeding is graded as pM1. Metastasis to a distant lymph node is pM1 and should not be considered in the pN stage. The definition of distant lymph nodes is different in the AJCC 8th edition than in the Japanese guideline, and we recommend following the AJCC 8th edition, in which superior mesenteric lymph node metastasis is pM1 [8].
- If the cancer cells show infiltration of the extranodal adipose tissue beyond the capsule of the lymph node, extranodal tumor extension (ENE) can be reported. ENE is associated with poor prognosis in GC [28-30].
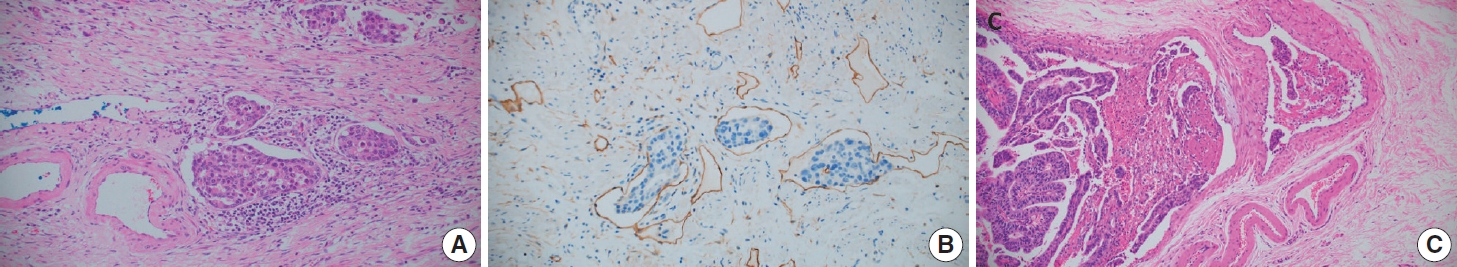
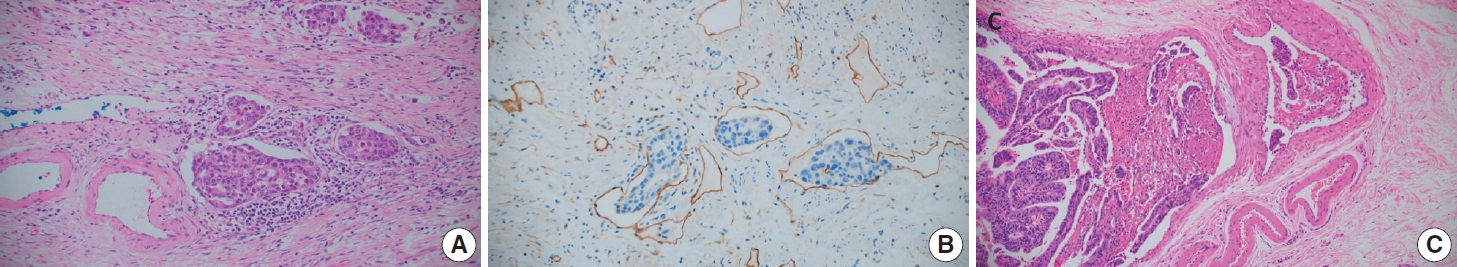
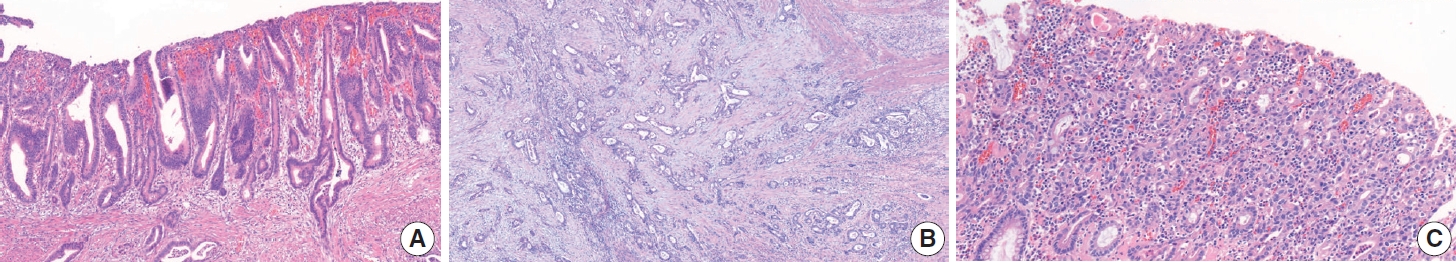
- Lymphovascular invasion includes both lymphatic and vascular invasion. Discrimination of lymphatics from blood vessels on H&E slides is often difficult, especially when they are small (Fig. 2A, B). Although IHC for D2-40 or CD31 can be used, the prognostic differences between lymphatic and blood vessel invasion have not been sufficiently evaluated in GC [12]; therefore, we recommend using ‘lymphovascular invasion.’ However, when tumor invasion or emboli are observed in large vessels with an identifiable smooth muscle layer or elastic lamina, it is called venous invasion and can be reported as a conditional data element (Fig. 2C). Venous invasion has been reported as a risk factor for recurrence in both early [31,32] and advanced GCs [33].
- Perineural invasion is reported when cancer cells are observed within or around the nerve [34].
- Pre-existing adenoma is reported when carcinoma is observed within an adenoma. If the adenoma is discrete from the carcinoma, it is reported as a separate lesion.
- Tumor perforation, serosal (peritoneal, mesenteric) seeding, and distant metastasis (including specific site) are reported when present.
- Peptic ulcers, adenomas, gastrointestinal stromal tumors, and other separate lesions are reported when present.
- Endoscopic resection specimens
- The size of the specimen is expressed as the length of the longest axis and the length perpendicular to the longest axis. The size of the tumor is indicated only by the length of the largest axis. The gross type of the tumor is described in the same way as for a surgical specimen.
- Apply ink to the entire deep margin and lateral margins of the specimen so that it can be viewed under a microscope. Prepare paraffin blocks by sequential parallel sectioning of the entire specimen at 2 mm intervals. Among the lateral margins of the four directions, the closest margin and the tumor should be included together in the sectioning direction.
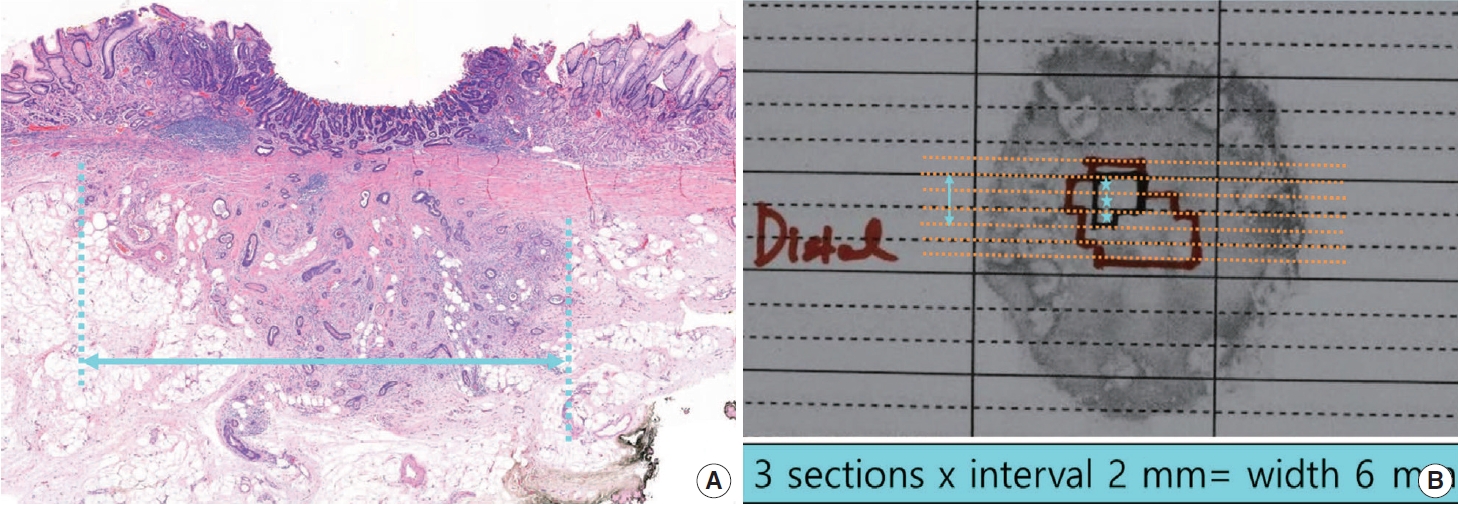
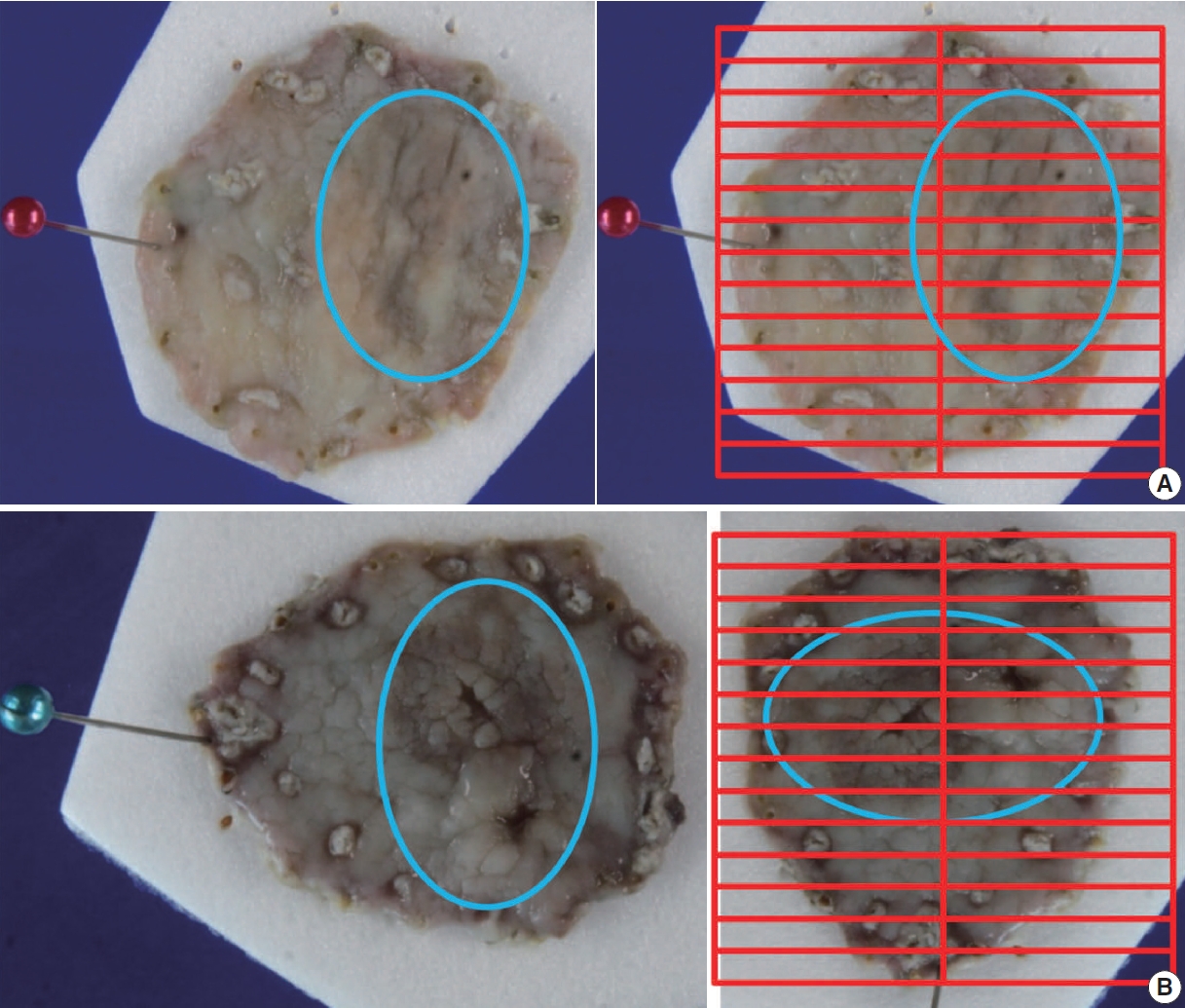
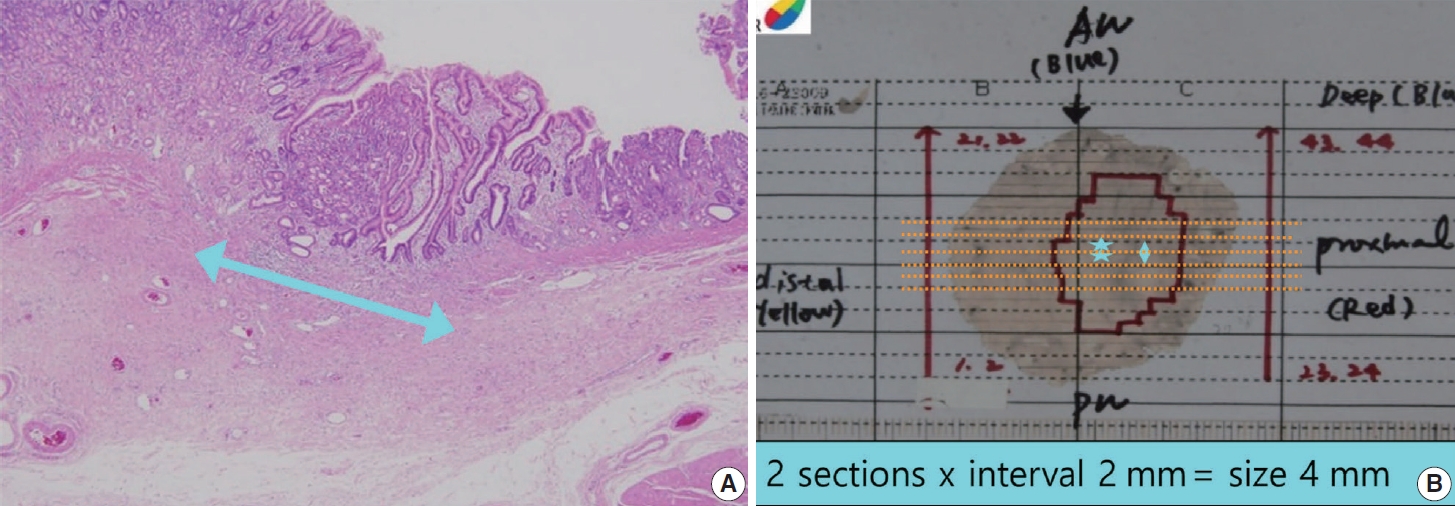
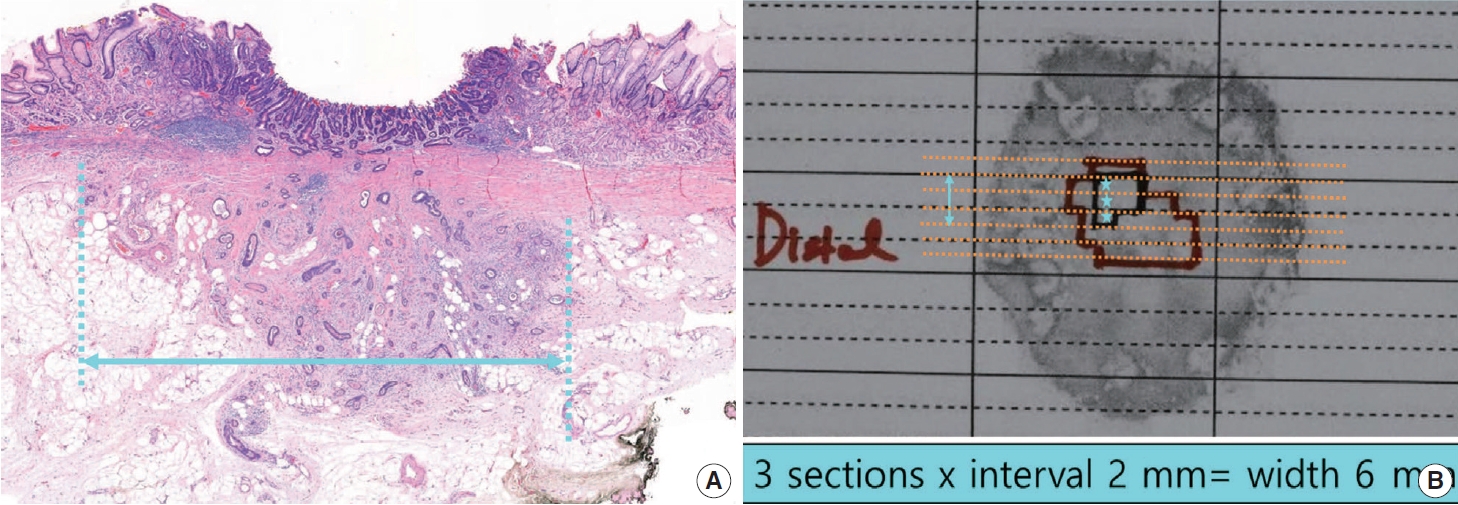
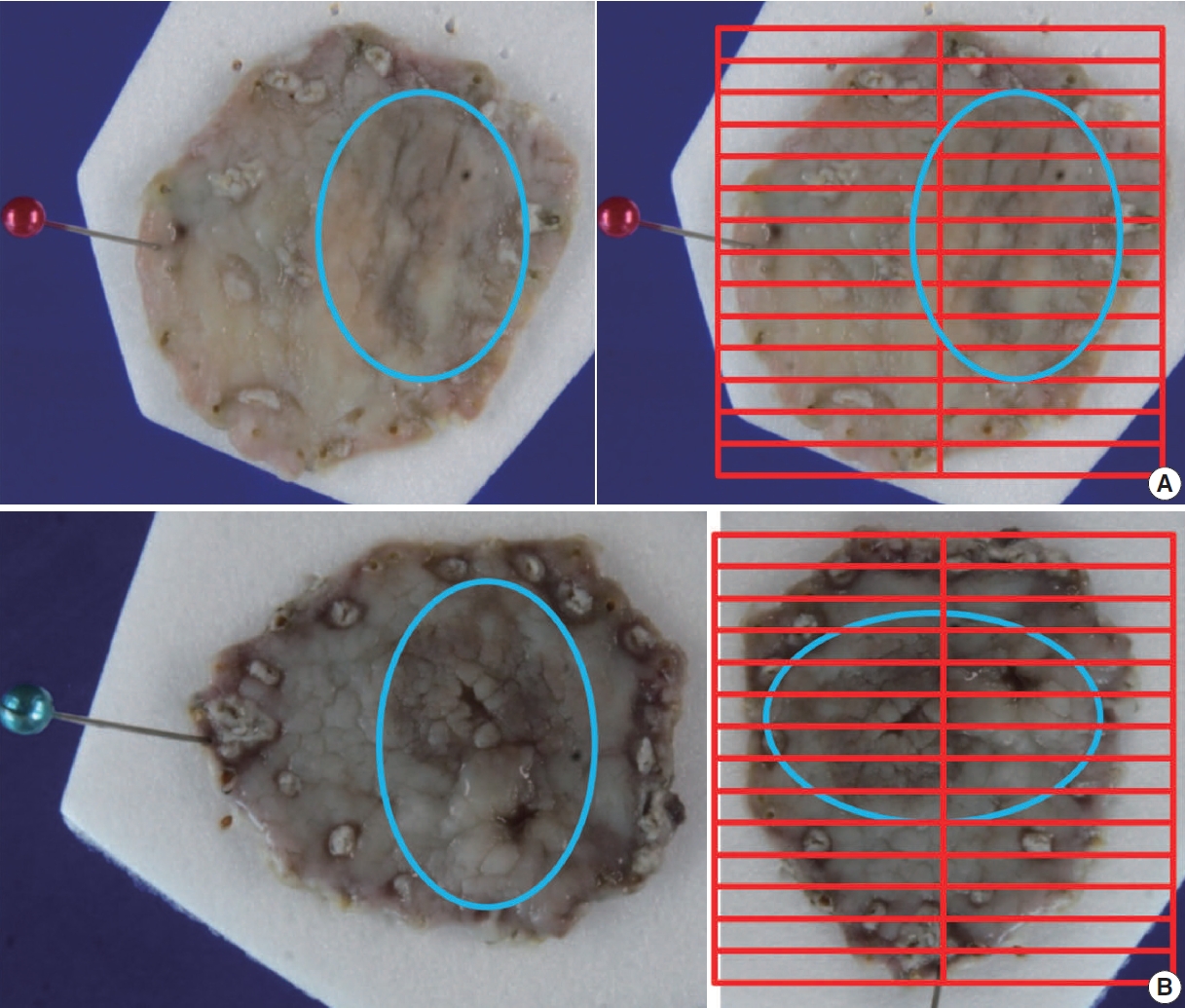
- For gastrointestinal specimens, the distal part is generally placed at the 9 o’clock position in a gross photograph. If the distances from each of the lateral margins are similar, serial sectioning of the specimen is commonly performed in the same direction. When visual observation indicates that the closest lateral margin is not included in this general sectioning direction, however, the direction of the sample or mapping frame should be turned so that the closest lateral margin and the tumor appear together on the section (Fig. 3).
- The histologic type of the tumor is described in the same way as for a surgical specimen. For the criteria and description of each type, refer to the information in the “Histologic classification” section below. The histologic type of the tumor should be described; the histologic diversity of tumor cells may be described separately as histologic components. If various morphologic components are observed within the tumor, all are described according to the histologic type. In such a case, the description should signify the quantitative majority of the tumor components. The description method can be selected according to institutional preferences. For example, record in order: well differentiated (WD)–moderately differentiated (MD)>poorly differentiated (PD)>signet ring cell (SRC) carcinoma; interval variable: WD-MD>50%, PD<50%, SRC<10%; and continuous variable: WD-MD65%, PD 30%, SRC 5%. Many studies have reported that tumors with a mixture of differentiated-type and undifferentiated-type components have a higher risk of lymph node metastasis than tumors with only one component [35-40]. Within the undifferentiated type, SRC has a lower lymph node metastasis frequency, which is reported to be at a level similar to that of the differentiated type [41-43]. In addition, some reports indicate that the lymph node metastasis frequency is lower in pure SRC cases than in SRC cases mixed with other component types [44-47]. However, only the histologic type is applied for determining whether an endoscopic resection is curative, and because differences in histologic components are not applied, they are reflected as conditional elements rather than a standard element. A pathological study of the criteria for determining whether an endoscopic resection is curative is currently underway by the GIPSG of the Korean Society of Pathology as a research project of the National Evidence–based Healthcare Collaborating Agency. If important results are obtained from that study, they should be reflected in the elements of this guideline.
- Only the length of the largest axis of a histologically confirmed tumor is recorded.
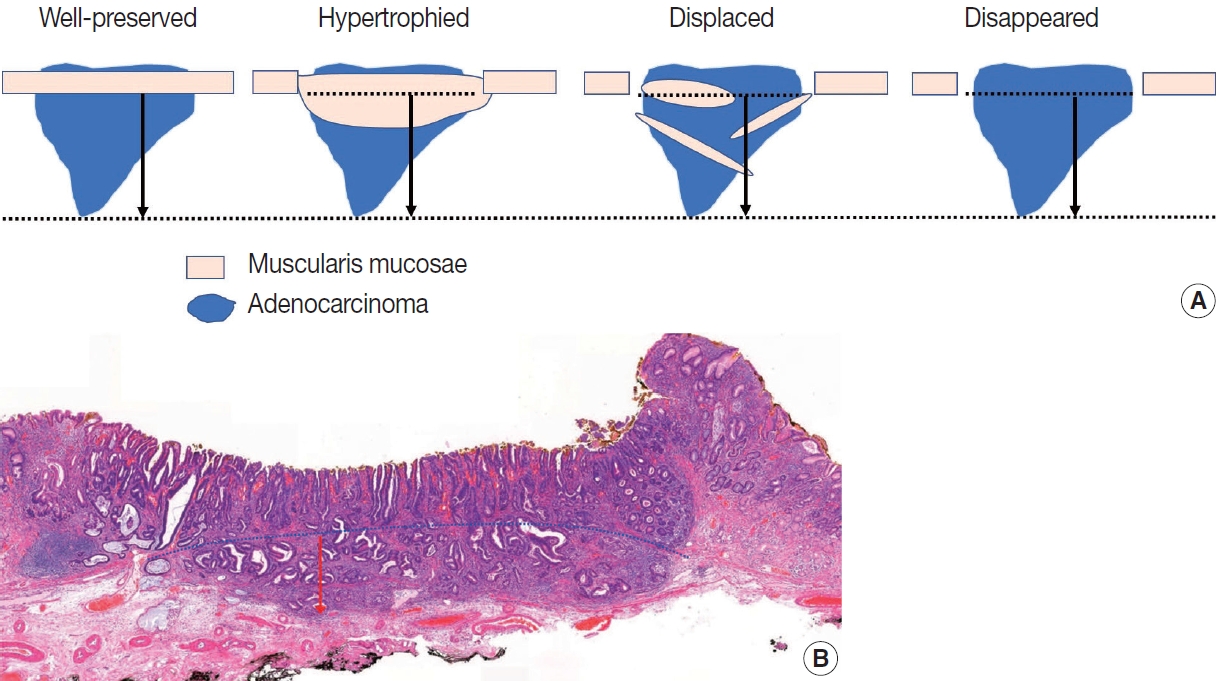
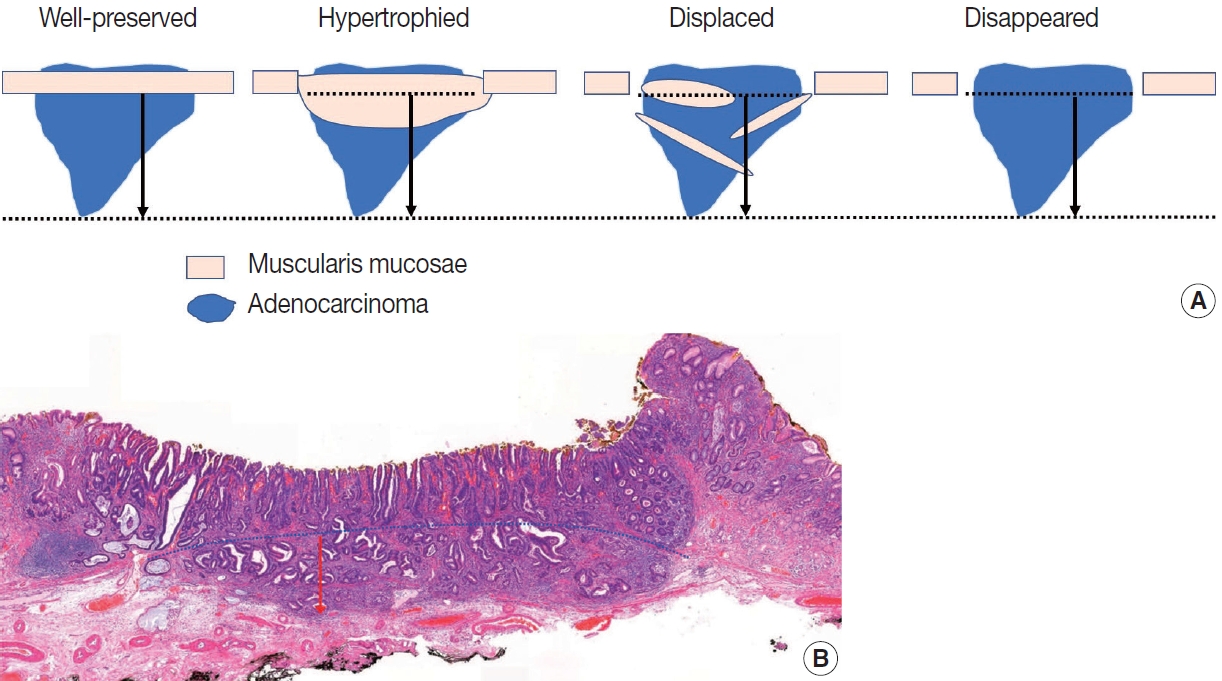
- The method for describing the depth of invasion is basically the same as for a surgical specimen. The difference is that the invasion depth in the submucosal layer is measured and described in cases of submucosal invasion, and it is measured in mm or µm. The measurement is the length from the lowest surface of the muscularis mucosae to the most deeply invaded point. In some cases, the muscularis mucosae are modified by tumor invasion (hypertrophied, displaced, completely disappeared). In these cases, depth is measured using an imaginary line extending from adjacent muscularis mucosae in the normal area not deformed by the tumor (Fig. 4A). Always ensure that the lowest surface of the original, unmodified muscularis mucosae is used as the reference point. If the progressing course of the adjacent muscularis mucosae forms a curve, the virtual line is set as a matching curve (Fig. 4B).
- No definitive description or research results indicate how to measure the depth of invasion when muscularis mucosae are modified. In the Japanese guideline, an explanation first appeared in the 14th edition from 2010: “if the muscularis mucosae are obscure due to ulcerative changes, the length should be measured on the virtual line based on the adjacent normal layer” [9]. In the 15th edition from 2017, it changed to recommend measuring from the surface of the tumor [48]. When muscularis mucosae are modified, some Korean pathologists measure from the lowest muscle fiber of the modified layer, and some measure from the imaginary line of the adjacent normal area. Two Korean studies reported that it is appropriate to measure from the imaginary line of the adjacent normal area in all modified situations [49,50]; accordingly, we use that recommendation as the standard measurement method in this guideline.
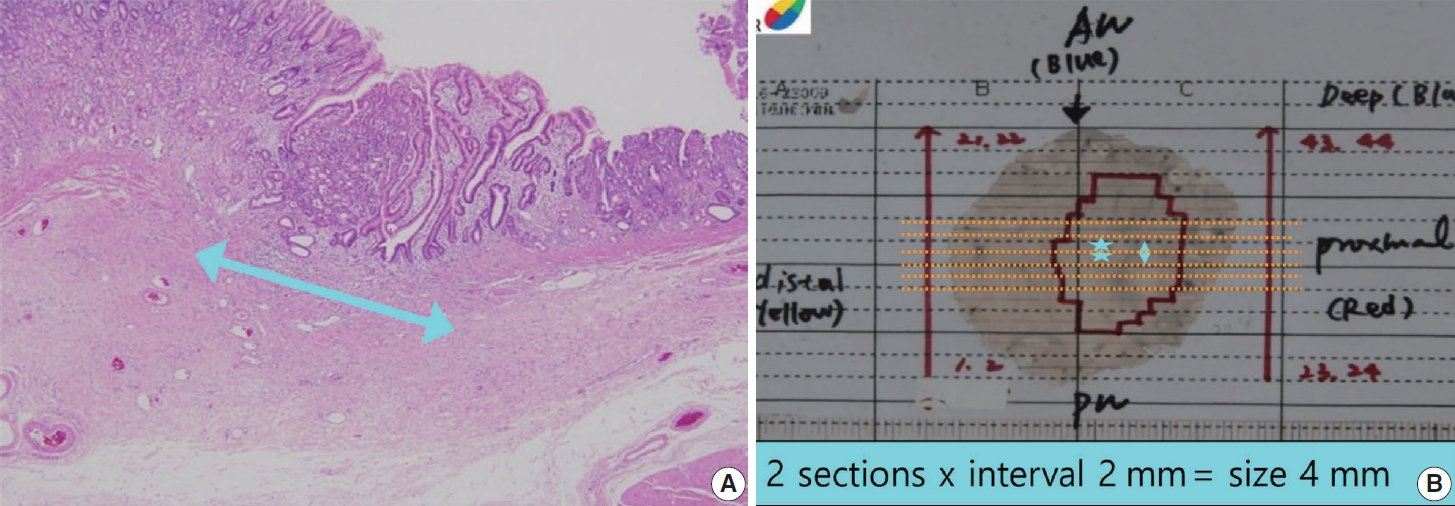
- In cases of submucosal invasion, studies have shown that not only the invasion depth, but also the invasion width are significant risk factors for lymph node metastasis [50,51]. However, because few multicenter studies have been done and it has not yet been applied to the curative resection criteria, the invasion width is a conditional data element. This point is being addressed in the ongoing GIPSG pathological study on the criteria for determining whether an endoscopic resection is curative. The method for measuring the invasion width is as follows (Fig. 5): if submucosal invasion is observed on only one section, write the actual size measured on the slide of that section. If submucosal invasion is observed across two or more slices, write the larger of the following two values: (1) the actual size measured on the slide with the largest invasion width, or (2) the number of slices spanned by the invasion×2 mm (thickness of slice).
- The resection margin is described for the nearest lateral margin and deep margin. If the lateral margin is close (≤0.2 cm) or is involved in the tumor, the corresponding directions should be written together. If multiple margins are involved, all should be written. This is the information needed by the gastroenterologist to decide whether to perform additional procedures (endoscopic resection, argon plasma coagulation, follow-up biopsy). As a conditional element, the distance in all four directions of the lateral margin can be described.
- The degree of invasion of the lateral resection margin and the probability of residual cancer are related. A high risk of residual cancer was reported when two or more of the four lateral margins were involved (multiple involvement) or when the length of involvement was large (more than 4 mm or 6 mm). However, it has not been determined whether additional treatment can be decided according to the degree of margin involvement because the risk is low but present in the group with a small degree of margin involvement.
- Ulceration is defined as a full-thickness disruption of muscularis mucosae, both active and scarring, and determined by histological findings, not endoscopic findings [5,9,52]. The presence or absence of an ulcer is an important criterion for judging whether an endoscopic resection is curative in mucosal cancer [5], so it must be described in the pathology report for mucosal cancer. Because ulcers are included in the indications for an endoscopic resection, the presence of ulcers is determined by endoscopic findings. Ultimately, however, it must be confirmed by pathological examination findings of the resected specimen. Endoscopic diagnosis is difficult in the absence of a mucosal break [53], and ulcer-negative endoscopy findings with ulcer-positive pathology findings were reported in 4.6%–5.5% of cases [54,55].
- Another problem that occurs in practice is a lack of clarity in the criteria for differentiating original small ulcers from biopsy-induced changes after endoscopic biopsy in a case that did not originally have ulcers. Due to the low accuracy of ulcer determination in endoscopic findings, a finding of no ulcer during endoscopy cannot guarantee a biopsy-induced change. Diagnostic criteria for this have been suggested by Shimoda et al. [56], and the Japanese gastric cancer treatment guidelines describe this as follows: “A biopsy-derived scar is usually observed histologically as fibrosis restricted to small areas just beneath the muscularis mucosae. If it cannot be discriminated from the ulcer scar, it should be classified as UL1.” [57]. According to JCOG1009/1010, a clinical study on undifferentiated-type EGC: “UL was judged as present if the muscularis propria was completely disrupted and if fibrosis in the submucosal layer was observed to be wider than the range of disrupted muscularis propria.” [58]. In our study group, ulcer size was measured in the ongoing GIPSG study on the criteria for curative resection, and the possibility of offering differentiation criteria for this problem was investigated. We found that the risk of lymph node metastasis with an ulcer of 4 mm or less was the same as in cases with no ulcer. Using that criterion, very small ulcers can be excluded from the risk factors for lymph node metastasis, which removes the need to differentiate them from biopsy-induced changes. The grading of ulcer size is reflected as a conditional element. The method for measuring the size of an ulcer (Fig. 6) is similar to that used to measure the submucosal invasion width. If an ulcer (full-thickness disruption of the muscularis mucosae) is observed on only one section, write the actual size measured on the slide. If it is observed across two or more slices, write the larger of the following two values: (1) the actual size measured on the slide with the largest disruption size or (2) the number of slices spanned by the disruption×2 mm (thickness of slice). The ulcer size is measured only within the tumor. If the ulcer spans the tumor and surrounding mucosa, measure the ulcer size only within the tumor area.
- The adenoma component should be described only when the histological findings of adenoma are clear, and the intratumoral region is distinct from the adenocarcinoma component.
- In diagnosis, only the adenocarcinoma contents should be described, and adenomas should be described separately as an additional item. For the size of the tumor, the size of the adenocarcinoma is described first, followed by the size of the total tumor. The distance from the resection margin describes the closest distance to any tumor component. If the resection margin is involved in a tumor or is less than 0.2 cm, the component should be described.
- Unlike colorectal cancer, GC occurs in the adenoma-adenocarcinoma pathway in only a small number of cases, and adenocarcinomas of very small size are common. In addition, in many cases of WD adenocarcinoma, structural abnormalities are not severe, so areas that are difficult to differentiate from adenoma can be mixed in the tumor. Therefore, a background adenoma is identified only when the histological findings are clear and the area within the tumor is distinct from the adenocarcinoma component. If it is difficult to distinguish the mixed components, the entire lesion is treated as an adenocarcinoma. For example, if one component corresponds to adenocarcinoma and another component is severely dysplastic but difficult to determine as adenocarcinoma, the whole is treated as an adenocarcinoma component. For an adenoma, only the presence of the adenoma component is briefly described in a separate section.
- Piecemeal resection or full-thickness tearing should be confirmed and documented in the histological examination. Even if the specimen is resected into several pieces, it is not piecemeal if the tumor is intact within one piece.
- Unlike surgical specimens, lymphatic and venous invasions are recorded separately in endoscopic resection specimens because of the differing risks of lymph node metastasis. Both lymphatic invasion and venous invasion are criteria for determining a non-curative resection. However, the risk of lymph node metastasis posed by lymphatic invasion is times higher than that from venous invasion, and a higher score is assigned in the risk prediction model [59]. This information is helpful when clinicians decide whether or not to perform gastrectomy; thus, it is recommended to report them separately. The standard method for differentiating lymphatic and venous invasion is H&E staining with the following criteria: it is determined as a lymphatic vessel when there is a thin wall or lymphatic fluid and as a venous vessel when there is a thick muscle wall or many red blood cells in the lumen. When it is difficult to distinguish between lymphatic vessels and small venules, classify them as lymphatic vessels.
- IHC staining may be performed to better observe lymphatic or venous vessels. However, because H&E and other immunostained slides are obtained from different levels, they should be interpreted separately. A specimen is deemed to be positive even if invasion is observed on only one slide.
- Histologic classification
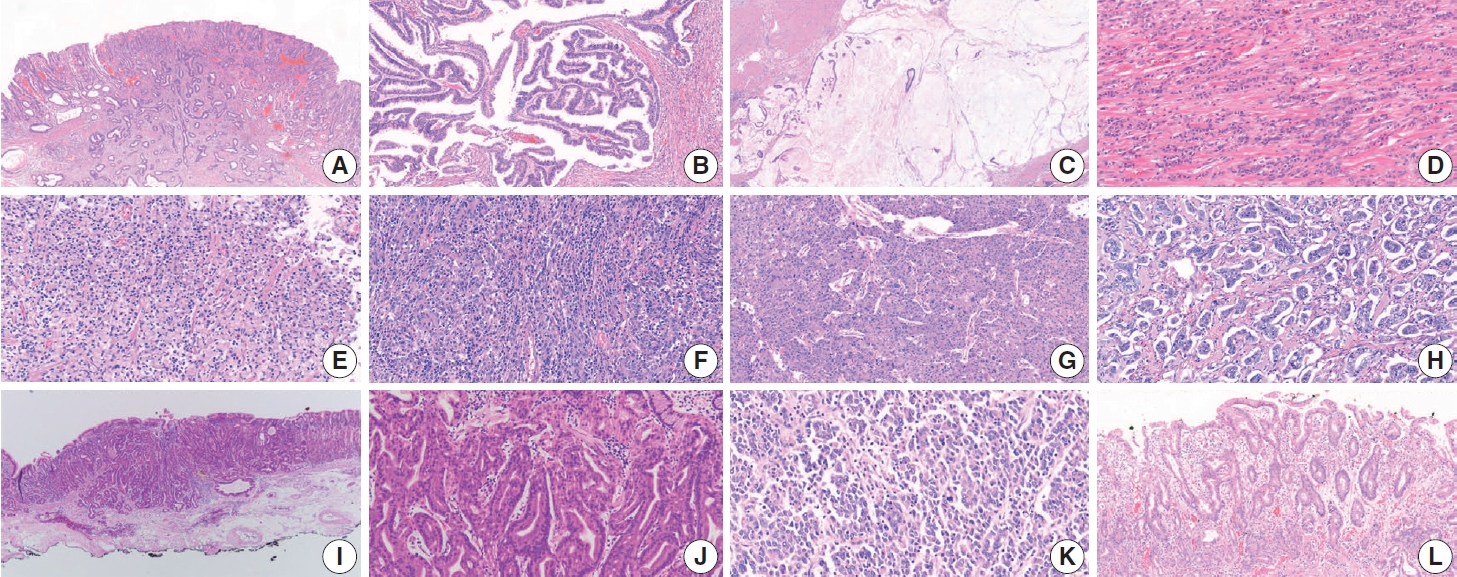
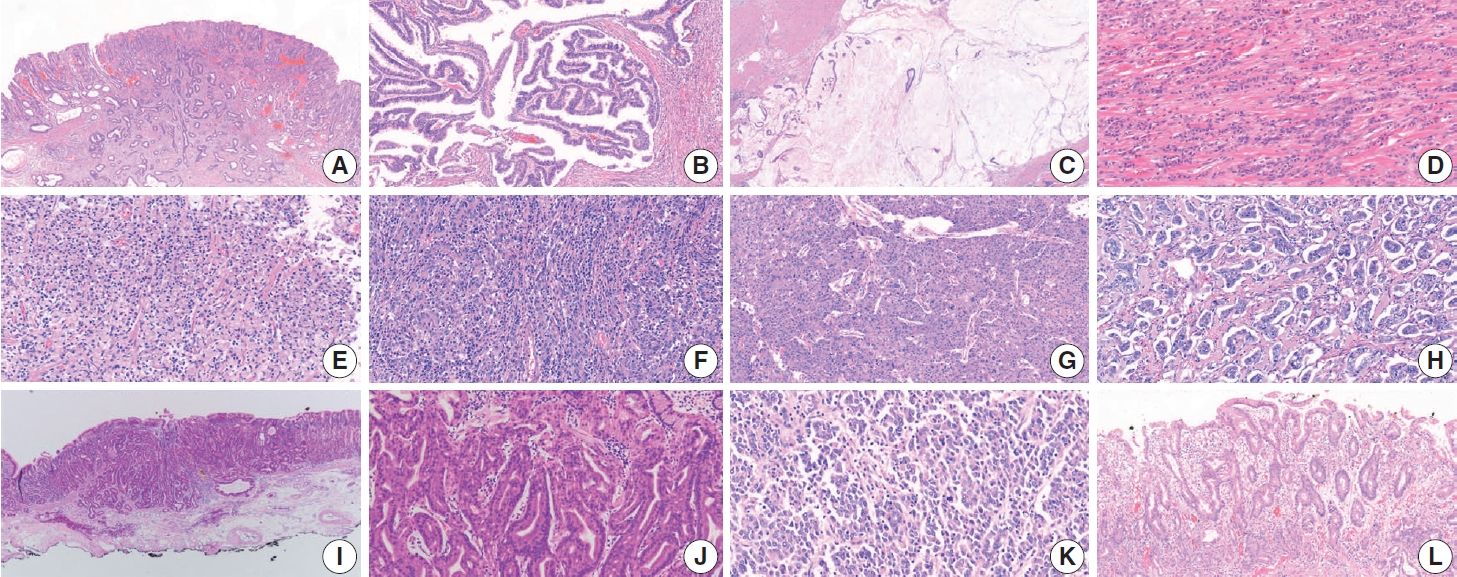
- Histologic classification of GC is based on the 5th edition of the World Health Organization (WHO) blue book [4]. Representative histopathologic types described in the WHO classification are summarized in Table 3 and Fig. 7. The diagnosis of GC is usually determined according to the component that occupies the largest portion of the tumor, but the diagnosis of special histologic subtypes is based on the diagnostic criteria of each subtype. The most common subtype is tubular adenocarcinoma, characterized by prominent dilated or slit-like tubules. Carcinomas composed of solid tumor clusters with rare tubule formation are also classified as tubular adenocarcinoma. Tumor cells can be columnar, cuboidal, or flat, and luminal mucin/cell debris is common.
- Papillary adenocarcinoma shows a papillary tumor structure with a central fibrovascular core and columnar or cuboidal tumor cells. For a diagnosis of papillary adenocarcinoma, more than 50% of the tumor area must contain the papillary tumor component [60-62]. High rates of liver metastasis, lymphovascular invasion, lymph node metastasis, and poor prognosis are reported in papillary adenocarcinoma [61-64].
- Mucinous adenocarcinoma is defined when more than 50% of the tumor area shows extracellular mucin. Tumor cells in mucinous adenocarcinoma can show a glandular growth pattern, solid pattern, or scattered single cell pattern, including SRC carcinoma [4]. Mucinous adenocarcinoma is classified as the intestinal, diffuse, or indeterminate type according to the main component of tumor cell differentiation [4]. Mucinous adenocarcinoma tends to be diagnosed at an advanced stage [65,66].
- Poorly cohesive carcinoma (PCC) is the second most common subtype of GC and is composed of isolated or small groups of tumor cells without gland formation [4]. Until the 3rd edition of the WHO classification, SRC carcinoma was an independent subtype, but since the 4th edition of WHO classification, SRC has been included in the PCC category. Recently, several studies have suggested that non-SRC PCC (PCC-NOS) has a relatively poor prognosis compared with SRC and that SRC and PCC-NOS have different molecular profiles [67-70]. The WHO classification defines SRC as “composed predominantly or exclusively of signetring cell components” [4]. A European group suggested a PCC classification definition according to the percentage of the SRC component (SRC, >90%; PCC-NOS, <10%; PCC with SRC component, 10%–90%), but no definite criteria for diagnosing PCC-NOS and SRC have been established, so more studies are required [71].
- Mixed adenocarcinomas, according to the WHO definition, are carcinomas with both glandular (tubular adenocarcinoma/papillary adenocarcinoma) and poorly cohesive (PCC/SRC) components [4]. Some reports recently suggested that mixed adenocarcinomas have poorer prognosis, such as frequent local recurrence and lymph node metastasis, than a pure subtype of carcinoma, especially in EGC [72,73]. However, no clear criteria have established a minimum ratio of glandular/poorly cohesive components for a diagnosis of mixed adenocarcinoma. Contrary to the WHO definition, many studies define mixed adenocarcinoma as PD adenocarcinoma or a PCC/SRC component mixed with glandforming components; those studies also report that the prognosis of mixed adenocarcinoma in EGC is worse than that of pure subtypes [39,74,75]. Although mixed adenocarcinoma does not have a clear definition, it seems that EGC has a poor prognosis when a glandular component coexists with other components in the same tumor; therefore, when both a glandular component and other components are observed in an EGC, it is recommended that they be mentioned separately.
- Adenocarcinoma with lymphoid stroma (medullary carcinoma with lymphoid stroma) was previously called ‘lymphoepithelioma-like carcinoma’ or ‘medullary carcinoma.’ Tumor cells of this subtype show irregular sheets, poorly defined clusters or tubules, trabeculae, or syncytial cells with dense lymphocytic infiltration and intraepithelial lymphocytes [4,76]. Such a tumor usually shows a well-defined margin without infiltrative growth and minimal desmoplasia. This type of tumor is frequently associated with Epstein-Barr virus (EBV) infection and sometimes shows microsatellite instability/mismatch repair deficiency [4,76]. Patients with this subtype show a lower number of lymph node metastases and better prognosis after surgery than those with other subtypes [77,78].
- Hepatoid adenocarcinoma is composed of hepatocyte-like tumor cells, which are large polygonal cells with eosinophilic-abundant cytoplasm arranged in a trabecular pattern [4,79]. This alpha-fetoprotein-positive tumor is often diagnosed preoperatively with multiple liver and lymph node metastases [4,79].
- Micropapillary adenocarcinoma is characterized by an insideout pattern of tumor clusters, which are small tumor clusters without a fibrovascular core, in clear spaces [4,80]. Micropapillary adenocarcinoma can be diagnosed when more than 10% of the tumor comprises micropapillary components [4,81]. This subtype is associated with poor prognosis and lymph node metastasis [4,80,81].
- Adenocarcinoma of the fundic-gland type is composed of tumor cells showing chief cell differentiation, parietal cell differentiation, or both. Because this tumor does not show obvious nuclear dysplasia or structural abnormalities, it would be reasonable to regard it as adenocarcinoma only when it invades the submucosal layer. Lymph node metastasis is very rare in this subtype [4,82,83].
- Undifferentiated carcinoma is composed of anaplastic cells without specific differentiation [4]. Grossly, a large ulcerating or fungating mass with necrosis is common. Tumor giant cells and rhabdoid tumor cells are common in this subtype, and spindle sarcomatoid cells can be seen [84,85]. Most patients show dismal prognosis with distant metastasis.
- Squamous cell carcinoma is a very rare gastric tumor and shows morphology similar to that found in other organs. Adenosquamous carcinoma has both glandular and squamous tumor components, with ≥25% squamous component [4]. Gastroblastoma is a biphasic tumor composed of spindle and epithelial cells.
- Crawling-type adenocarcinomas are characterized by complex branching or anastomotic structures and low-grade nuclei and have not yet been classified as a distinct subtype in the WHO classification [4]. Because of their low-grade nuclear atypia, reactive looking structural change, and mucosal location, crawling-type adenocarcinomas were once called a very WD form of gastric adenocarcinoma. Recent studies have shown that large crawling-type adenocarcinomas are often accompanied by PD components, and one report indicates that lymph node metastasis occurs frequently when the cancer invades beyond the submucosal layer [86,87]. Although it has not yet been classified as a formal subtype, some research results on crawling-type adenocarcinoma have recently been published, and attention needs to be paid in terms of prognosis.
- Tubular adenocarcinoma and papillary adenocarcinoma can be graded. When two or more differentiations are mixed in an adenocarcinoma, the differentiation grade reflects the largest tumor area. A distinct glandular structure composed of columnar cells is classified as WD, and a small glandular structure composed of cuboidal or flat cells is classified as MD. In a tumor with an indistinct glandular structure, carcinoma forming frequent luminal structures is classified as MD, and that with a rare luminal structure is classified as PD (Fig. 8) [3]. Although the WHO recommends a two-tier grading system of low- (WD and MD) and high-grade (PD), most pathologists and clinicians use a three-tier grading system. We have agreed to use a three-tier grading system that can be easily switched to a two-tier grading system.
- In endoscopic gastric biopsy samples, it is often difficult to diagnose a specific subtype of gastric carcinoma. However, histologic subtypes and differentiation are important in the selection of a treatment modality. We recommend reporting a histologic component or subtype if there is a PD component or subtypes associated with poor prognosis (such as PCC, PD tubular adenocarcinoma, or micropapillary feature), irrespective of the proportion. Some peculiar subtypes of adenocarcinomas, such as adenocarcinoma of the fundic-gland type and EBV-associated gastric carcinoma, have a lower rate of lymph node metastasis than other subtypes with similar invasion depth, especially in EGC [82,88,89]. Reporting these subtypes and testing for EBV in situ in biopsy specimens could thus be helpful for patient management [89].
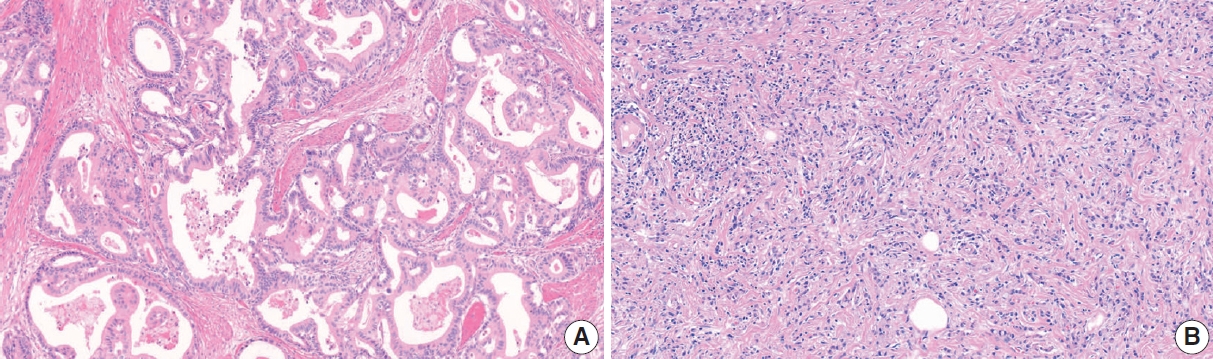
- The Lauren classification has been one of the most commonly used classification systems for GC worldwide since its publication in 1965 (Table 3) [90]. According to the WHO 5th edition, WD and MD papillary adenocarcinoma and tubular adenocarcinoma are classified as the intestinal type, and PCC and SRC are classified as the diffuse type (Fig. 9). In the Lauren classification, the mixed type (not the same as mixed adenocarcinoma in the histological classification) is used when intestinal and diffuse tumor components coexist in similar proportions. Although a table in the WHO blue book indicates that solid type, PD adenocarcinoma is classified as indeterminate type, this does not mean that all PD adenocarcinoma should be classified as such, and there is some disagreement among pathologists about the definition of the indeterminate type. Further discussion is needed to decide whether other special histological types of adenocarcinoma are excluded from the Lauren classification or whether they can be classified as intestinal, diffuse, or indeterminate according to their morphology.
- To determine the feasibility of an endoscopicresection of tumors, most clinical guidelines and studies apply the differentiated type (papillary adenocarcinoma, tubular adenocarcinoma, WD and MD)/undifferentiated type (tubular adenocarcinoma, PD and poorly cohesive carcinoma, including SRC carcinoma) criteria of the Japanese guidelines [57]. In these criteria, PD adenocarcinoma is classified as the undifferentiated type. To prevent confusion with undifferentiated carcinoma, we do not recommend using the ‘differentiated type/undifferentiated type’ criteria in pathology reports. Instead, using the histologic classification and/or Lauren classification can provide sufficient information to clinicians and researchers.
- Neoplastic epithelial proliferation without stromal invasion is called either adenoma or dysplasia. This intraepithelial neoplasia is usually called an adenoma by Western pathologists when the tumor shows a protruding, polypoid appearance with a distinct border and dysplasia when the tumor appears as a flat, depressed lesion or elevated indistinct lesion [4]. The Japanese classification tends to refer to elevated, flat, and depressed intraepithelial lesions as adenomas. Both adenoma and dysplasia can be used as terms for intraepithelial neoplasia in Korea.
- Gastric adenomas can be subclassified into the intestinal type, foveolar type, pyloric gland type, and oxyntic gland type. Intestinal-type adenomas are the most common adenomas and usually show tubule formation and columnar cells with elongated nuclei, with or without goblet cells and Paneth cells [4]. Foveolar-type adenomas are the second most common type of gastric adenoma, and an apical mucin cap is characteristic [91]. Pyloric gland type adenomas consist of columnar cells with ground-glasslike cytoplasm, basally located nuclei, and closely packed tubular glands with occasional dilatation [92]. Oxyntic gland type adenomas, also called oxyntic gland neoplasms because they can be diagnosed as adenocarcinoma only when submucosal invasion is confirmed, can progress into adenocarcinoma of the fundic gland type. This adenoma is composed of tumor cells with an oxyntic gland (chief cells, parietal cells, and mucous neck cells) and exhibits structural irregularity and minimal to mild nuclear atypia [82,88].
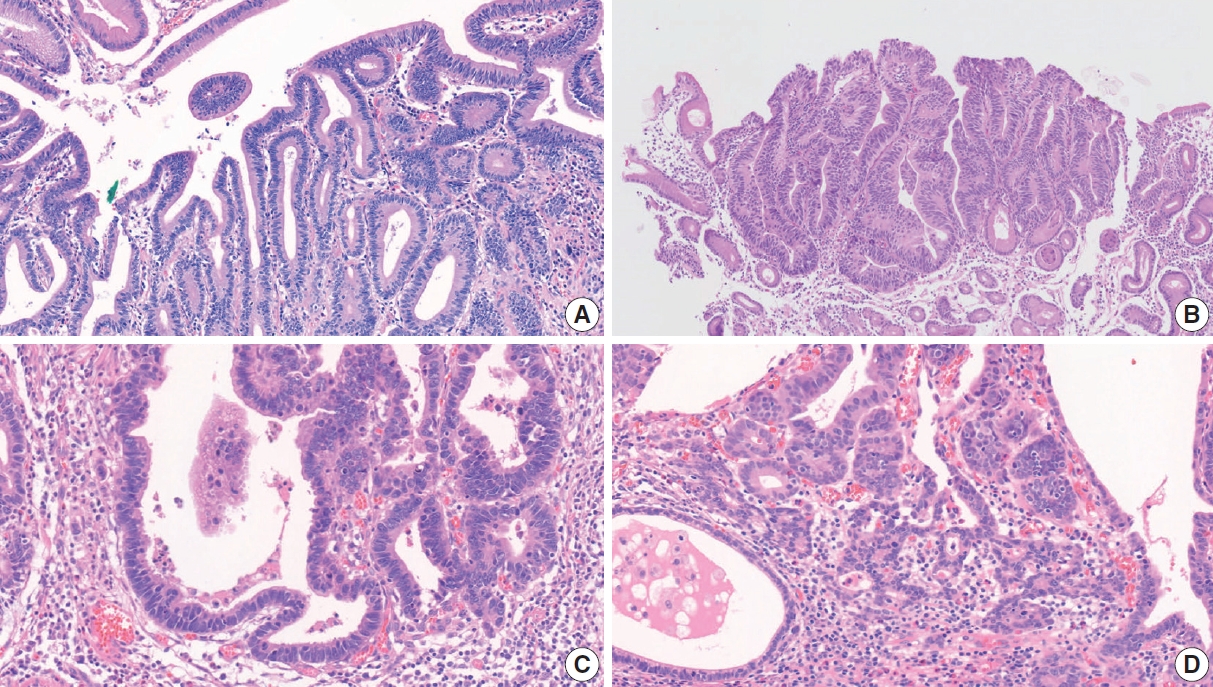
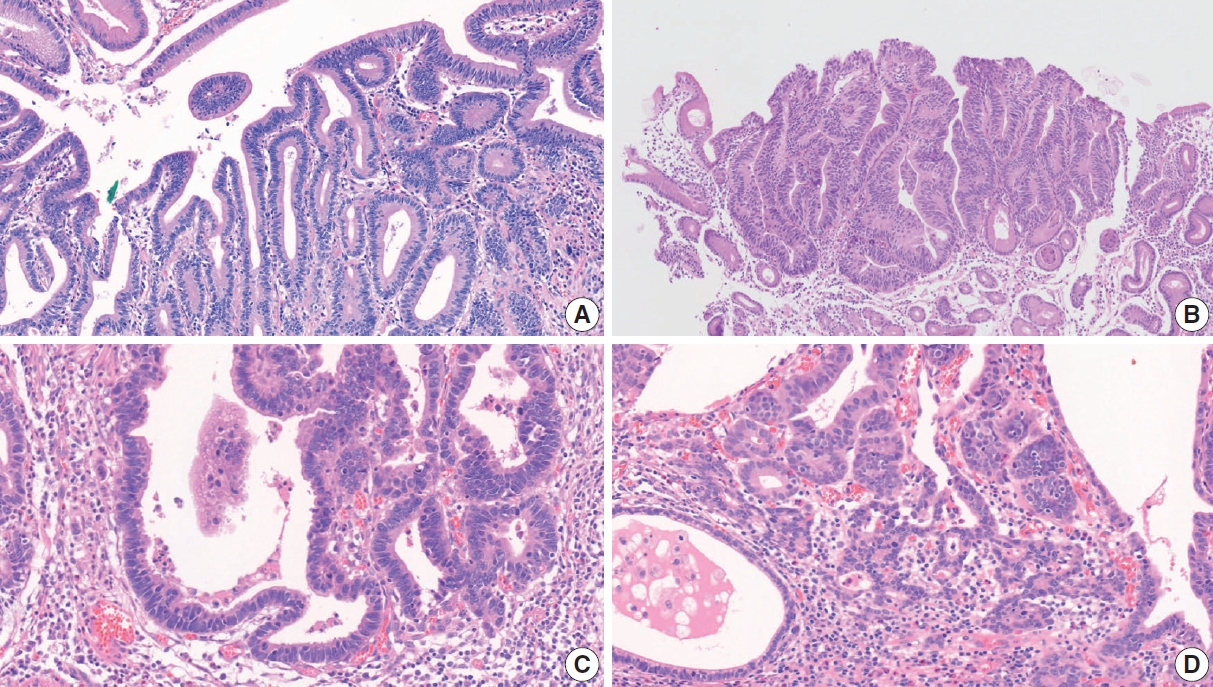
- A two-tier system (low-grade/high-grade) is recommended for grading adenomas. Low-grade adenomas are characterized by a simple tubular or papillary architecture, hyperchromatic elongated or ovoid nuclei without severe atypia, preserved cellular polarity with basally located nuclei, and relatively regular intervening stroma without structural disruption. Goblet cells, apoptotic features, and mild to moderate mitotic features can be observed in low-grade adenomas (Fig. 10A). High-grade adenomas show more complex structures such as fusion, crowding, and budding of glands and the formation of glands with varying diameters. Cellular atypia is more pronounced in high-grade adenomas, such as loss of polarity, a high nuclear/cytoplasm ratio, pleomorphic nuclei, frequent mitosis, and atypical mitosis [93,94]. Intraglandular necrotic debris is also a diagnostic clue for high-grade dysplasia and, more commonly, adenocarcinoma (Fig. 10B) [95]. A diagnosis of adenocarcinoma should be considered when more than one of the following is present: evidence of stromal invasion (including single cell invasion into stroma and desmoplastic reaction), marked structural atypia, and marked glandular crowding (Fig. 10C) [94].
- H. pylori infection is the most common cause of gastric adenocarcinoma, and eradication of H. pylori is associated with metachronous GC [96,97]. To detect H. pylori infection in a pathology specimen, additional staining (such as the Wright-Giemsa stain or Warthin-starry stain) is recommended. The proportion of drug-resistant H. pylori is increasing, and in patients with clarithromycin-resistant H. pylori infection, the failure rate of standard eradication treatment is also increasing. In patients with H. pylori infection, testing for clarithromycin-resistance is helpful for H. pylori eradication.
- Molecular markers
- All molecular tests are optional, conditional data elements. All report forms for the pathologic diagnosis of molecular markers are shown in Table 4.
- Determination of human epidermal growth factor receptor 2 (HER2) status is critical to identify patients with advanced-stage cancer for appropriate precision therapy. HER2-positive GC patients are currently treated with trastuzumab in combination with chemotherapy as first-line therapy, and fam-trastuzumab deruxtecan-nxki, a.k.a. trastuzumab deruxtecan, was recently approved by the Food and Drug Administration as a third- or later-line treatment [5,7,98,99]. HER2 status is principally determined by IHC or in situ hybridization (ISH) assays. HER2-positivity is defined as IHC 3+ or IHC 2+/ISH-positive [100,101]. HER2 testing requires formalin-fixed paraffin-embedded biopsy tissues with an adequate number of tumor fragments (ideally at least four) or representative surgical specimens with more differentiated components [102,103].
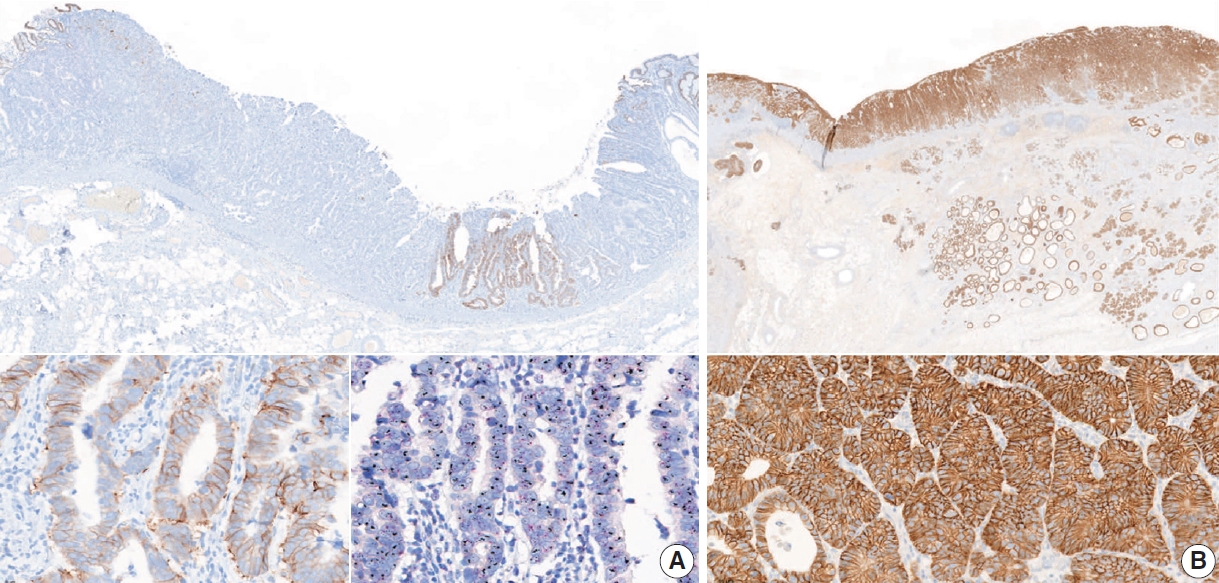
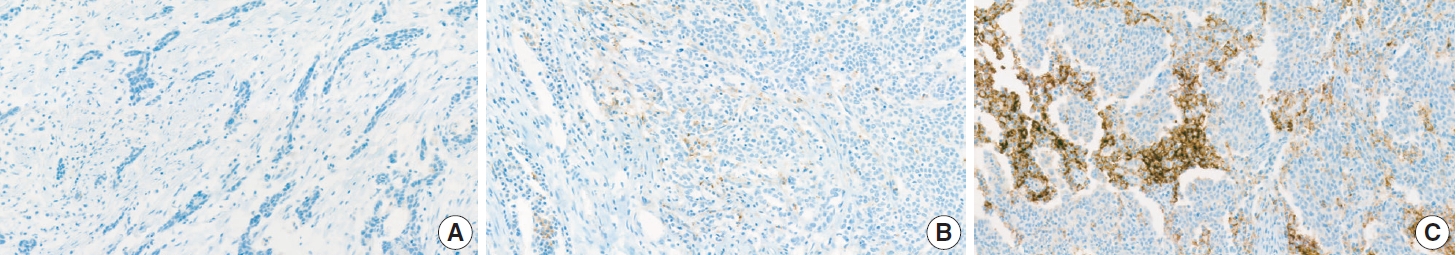
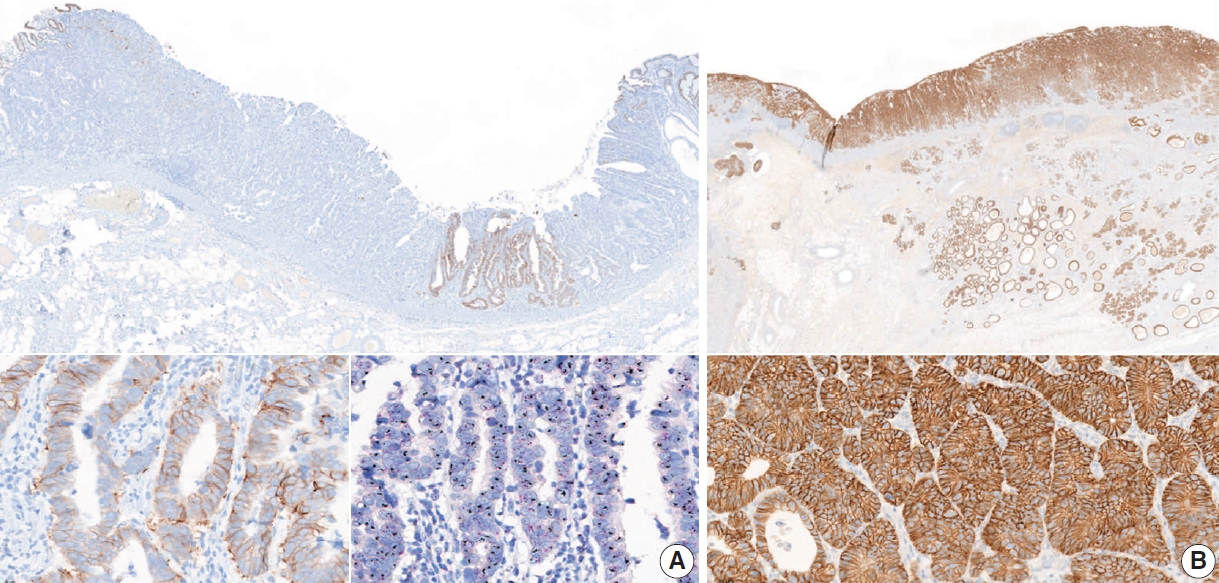
- In currently recommended testing algorithms, HER2 status should be initially established using IHC [7,100] to estimate the immunoreactive intensity and percentage of basolateral membranous expression on cancer cells [7,104]. The score ranges from 0 to 3 based on ≥10% cutoff level of HER2 expression in surgical specimens and ≥5 clustered cells in biopsy specimens as follows: 0 (negative), no reactivity or membranous reactivity in <10% of cancer cells from surgical specimens or any cancer cells in biopsy specimens; 1+ (negative), faint or barely perceptible membrane reactivity; 2+ (equivocal), weak to moderate complete or basolateral membrane reactivity; and 3+ (positive), strong complete or basolateral membrane reactivity (Fig. 11).
- Cases with a score of 2+ or indeterminate by IHC should be confirmed with ISH techniques to determine the final HER2 status [7,100]. Positive HER2 amplification is defined as a HER2: CEP17 (centromeric region of chromosome 17) ratio ≥ 2.0. To evaluate the ISH results, first check the HER2 IHC slide to select the most strongly stained region that might predict a higher level of HER2 amplification. Next, at least 20 evaluable, non-overlapping invasive tumor cells should be counted. If CEP17 signals are ≥3 and the ratio of HER2:CEP17 is <2.0, an average HER2 copy number >6 signals/cell is considered positive for HER2 amplification by ISH and <4 signals/cell is considered negative. If an average HER2 copy number is between four and six signals/cell, another 20 cells should be counted in a different area. Sometimes, the determination of HER2 status is uncertain due to sample problems or technical issues [103,105]. In that case, the test should be reported as “cannot be determined.”
- Some studies have revealed a significant correlation between HER2 expression and histologic subtype in GC. The Trastuzumab for Gastric Cancer (ToGA) trial and other published studies showed that the HER2 positivity rate was higher in differentiated subtypes (Lauren intestinal type and WD and MD type) than in the Lauren diffuse type or PD type [106-108]. Furthermore, intratumor heterogeneity of HER2 expression was reported in approximately 50% of GC cases [106,109]. Inter-lesional heterogeneity of HER2 expression for either positive or negative shifting has been reported between primary carcinomas and synchronous or metachronous locoregional/distant metastases at a rate of 2%–14% [110-115].
- Therefore, HER2 status should be re-evaluated for all newly diagnosed secondary, recurrent, and metastatic lesions, regardless of the HER2 status of the primary cancer because it affects the therapeutic strategy and prognosis of patients [116,117].
- Microsatellites, also called short tandem repeats, consist of repeats of a sequence that ranges from 1–6 nucleotides in length [103,118,119]. DNA mismatch repair (MMR) is a highly conserved mechanism to recognize and replace or repair mismatched nucleotides during DNA replication [119]. MMR deficiency (dMMR) is commonly caused by a germline mutation or sporadic epigenetic silencing and leads to insertions or deletions of nucleotides in microsatellite regions during DNA replication [119,120]. The four genes that play an important role in this process are mutL homolog 1 (MLH1), mutS homolog 2 (MSH2), mutS homolog 6 (MSH6),and PMS1 homolog 2 (PMS2) [103,119-121]. When MMR does not function normally, it is called microsatellite instability (MSI) [119,122].
- MSI is the hallmark of Lynch syndrome and is found in many sporadic cancers [103,123]. MSI-high (MSI-H) is observed in 6.9%–22.7% of sporadic GC cases [124-127]. As a distinct molecular subtype, MSI-GC is characterized by the gastric CpG island methylator phenotype with MLH1 silencing [124]. The clinical characteristics of MSI-GC are antrum (distal) locations, intestinal type of Lauren histology, early disease stage, and favorable prognosis [5,103,125,126]. Clinically, MSI is an actionable predictive biomarker for resistance to 5-fluorouracil-based adjuvant chemotherapy and indicates good suitability for immunotherapy [128-132]. For this reason, clinician requests for MSI and/or MMR test are increasing. In the National Comprehensive Cancer Network Guidelines for Gastric Cancer V.2.2022, universal MSI and MMR testing is recommended for all newly diagnosed GC patients, in accordance with the CAP DNA Mismatch Repair Biomarker Reporting Guidelines [100].
- The three main methods used to detect MSI/dMMR are as follows: (1) polymerase chain reaction (PCR) amplification of microsatellite sequences; (2) IHC staining to determine the expression of the four MMR proteins MLH1, MSH2, MSH6, and PMS2; and (3) next-generation sequencing (NGS) [103,119,120,133]. Additionally, a new kit enables diagnosis of MSI according to the number of deleted base mutations by using a melting curve analysis with a peptide nucleic acid (PNA) probe [134].
- PCR can compare the allelic position of the microsatellite locus in the tumor with that in normal tissue [103,120,133]. The National Cancer Institute recommends the so-called Bethesda Panel as reference [133,135]. This panel is composed of two mononucleotide repeats (BAT-25 and BAT-26) and three dinucleotide repeats (D2S123, D5S346, and D17S250) [22,103,133,135]. These regions are amplified in parallel using fluorescent PCR, and their sizes are assessed by capillary electrophoresis [133,136]. However, because the dinucleotide markers are less sensitive and specific than the mononucleotide markers [137], an alternative panel with five poly-A mononucleotide repeats (NR-21, NR-24, NR-27 [or Mono-27], BAT-25, and BAT-26) has also been suggested [22,103,119].
- MSI-H is defined as instability of two or more of five microsatellite loci; MSI-low (MSI-L) is defined as instability of one site, and microsatellite stable (MSS) is defined as no instability at any site. Currently, clinical studies tend to categorize MSI-L and MSS as one type. This PCR method enables a functional measure of dMMR by directly measuring DNA changes. However, the method does not identify the MMR gene to be investigated. When the PCR test fails or the interpretation of the results is difficult, the test should be reported as “undetermined,” and IHC testing or NGS is recommended.
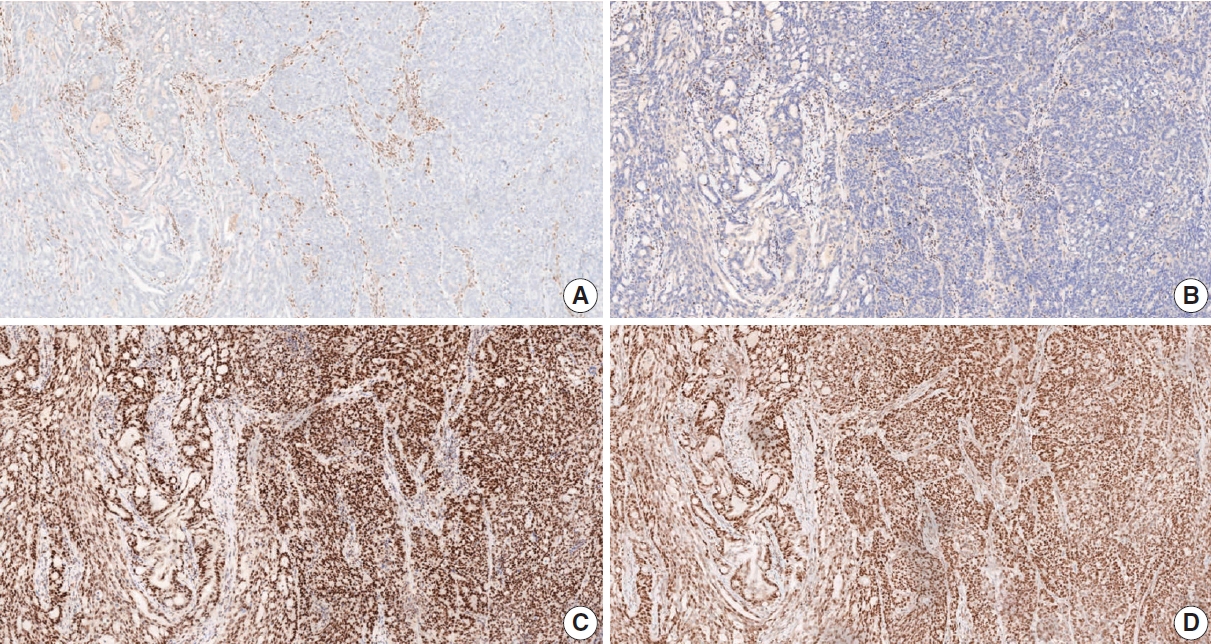
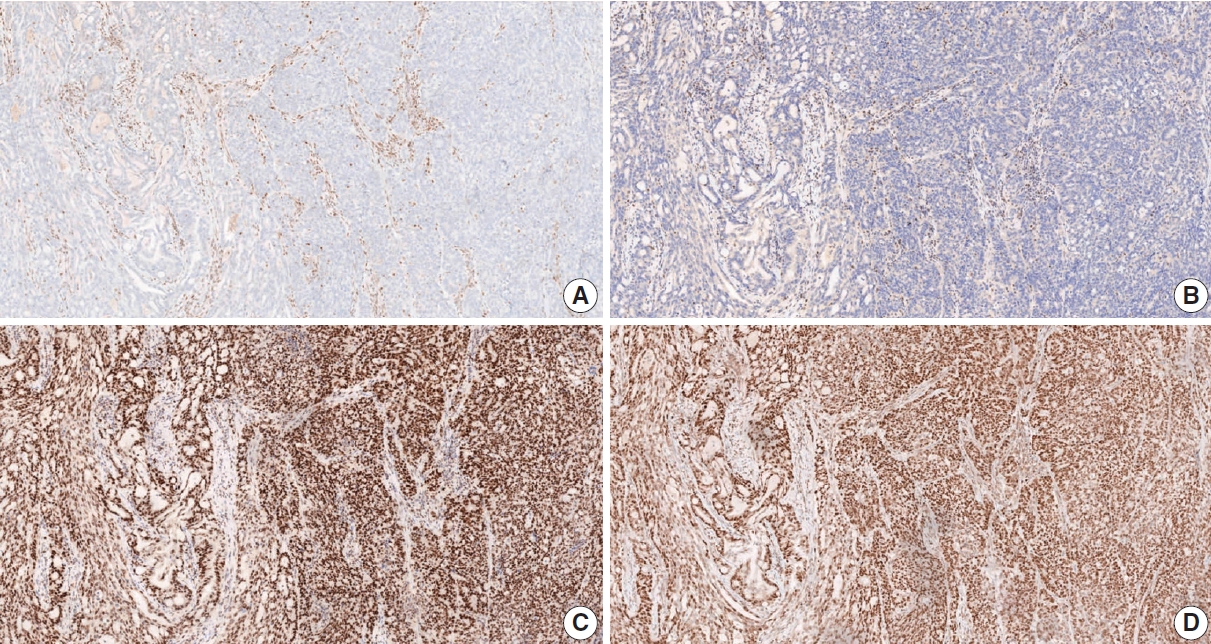
- IHC for MMR proteins in GC samples is a simple and useful practice to determine dMMR. This method shows performance characteristics similar to MSI detection by PCR and a high concordance rate (>90%) [138]. The use of all four proteins, MLH1, MSH2, MSH6, and PMS2, is recommended for the IHC test. However, in more than 90% of cases, MSI-GC is associated with MLH1 and/or PMS2 losses by hypermethylation of the MLH1 gene. Because this IHC method is based on the ubiquitous expression of the MMR proteins in cell nuclei, nuclear staining should be checked when determining MMR positivity [22,119]. The presence of internal positive controls such as normal mucosa, lymphocytes, or stromal cells is essential for the interpretation of results [119]. dMMR is determined when the nuclear expression of at least one MMR protein is absent (Fig. 12) [22]. Heterogeneity of IHC or abnormal staining (cytoplasmic or membranous staining) is sometimes observed [138-143]. When it is difficult to interpret the IHC results, the test should be reported as “undetermined,” and PCR-based testing or NGS is recommended to confirm the MMR status. Using both IHC and PCR analyses for the detection of MSI-H/dMMR can reduce indeterminacy in the results.
- EBV-associated gastric carcinoma belongs to one of four types of molecular classification suggested by the Cancer Genome Atlas (TCGA) [124]. Virus-host interactions play a pivotal role in EBV-induced carcinogenesis [144]. In EBV-associated gastric carcinoma, BamHI-A rightward frame 1 (BARF1) and latent membrane 2A (LMP2A) are putative viral oncogenes [145-147]. Once EBV enters the epithelium, EBV DNA methylation occurs globally. Hypermethylation of the CpG island promoter occurs throughout human cellular progress, which inactivates tumor suppressor genes [148]. Unique methylation leading to CDKN2A (p16) downregulation seems to be essential [124]. Eventually, EBV-infected gastric epithelial cells begin clonal growth, and gene mutations in EBV-infected cells lead to carcinogenesis [144]. EBV-associated gastric carcinoma is molecularly characterized by frequent mutations in phosphatidylinositol-4,5-biphosphate 3-kinase catalytic subunit α (PIK3CA) [124] and ATrich interaction domain 1A(ARID1A)[125], rare TP53 mutations [124], and the overexpression of interferon-γ [149] and programmed death ligand 1 (PD-L1) [124,150].
- EBV-associated gastric carcinoma has distinct histologic, genetic, and immune microenvironmental features. Notably, EBV-associated gastric carcinomas exhibit a dramatic response to pembrolizumab immunotherapy (100% overall response rate) [130]. EBV positivity can be a good indication for immunotherapy in GC. Moreover, in submucosal invasive GC, EBV positivity has been associated with a low risk of lymph node metastasis [151,152].
- ISH for EBV-encoded small RNAs (EBERs) is the most suitable and widely used method to detect EBV in formalin-fixed paraffin-embedded tissues and cytology specimens [153,154]. It is a highly sensitive detection method because of the large number of EBERs (106–107 copies/cell) [19], but it cannot be used for quantitative analysis of viral particles. Several commercial probes for EBERs are available, in which EBERs labeled with biotin, digoxigenin, or fluorescein can be visualized by microscopic examination. In most EBV-associated GCs, EBER signals are observed with strong intensity in almost all cancer cell nuclei. In certain cases, EBER signals are heterogeneous, i.e., positive only in a focal portion of the cancer or mixed—weak to strong—intensity (Fig. 13). Recently, focal positivity of EBER signals was reported in 18% of EBV-associated GC cases in Germany [155]. In daily practice in Korea, however, intratumoral heterogeneity of EBER signals is not as high as in those German cases. Whether focal negative/weak intensity represents an absence of EBV infection or a subcritical or insufficient copy number of EBERs remains unclear [156]. EBER signals are rarely detected in intratumoral or peritumoral lymphocytes, which originate from peripheral B lymphocytes infected with EBV in a latent state.
- The programmed death-1 receptor (PD-1)–PD-L1 interaction is one of the major mechanisms of immune modulation that allow T-cell inactivation and tumor immune evasion [157]. Blocking the PD-1/PD-L1 pathway is a standard therapeutic strategy for various solid tumors, including GCs [158].
- Pembrolizumab was granted accelerated FDA-approval as a third-line treatment of GC based on the findings of the phase 2 KEYNOTE-059 trial, which demonstrated its treatment benefit in advanced GC patients with PD-L1 combined positive score (CPS) positivity (CPS ≥1). Accompanying approval was granted for the PD-L1 IHC 22C3 pharmDx assay on the Autostainer Link 48 platform as a companion diagnostic assay [159]. However, the subsequent phase 3 KEYNOTE-061 trial failed to demonstrate a significant survival improvement in PD-L1-positive GC patients [160].
- Another phase 3 trial, CheckMate-649, demonstrated the efficacy of nivolumab in combination with fluoropyrimidine and platinum-based chemotherapy as a first-line treatment for HER2-negative advanced or metastatic GC, gastroesophageal junction cancer, and esophageal adenocarcinoma patients with PD-L1 CPS ≥5 [161]. In that trial, PD-L1 expression was determined using the PD-L1 IHC 28-8 pharmDx assay on the Autostainer Link 48 platform. Recently, that assay earned the CE-IVD mark in Europe as a companion diagnostic for identifying candidates for nivolumab treatment.
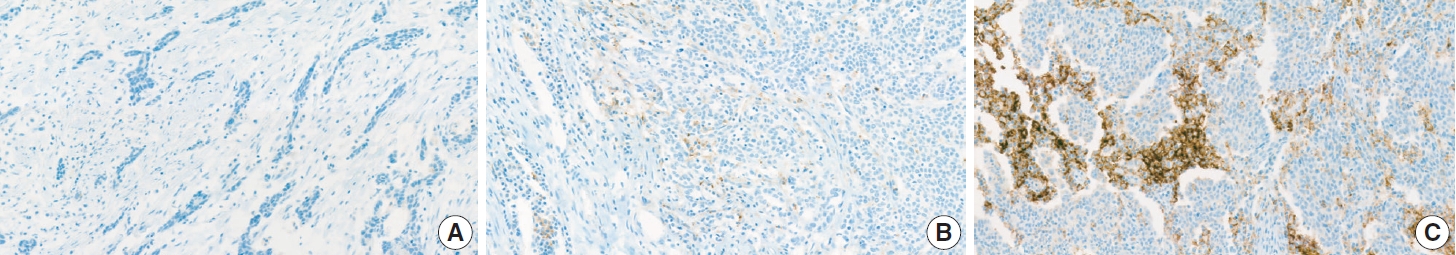
- Both assays share the CPS scoring system to determine PD-L1 expression, which is the number of PD-L1–stained cells (tumor cells, lymphocytes, and macrophages) divided by the total number of viable tumor cells, multiplied by 100. For adequate evaluation, a specimen containing a minimum of 100 viable tumor cells is required [162]. A PD-L1–stained tumor cell should present partial or complete membrane staining of viable cells with more than faint staining intensity (≥1+). PD-L1–stained immune cells include only mononuclear inflammatory cells (lymphocytes or macrophages) within tumor nests and adjacent stroma and show membrane and/or cytoplasmic staining. Other stromal cells such as fibroblasts, neutrophils, and plasma cells should be excluded from the CPS numerator. If the result of the calculation exceeds 100, it is presented as a maximum score of 100. If the PD-L1 staining shows heterogeneous results, the final CPS should be estimated by calculating each area’s CPS result (Fig. 14).
- Because two different PD-L1 assays have been approved based on different CPS cutoff values, the interpretation of PD-L1 positivity should be based on the CPS cutoff value appropriate to the assay used for evaluation. The PD-L1 IHC 22C3 pharmDx assay uses CPS ≥1 for CPS positivity, and the 28-8 pharmDx assay uses CPS ≥5. The report should specify the assay type and appropriate cutoff value used for the PD-L1 positivity interpretation.
- Previous studies have reported changes in PD-L1 expression during chemotherapy [163,164] and discrepancies between primary and metastatic lesions [164,165]. Therefore, re-evaluation of PD-L1 IHC in secondary, recurrent, and metastatic lesions is recommended for GC patients.
- Recently identified molecular profiles are not only important for improving our understanding of driver alterations involved in gastric carcinogenesis, but also for identifying clinically relevant biomarkers and new potential therapeutic targets [124,125]. Therefore, the clinical need for NGS in AGCs is increasing.
- According to the recent National Comprehensive Cancer Network (NCCN) guideline, the biomarkers implicated in clinical management of AGC include HER2, MSI, PD-L1, tumor mutation burden (TMB) status, and neurotrophic tyrosine receptor kinase (NTRK) gene fusion [100]. Among these, TMB can only be assessed using NGS, and NTRK fusion is best evaluated using NGS (preferential RNA sequencing) [166]. Alternatively, it can be screened with TRK IHC, and then sequencing can be performed in positive cases [166]. Some other targets also showed promising clinical results in advanced GC, such as fibroblast growth factor receptor 2 (FGFR2) amplification [167], epidermal growth factor receptor (EGFR) amplification [168], MET amplification [169], and alterations of homologous recombination deficiency–related genes [170]. In addition, there are very rare (prevalence <1%) targetable tissue–agnostic variants [171] such as BRAF V600E [172], anaplastic lymphoma kinase (ALK) fusion [173], and reactive oxygen species 1 (ROS1) fusion [174].
- TMB is defined as the total number of somatic coding mutations in a tumor and represents an emerging biomarker for immunotherapy response in cancer patients [175]. The exploratory analysis for KEYNOTE-062 suggested an association between TMB and the clinical efficacy of first-line pembrolizumab-based therapy in patients with advanced GC [176]. Although whole exome sequencing is considered the gold standard for TMB, recent targeted gene panels have also provided accurate quantification [175]. The lack of harmonization in panel-based TMB quantification and lack of robust predictive cutoffs are currently some of the main limitations of TMB as a biomarker in clinical practice [175].
- The gold standard for MSI detection is PCR or IHC. Recently, several MSI detection methods based on NGS have shown high concordance (>95%) with the conventional PCR-based assay [171,177,178]. The recent NCCN guidelines indicate that sequencing via a validated NGS assay may be used to determine MSI status and other biomarkers when limited tissue is available for testing [100].
- Tissue preparation is one of the most important factors for getting accurate and reliable results from NGS. In general, the total DNA and RNA requirements range from 10 to 300 ng for targeted gene panels [179]. Tissue specimen requirements are formalin-fixed, paraffin-embedded tissue or cytology specimens [179]. The minimum sample requirement for reliable sequencing results is a specimen with a tumor fraction and surface area >10%–20% and 5 mm2, respectively [179].
- GC is classified as the gastric type, intestinal type, mixed type, or unclassified type based on the expression of MUC5AC, MUC6, MUC2, and CD10 [3]. The gastric type is positive for MUC5AC and/or MUC6, and the intestinal type is positive for MUC2 and/ or CD10. The mixed type is positive for both gastric and intestinal mucins, and the unclassified type is negative for both.
- Molecular profiles of GCs have been published in recent studies by TCGA and the Asian Cancer Research Group (ACRG). TCGA classified GCs into EBV, MSI, genomically stable, and chromosomally unstable [124]. In contrast, ACRG published a molecular classification of MSI, microsatellite stable/epithelial mesenchymal transition (MSS/EMT), MSS/TP53+, and MSS/TP53– [125]. The MSS/EMT subtype is closely associated with the SRC and PCC histology and Lauren’s diffuse type, and patient survival is poor. The EBV and MSI subtypes are related to the histologic type of adenocarcinoma with lymphoid stroma and have relatively better prognosis. High TMB and increased expression of PD-L1 are commonly reported in the EBV and MSI subtypes.
- Several studies have reported that these molecular classifications could be reproduced in GCs using simple techniques, including EBV ISH, MSI testing, MMR IHC, E-cadherin IHC, and p53 IHC [127,180,181]. Using those tests, GC is classified as EBV, MSI, EMT, altered p53, and not altered p53. Those molecular subtypes showed distinct clinicopathologic characteristics.
APPLICATION OF STANDARD PATHOLOGY REPORT
Gastrectomy (specimen) type
Gross type
Previous treatment
Tumor focality
Tumor location
Tumor size
Tumor regression grade
Depth of invasion
Regional lymph node metastasis
Extranodal tumor extension
Lymphovascular invasion
Perineural invasion
Pre-existing adenoma
Associated findings
Separate lesions
Description of the specimen
Sectioning of the specimen
Histologic type and components
Tumor size
Depth of invasion
Resection margin
Ulcer
Cases with adenoma components
En bloc resection
Lymphatic/venous invasion
Histologic types in biopsy specimens
Lauren classification
Adenoma
Helicobacter pylori
Human epidermal growth factor receptor 2 testing
Microsatellite instability and mismatch repair deficiency
EBV testing
PD-L1 immunohistochemistry
Next generation sequencing
Mucin phenotype
Easy methods for molecular classification
Supplementary Information
Ethics Statement
Not applicable.
Availability of Data and Material
Data sharing not applicable to this article as no datasets were generated or analyzed during the study.
Code Availability
Not applicable.
Author contributions
Conceptualization: YSP, MCK, BHK, HSL, KMK, JMK, SHL, MYC. Project administration: KMK, JMK, SHL, MYC. Supervision: DWK, YC, HK, KMK, DYP, SYJ, JMK, YJC, HKC, MSC, MYC. Writing—original draft preparation: YSP, MCK, BHK, HSL, SHL. Writing—review & editing: YSP, MCK, BHK, HSL, DWK, MJG, ORS, YC, WL, HK, HIS, KMK, HSK, GK, DYP, SYJ, JMK, YJC, HKC, SA, MSC, SHH, YK, ANS, SHL, MYC. Approval of final manuscript: all authors.
Conflicts of Interest
S.H.L., a contributing editor of the Journal of Pathology and Translational Medicine, was not involved in the editorial evaluation or decision to publish this article. All remaining authors have declared no conflicts of interest.
Funding Statement
This research was supported by a grant of Patient-Centered Clinical Research Coordinating Center (PACEN) funded by the Ministry of Health & Welfare, Republic of Korea (grant number: HC20C0123).

| Standard and Conditional data elements | |||
|---|---|---|---|
| Gastrectomy (specimen) typea | |||
| □ Total gastrectomy | |||
| □ Distal (subtotal) gastrectomy | |||
| □ Proximal gastrectomy | |||
| □ Wedge resection | |||
| □ Others ( _______________ ) | |||
| Gross typea | |||
| □ EGC type | |||
| □ EGC type I/IIa/IIb/IIc/III | |||
| □ Mixed EGC type ( ____________ ) | |||
| □ AGC type | |||
| □ Borrmann type 1/2/3/4/unclassifiable | |||
| □ Others ( _______________ ) | |||
| Residual with previous treatmenta (when applicable) | |||
| □ Residual | |||
| □ Previous treatment | |||
| □ Chemotherapy | |||
| □ Chemoradiotherapy | |||
| □ Endoscopic mucosal resection | |||
| □ Endoscopic submucosal dissection | |||
| □ Unknown | |||
| □ Others ( ____________ ) | |||
| Tumor focalitya | |||
| □ Single | |||
| □ Multiple | |||
| Tumor locationa | |||
| □ Involvement | |||
| □ Esophagus/Upper/Middle/Lower third of the stomach/Duodenum | |||
| □ Center | |||
| □ Cardia/Fundus/Body/Antrum/Pylorus | |||
| □ Lesser curvature/Greater curvature/Anterior wall/Posterior wall | |||
| □ Others ( ____________ ) | |||
| Tumor sizea | |||
| One largest dimension | |||
| □ ___ cm | |||
| Tumor sizeb | |||
| Secondary or tertiary tumor dimensions | |||
| □ ___ × ___ cm | |||
| □ ___ × ___ × ___ cm | |||
| Histologic typea | |||
| According to the principles described in “Histologic classification” section | |||
| □ WHO | |||
| □ Lauren | |||
| Tumor regression gradea (when applicable) | |||
| □ Grade 0: Complete response (no viable cancer cells) | |||
| □ Grade 1: Near complete response (single cells or rare small groups of cancer cells) | |||
| □ Grade 2: Partial response (residual cancer with evident tumor regression, but more than single cells or rare small groups of cancer cells) | |||
| □ Grade 3: Poor or no response (extensive residual cancer with no evident tumor regression) | |||
| Lymph node tumor regressionb (when applicable) | |||
| □ Not identified | |||
| □ Present | |||
| Depth of invasion (pT)a | |||
| □ Invades lamina propria (pT1a) | |||
| □ Invades muscularis mucosae (pT1a) | |||
| □ Invades submucosa (sm1/sm2/sm3) (pT1b) | |||
| □ Invades proper muscle (pT2) | |||
| □ Invades subserosa (pT3) | |||
| □ Invades serosa (visceral peritoneum) (pT4a) | |||
| □ Directly invades adjacent structure (pT4b) | |||
| specify ( _____________ ) | |||
| Resection margina | |||
| □ Proximal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Distal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Circumferential resection marginb | |||
| Applied in EGJ or cardia cancer | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Regional lymph node metastasisa | |||
| At least 16 regional lymph nodes should be assessed | |||
| □ no metastasis in ____ regional lymph nodes | |||
| □ metastasis in ___ out of ___ regional lymph nodes | |||
| Extranodal tumor extensionb | |||
| □ Not identified | |||
| □ Present | |||
| Isolated tumor cell clustersb | |||
| Applied in incidentally identified tumor cell cluster less than 0.2 mm in greatest dimension with no other regional lymph node metastasis (pN0) | |||
| □ Present [pN0 (i+)] | |||
| Lymphovascular invasiona | |||
| □ Not identified | |||
| □ Present | |||
| Venous invasionb | |||
| Applied when identified in large vessels with an identifiable smooth muscle layer or elastic lamina | |||
| □ Not identified | |||
| □ Present | |||
| Perineural invasiona | |||
| □ Not identified | |||
| □ Present | |||
| Pre-existing adenomaa (when present) | |||
| Used if the carcinoma is within the adenoma | |||
| □ Tubular/Tubulovillous/Villous adenoma | |||
| □ Low grade dysplasia/High grade dysplasia | |||
| Associated findingsa (when present) | |||
| □ Tumor perforation | |||
| □ Serosal (peritoneal, mesenteric) seeding | |||
| □ Distant metastasis | |||
| Other organ, specify: ______________ | |||
| Distant lymph node | |||
| Separate lesionsa (when present) | |||
| □ Peptic ulcer | |||
| □ Adenoma | |||
| □ GIST | |||
| □ Others ( ____________ ) | |||
| Standard and Conditional data elements | |||
|---|---|---|---|
| Specimen sizea | |||
| □ ___ × ___ cm | |||
| Gross type of tumora | |||
| Same as method of surgical specimen | |||
| Tumor sizea | |||
| One largest dimension | |||
| □ _____ cm | |||
| Histologic typea | |||
| According to the principles described in “Histologic classification” section | |||
| □ WHO | |||
| □ Lauren | |||
| Histologic componentsb | |||
| All morphologic components of tumor cell may be described | |||
| Depth of invasion (pT)a | |||
| □ Invades lamina propria (pT1a) | |||
| □ Invades muscularis mucosae (pT1a) | |||
| □ Invades submucosa (submucosal depth: _____ mm or μm) | |||
| □ Invades proper muscle (pT2) | |||
| Depth of invasion (pT)b | |||
| In case of submucosa invasion, the invasion width can be additionally described | |||
| □ invades submucosa (submucosal depth: _____ mm or μm) (submucosal width: _____ mm) | |||
| Resection margina | |||
| □ Lateral margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Deep margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Resection marginb | |||
| □ Proximal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Distal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Anterior margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Posterior margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Deep margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Ulcerationa | |||
| □ Absent | |||
| □ Present | |||
| Ulcerationb | |||
| □ Absent | |||
| □ Non-significant (≤ 4 mm) | |||
| □ Significant (> 4 mm) | |||
| Cases with adenoma componentsa | |||
| □ Absent | |||
| □ Present | |||
| specify: ______________ | |||
| En bloc resectiona | |||
| □ Yes | |||
| □ No (piecemeal/tearing) | |||
| Lymphatic invasiona | |||
| □ Not identified | |||
| □ Present | |||
| Venous invasiona | |||
| □ Not identified | |||
| □ Present | |||
| Molecular markers | ||
|---|---|---|
| All molecular markers are “conditional data element” | ||
| HER2 immunohistochemistry | ||
| □ Negative (0/1+) | ||
| □ Equivocal (2+) | ||
| □ Positive (3+) | ||
| □ Undetermined (explain): | ||
| HER2 (ERBB2) in situ hybridization | ||
| Number of invasive cancer cells counted: ______ cells | ||
| □ Using dual-probe assay | ||
| □ HER2 (ERBB2)/CEP17 ratio: ______ | ||
| □ Average number of HER2 (ERBB2) signals per cancer cell: ______ | ||
| □ Average number of CEP17 signals per cancer cell: ______ | ||
| □ Using single-probe assay | ||
| □ Average number of HER2 (ERBB2) signals per cancer cell: ______ | ||
| Summary: Negative/Positive for HER2 (ERBB2) gene amplification | ||
| □ Undetermined (explain): | ||
| Microsatellite instability (MSI) | ||
| Summary: | ||
| □ Microsatellite stable (MSS) | ||
| □ Microsatellite instability-low (MSI-L) | ||
| □ Microsatellite instability-high (MSI-H) | ||
| □ Undetermined (explain)a | ||
| DNA mismatch repair immunohistochemistry | ||
| MLH1: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| MSH2: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| PMS2: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| MSH6: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| Summary: | ||
| □ DNA mismatch repair deficiency (was/was not) observed | ||
| □ Because it is difficult to determine DNA mismatch repair deficiency, PCR-based testing and/or NGS for MSI is recommended. | ||
| In situ hybridization for Epstein-Barr virus–encoded small RNAs | ||
| □ Positive [diffuse/heterogenous (focal and/or mixed intensity)]b,c | ||
| □ Negative | ||
| Summary: Epstein-Barr virus–associated gastric carcinoma | ||
| PD-L1 immunohistochemistry | ||
| PD-L1 [Antibody (22C3 PharmDx/22C3 conc. Ventana/28-8 PharmDx/others:______)]: | ||
| □ CPS = ________ | ||
HER2, human epidermal growth factor receptor 2; CEP17, centromeric region of chromosome 17; MLH1, mutL homolog 1; MSH2, mutS homolog 2; PMS2, PMS1 homolog 2; MSH6, mutS homolog 6; PCR, polymerase chain reaction; NGS, next-generation sequencing; PD-L1, programmed death ligand 1; CPS, combined positive score.
aBecause it is difficult to determine MSI status, mismatch repair immunohistochemistry and/or NGS is recommended;
bChecking the signal pattern is optional;
cThe term “Epstein-Barr virus–associated gastric carcinoma” applies to positive cases.
- 1. Sung H, Ferlay J, Siegel RL, et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021; 71: 209-49. ArticlePubMedPDF
- 2. Hong S, Won YJ, Lee JJ, et al. Cancer statistics in Korea: incidence, mortality, survival, and prevalence in 2018. Cancer Res Treat 2021; 53: 301-15. ArticlePubMedPMCPDF
- 3. Kim WH, Park CK, Kim YB, et al. A standardized pathology report for gastric cancer. Korean J Pathol 2005; 39: 106-13.
- 4. World Health Organization. WHO classification of tumours: digestive system tumours. Geneva: World Health Organization, 2019.
- 5. Guideline Committee of the Korean Gastric Cancer Association (KGCA); Development Working Group and Review Panel. Korean practice guideline for fastric cancer 2018: an evidence-based, multi-disciplinary approach. J Gastric Cancer 2019; 19: 1-48. ArticlePubMedPMCPDF
- 6. Muro K, Chung HC, Shankaran V, et al. Pembrolizumab for patients with PD-L1-positive advanced gastric cancer (KEYNOTE-012): a multicentre, open-label, phase 1b trial. Lancet Oncol 2016; 17: 717-26. ArticlePubMed
- 7. Bang YJ, Van Cutsem E, Feyereislova A, et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet 2010; 376: 687-97. PubMed
- 8. Amin MB, Edge SB, Greene FL, et al. AJCC cancer staging manual. 8th ed. Cham: Springer, 2017.
- 9. Japanese Gastric Cancer Association. Japanese classification of gastric carcinoma: 3rd English edition. Gastric Cancer 2011; 14: 101-12. ArticlePubMedPDF
- 10. Fritz A, Percy C, Jack A, et al. International Classification of Diseases for Oncology (ICD-O). 3rd ed. Lyon: International Agency for Research on Cancer, 2013.
- 11. Shi C, Berlin J, Branton PA, et al. Protocol for the examination of specimens from patients with carcinoma of the stomach [Internet]. Northfield: College of American Pathologists, 2020 [cited 2022 Dec 1]. Available from: https://documents.cap.org/protocols/cp-giupper-esophagus-20-4100.pdf.
- 12. Shi C, Badgwell BD, Grabsch HI, et al. Data set for reporting carcinoma of the stomach in gastrectomy. Arch Pathol Lab Med 2022; 146: 1072-83. ArticlePubMedPDF
- 13. Cunningham D, Allum WH, Stenning SP, et al. Perioperative chemotherapy versus surgery alone for resectable gastroesophageal cancer. N Engl J Med 2006; 355: 11-20. ArticlePubMed
- 14. Zhang X, Liang H, Li Z, et al. Perioperative or postoperative adjuvant oxaliplatin with S-1 versus adjuvant oxaliplatin with capecitabine in patients with locally advanced gastric or gastro-oesophageal junction adenocarcinoma undergoing D2 gastrectomy (RESOLVE): an open-label, superiority and non-inferiority, phase 3 randomised controlled trial. Lancet Oncol 2021; 22: 1081-92. ArticlePubMed
- 15. Kang YK, Yook JH, Park YK, et al. PRODIGY: a phase III study of neoadjuvant docetaxel, oxaliplatin, and S-1 plus surgery and adjuvant S-1 versus surgery and adjuvant S-1 for resectable advanced gastric cancer. J Clin Oncol 2021; 39: 2903-13. PubMedPMC
- 16. Langer R, Becker K. Tumor regression grading of gastrointestinal cancers after neoadjuvant therapy. Virchows Arch 2018; 472: 175-86. ArticlePubMedPDF
- 17. Mandard AM, Dalibard F, Mandard JC, et al. Pathologic assessment of tumor regression after preoperative chemoradiotherapy of esophageal carcinoma: clinicopathologic correlations. Cancer 1994; 73: 2680-6. ArticlePubMed
- 18. Dworak O, Keilholz L, Hoffmann A. Pathological features of rectal cancer after preoperative radiochemotherapy. Int J Colorectal Dis 1997; 12: 19-23. ArticlePubMedPDF
- 19. Becker K, Mueller JD, Schulmacher C, et al. Histomorphology and grading of regression in gastric carcinoma treated with neoadjuvant chemotherapy. Cancer 2003; 98: 1521-30. ArticlePubMed
- 20. Mirza A, Naveed A, Hayes S, et al. Assessment of histopathological response in gastric and gastro-oesophageal junction adenocarcinoma following neoadjuvant chemotherapy: which scoring system to use? ISRN Pathol 2012; 2012: 519351.ArticlePDF
- 21. Karamitopoulou E, Thies S, Zlobec I, et al. Assessment of tumor regression of esophageal adenocarcinomas after neoadjuvant chemotherapy: comparison of 2 commonly used scoring approaches. Am J Surg Pathol 2014; 38: 1551-6. PubMed
- 22. Kim BH, Kim JM, Kang GH, et al. Standardized pathology report for colorectal cancer, 2nd edition. J Pathol Transl Med 2020; 54: 1-19. ArticlePubMedPDF
- 23. Tsekrekos A, Vieth M, Ndegwa N, et al. Interobserver agreement of a gastric adenocarcinoma tumor regression grading system that incorporates assessment of lymph nodes. Hum Pathol 2021; 116: 94-101. ArticlePubMed
- 24. Davies AR, Myoteri D, Zylstra J, et al. Lymph node regression and survival following neoadjuvant chemotherapy in oesophageal adenocarcinoma. Br J Surg 2018; 105: 1639-49. ArticlePubMedPDF
- 25. Reim D, Novotny A, Friess H, et al. Significance of tumour regression in lymph node metastases of gastric and gastro-oesophageal junction adenocarcinomas. J Pathol Clin Res 2020; 6: 263-72. ArticlePubMedPMCPDF
- 26. Smyth EC, Fassan M, Cunningham D, et al. Effect of pathologic tumor response and nodal status on survival in the Medical Research Council adjuvant gastric infusional chemotherapy Trial. J Clin Oncol 2016; 34: 2721-7. ArticlePubMedPMC
- 27. Koh YW, Park YS, Ryu MH, et al. Postoperative nodal status and diffuse-type histology are independent prognostic factors in resectable advanced gastric carcinomas after preoperative chemotherapy. Am J Surg Pathol 2013; 37: 1022-9. ArticlePubMed
- 28. Veronese N, Fassan M, Wood LD, et al. Extranodal extension of nodal metastases is a poor prognostic indicator in gastric cancer: a systematic review and meta-analysis. J Gastrointest Surg 2016; 20: 1692-8. ArticlePubMedPDF
- 29. Lee IS, Park YS, Ryu MH, et al. Impact of extranodal extension on prognosis in lymph node-positive gastric cancer. Br J Surg 2014; 101: 1576-84. ArticlePubMedPDF
- 30. Lee IS, Kang HJ, Park YS, et al. Prognostic impact of extranodal extension in stage 1B gastric carcinomas. Surg Oncol 2018; 27: 299-305. ArticlePubMed
- 31. Araki I, Hosoda K, Yamashita K, et al. Prognostic impact of venous invasion in stage IB node-negative gastric cancer. Gastric Cancer 2015; 18: 297-305. ArticlePubMedPDF
- 32. Takeuchi A, Ojima T, Katsuda M, et al. Venous invasion is a risk factor for recurrence of pT1 gastric cancer with lymph node metastasis. J Gastrointest Surg 2022; 26: 757-63. ArticlePubMedPDF
- 33. Nishibeppu K, Komatsu S, Ichikawa D, et al. Venous invasion as a risk factor for recurrence after gastrectomy followed by chemotherapy for stage III gastric cancer. BMC Cancer 2018; 18: 108.ArticlePubMedPMCPDF
- 34. Liebig C, Ayala G, Wilks JA, Berger DH, Albo D. Perineural invasion in cancer: a review of the literature. Cancer 2009; 115: 3379-91. ArticlePubMed
- 35. Hanaoka N, Tanabe S, Mikami T, Okayasu I, Saigenji K. Mixedhistologic-type submucosal invasive gastric cancer as a risk factor for lymph node metastasis: feasibility of endoscopic submucosal dissection. Endoscopy 2009; 41: 427-32. ArticlePubMed
- 36. Takizawa K, Ono H, Kakushima N, et al. Risk of lymph node metastases from intramucosal gastric cancer in relation to histological types: how to manage the mixed histological type for endoscopic submucosal dissection. Gastric Cancer 2013; 16: 531-6. ArticlePubMedPDF
- 37. Lee JH, Choi IJ, Han HS, et al. Risk of lymph node metastasis in differentiated type mucosal early gastric cancer mixed with minor undifferentiated type histology. Ann Surg Oncol 2015; 22: 1813-9. ArticlePubMedPDF
- 38. Sekiguchi M, Oda I, Taniguchi H, et al. Risk stratification and predictive risk-scoring model for lymph node metastasis in early gastric cancer. J Gastroenterol 2016; 51: 961-70. ArticlePubMedPDF
- 39. Seo HS, Lee GE, Kang MG, Han KH, Jung ES, Song KY. Mixed histology is a risk factor for lymph node metastasis in early gastric cancer. J Surg Res 2019; 236: 271-7. ArticlePubMed
- 40. Horiuchi Y, Ida S, Yamamoto N, et al. Feasibility of further expansion of the indications for endoscopic submucosal dissection in undifferentiated-type early gastric cancer. Gastric Cancer 2020; 23: 285-92. ArticlePubMedPDF
- 41. Ha TK, An JY, Youn HK, Noh JH, Sohn TS, Kim S. Indication for endoscopic mucosal resection in early signet ring cell gastric cancer. Ann Surg Oncol 2008; 15: 508-13. ArticlePubMedPDF
- 42. Kim BS, Oh ST, Yook JH, Kim BS. Signet ring cell type and other histologic types: differing clinical course and prognosis in T1 gastric cancer. Surgery 2014; 155: 1030-5. ArticlePubMed
- 43. Guo CG, Zhao DB, Liu Q, et al. Risk factors for lymph node metastasis in early gastric cancer with signet ring cell carcinoma. J Gastrointest Surg 2015; 19: 1958-65. ArticlePubMedPDF
- 44. Huh CW, Jung DH, Kim JH, et al. Signet ring cell mixed histology may show more aggressive behavior than other histologies in early gastric cancer. J Surg Oncol 2013; 107: 124-9. ArticlePubMedPDF
- 45. Lee IS, Lee S, Park YS, Gong CS, Yook JH, Kim BS. Applicability of endoscopic submucosal dissection for undifferentiated early gastric cancer: Mixed histology of poorly differentiated adenocarcinoma and signet ring cell carcinoma is a worse predictive factor of nodal metastasis. Surg Oncol 2017; 26: 8-12. ArticlePubMed
- 46. Kim YH, Park JH, Park CK, et al. Histologic purity of signet ring cell carcinoma is a favorable risk factor for lymph node metastasis in poorly cohesive, submucosa-invasive early gastric carcinoma. Gastric Cancer 2017; 20: 583-90. ArticlePubMedPDF
- 47. Chu Y, Mao T, Li X, et al. Predictors of lymph node metastasis and differences between pure and mixed histologic types of early gastric signet-ring cell carcinomas. Am J Surg Pathol 2020; 44: 934-42. ArticlePubMedPMC
- 48. Japanese Gastric Cancer Society. Regulations for the treatment of gastric cancer. 15th ed. Tokyo: Kanehara Publishing, 2017.
- 49. Kim JY, Kim WG, Jeon TY, et al. Lymph node metastasis in early gastric cancer: evaluation of a novel method for measuring submucosal invasion and development of a nodal predicting index. Hum Pathol 2013; 44: 2829-36. ArticlePubMed
- 50. Ma DW, Lee SJ, Kook MC, et al. The suggestion of revised criteria for endoscopic resection of differentiated-type submucosal gastric cancer. Ann Surg Oncol 2020; 27: 795-801. ArticlePDF
- 51. Choi JY, Park YS, Jung HY, et al. Identifying predictors of lymph node metastasis after endoscopic resection in patients with minute submucosal cancer of the stomach. Surg Endosc 2015; 29: 1476-83. ArticlePubMedPDF
- 52. Gotoda T, Yanagisawa A, Sasako M, et al. Incidence of lymph node metastasis from early gastric cancer: estimation with a large number of cases at two large centers. Gastric Cancer 2000; 3: 219-25. ArticlePubMedPDF
- 53. Park SM, Kim BW, Kim JS, Kim YW, Kim GJ, Ryu SJ. Can endoscopic ulcerations in early gastric cancer be clearly defined before endoscopic resection? A survey among endoscopists. Clin Endosc 2017; 50: 473-8. ArticlePubMedPMCPDF
- 54. Kim JM, Sohn JH, Cho MY, et al. Pre- and post-ESD discrepancies in clinicopathologic criteria in early gastric cancer: the NECA-Korea ESD for Early Gastric Cancer Prospective Study (NKeep). Gastric Cancer 2016; 19: 1104-13. ArticlePubMedPDF
- 55. Yabuuchi Y, Takizawa K, Kakushima N, et al. Discrepancy between endoscopic and pathological ulcerative findings in clinical intramucosal early gastric cancer. Gastric Cancer 2021; 24: 691-700. ArticlePubMedPDF
- 56. Shimoda T, Kushima R, Ono H. Histological features of peptic ulceration and biopsy scar of gastric ESD specimen. Stomach Intestine 2013; 48: 16-24.
- 57. Japanese Gastric Cancer Association. Japanese gastric cancer treatment guidelines 2018 (5th edition). Gastric Cancer 2021; 24: 1-21. ArticlePubMedPDF
- 58. Takizawa K, Ono H, Hasuike N, et al. A nonrandomized, singlearm confirmatory trial of expanded endoscopic submucosal dissection indication for undifferentiated early gastric cancer: Japan Clinical Oncology Group study (JCOG1009/1010). Gastric Cancer 2021; 24: 479-91. ArticlePubMedPDF
- 59. Hatta W, Gotoda T, Oyama T, et al. A scoring system to stratify curability after endoscopic submucosal dissection for early gastric cancer: "eCura system". Am J Gastroenterol 2017; 112: 874-81. ArticlePubMed
- 60. Uesugi N, Sugai T, Sugimoto R, et al. Clinicopathological and molecular stability and methylation analyses of gastric papillary adenocarcinoma. Pathology 2017; 49: 596-603. ArticlePubMed
- 61. Lee HJ, Kim GH, Park DY, et al. Endoscopic submucosal dissection for papillary adenocarcinoma of the stomach: is it really safe? Gastric Cancer 2017; 20: 978-86. ArticlePubMedPDF
- 62. Min BH, Byeon SJ, Lee JH, et al. Lymphovascular invasion and lymph node metastasis rates in papillary adenocarcinoma of the stomach: implications for endoscopic resection. Gastric Cancer 2018; 21: 680-8. ArticlePubMedPDF
- 63. Yasuda K, Adachi Y, Shiraishi N, Maeo S, Kitano S. Papillary adenocarcinoma of the stomach. Gastric Cancer 2000; 3: 33-8. ArticlePubMedPDF
- 64. Yu H, Fang C, Chen L, et al. Worse prognosis in papillary, compared to tubular, early gastric carcinoma. J Cancer 2017; 8: 117-23. ArticlePubMedPMC
- 65. Kawamura H, Kondo Y, Osawa S, et al. A clinicopathologic study of mucinous adenocarcinoma of the stomach. Gastric Cancer 2001; 4: 83-6. ArticlePubMedPDF
- 66. Lee HH, Song KY, Park CH, Jeon HM. Undifferentiated-type gastric adenocarcinoma: prognostic impact of three histological types. World J Surg Oncol 2012; 10: 254.ArticlePubMedPMCPDF
- 67. Nakamura K, Eto K, Iwagami S, et al. Clinicopathological characteristics and prognosis of poorly cohesive cell subtype of gastric cancer. Int J Clin Oncol 2022; 27: 512-9. ArticlePubMedPDF
- 68. Roviello F, Marano L, Ambrosio MR, et al. Signet ring cell percentage in poorly cohesive gastric cancer patients: a potential novel predictor of survival. Eur J Surg Oncol 2022; 48: 561-9. ArticlePubMed
- 69. Garcia-Pelaez J, Barbosa-Matos R, Gullo I, Carneiro F, Oliveira C. Histological and mutational profile of diffuse gastric cancer: current knowledge and future challenges. Mol Oncol 2021; 15: 2841-67. ArticlePubMedPMCPDF
- 70. Kwon CH, Kim YK, Lee S, et al. Gastric poorly cohesive carcinoma: a correlative study of mutational signatures and prognostic significance based on histopathological subtypes. Histopathology 2018; 72: 556-68. ArticlePubMedPDF
- 71. Mariette C, Carneiro F, Grabsch HI, et al. Consensus on the pathological definition and classification of poorly cohesive gastric carcinoma. Gastric Cancer 2019; 22: 1-9. ArticlePubMedPDF
- 72. Han JP, Hong SJ, Kim HK. Long-term outcomes of early gastric cancer diagnosed as mixed adenocarcinoma after endoscopic submucosal dissection. J Gastroenterol Hepatol 2015; 30: 316-20. ArticlePubMedPDF
- 73. Park HK, Lee KY, Yoo MW, Hwang TS, Han HS. Mixed carcinoma as an independent prognostic factor in submucosal invasive gastric carcinoma. J Korean Med Sci 2016; 31: 866-72. ArticlePubMedPMCPDF
- 74. Horiuchi Y, Fujisaki J, Yamamoto N, et al. Undifferentiated-type predominant mixed-type early gastric cancer is a significant risk factor for requiring additional surgeries after endoscopic submucosal dissection. Sci Rep 2020; 10: 6748.ArticlePubMedPMCPDF
- 75. Ozeki Y, Hirasawa K, Sawada A, et al. Mixed histology poses a greater risk for noncurative endoscopic resection in early gastric cancers regardless of the predominant histologic types. Eur J Gastroenterol Hepatol 2021; 32: 186-93. ArticlePubMed
- 76. Shin DH, Kim GH, Lee BE, et al. Clinicopathologic features of early gastric carcinoma with lymphoid stroma and feasibility of endoscopic submucosal dissection. Surg Endosc 2017; 31: 4156-64. ArticlePubMedPDF
- 77. Iwasaki K, Suda T, Takano Y, et al. Postoperative outcomes of gastric carcinoma with lymphoid stroma. World J Surg Oncol 2020; 18: 102.ArticlePubMedPMCPDF
- 78. Huh CW, Jung DH, Kim H, et al. Clinicopathologic features of gastric carcinoma with lymphoid stroma in early gastric cancer. J Surg Oncol 2016; 114: 769-72. ArticlePubMedPDF
- 79. Inagawa S, Shimazaki J, Hori M, et al. Hepatoid adenocarcinoma of the stomach. Gastric Cancer 2001; 4: 43-52. ArticlePubMedPDF
- 80. Roh JH, Srivastava A, Lauwers GY, et al. Micropapillary carcinoma of stomach: a clinicopathologic and immunohistochemical study of 11 cases. Am J Surg Pathol 2010; 34: 1139-46. PubMed
- 81. Fujita T, Gotohda N, Kato Y, et al. Clinicopathological features of stomach cancer with invasive micropapillary component. Gastric Cancer 2012; 15: 179-87. ArticlePubMedPDF
- 82. Benedict MA, Lauwers GY, Jain D. Gastric adenocarcinoma of the fundic gland type: update and literature review. Am J Clin Pathol 2018; 149: 461-73. PubMed
- 83. Miyazawa M, Matsuda M, Yano M, et al. Gastric adenocarcinoma of the fundic gland (chief cell-predominant type): a review of endoscopic and clinicopathological features. World J Gastroenterol 2016; 22: 10523-31. ArticlePubMedPMC
- 84. Agaimy A, Rau TT, Hartmann A, Stoehr R. SMARCB1 (INI1)- negative rhabdoid carcinomas of the gastrointestinal tract: clinicopathologic and molecular study of a highly aggressive variant with literature review. Am J Surg Pathol 2014; 38: 910-20. PubMed
- 85. Willems S, Carneiro F, Geboes K. Gastric carcinoma with osteoclast-like giant cells and lymphoepithelioma-like carcinoma of the stomach: two of a kind? Histopathology 2005; 47: 331-3. ArticlePubMed
- 86. Woo HY, Bae YS, Kim JH, et al. Distinct expression profile of key molecules in crawling-type early gastric carcinoma. Gastric Cancer 2017; 20: 612-9. ArticlePubMedPDF
- 87. Okamoto N, Kawachi H, Yoshida T, et al. “Crawling-type” adenocarcinoma of the stomach: a distinct entity preceding poorly differentiated adenocarcinoma. Gastric Cancer 2013; 16: 220-32. ArticlePubMedPDF
- 88. Ushiku T, Kunita A, Kuroda R, et al. Oxyntic gland neoplasm of the stomach: expanding the spectrum and proposal of terminology. Mod Pathol 2020; 33: 206-16. ArticlePubMedPDF
- 89. Tsuji Y, Ushiku T, Shinozaki T, et al. Risk for lymph node metastasis in Epstein-Barr virus-associated gastric carcinoma with submucosal invasion. Dig Endosc 2021; 33: 592-7. ArticlePubMedPDF
- 90. Lauren P. The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma: an attempt at a histo-clinical classification. Acta Pathol Microbiol Scand 1965; 64: 31-49. PubMed
- 91. Park DY, Srivastava A, Kim GH, et al. Adenomatous and foveolar gastric dysplasia: distinct patterns of mucin expression and background intestinal metaplasia. Am J Surg Pathol 2008; 32: 524-33. ArticlePubMed
- 92. Choi WT, Brown I, Ushiku T, et al. Gastric pyloric gland adenoma: a multicentre clinicopathological study of 67 cases. Histopathology 2018; 72: 1007-14. ArticlePubMedPDF
- 93. Kim JM, Sohn JH, Cho MY, et al. Inter-observer reproducibility in the pathologic diagnosis of gastric intraepithelial neoplasia and early carcinoma in endoscopic submucosal dissection specimens: a multi-center study. Cancer Res Treat 2019; 51: 1568-77. ArticlePubMedPMCPDF
- 94. Kim JM, Cho MY, Sohn JH, et al. Diagnosis of gastric epithelial neoplasia: dilemma for Korean pathologists. World J Gastroenterol 2011; 17: 2602-10. ArticlePubMedPMC
- 95. Watanabe Y, Shimizu M, Itoh T, Nagashima K. Intraglandular necrotic debris in gastric biopsy and surgical specimens. Ann Diagn Pathol 2001; 5: 141-7. ArticlePubMed
- 96. Choi IJ, Kook MC, Kim YI, et al. Helicobacter pylori therapy for the prevention of metachronous gastric cancer. N Engl J Med 2018; 378: 1085-95. ArticlePubMed
- 97. Ono H, Yao K, Fujishiro M, et al. Guidelines for endoscopic submucosal dissection and endoscopic mucosal resection for early gastric cancer (second edition). Dig Endosc 2021; 33: 4-20. ArticlePubMedPDF
- 98. Shitara K, Bang YJ, Iwasa S, et al. Trastuzumab deruxtecan in previously treated HER2-positive gastric cancer. N Engl J Med 2020; 382: 2419-30. ArticlePubMed
- 99. FDA approves fam-trastuzumab deruxtecan-nxki for HER2-positive gastric adenocarcinomas [Internet]. Silver Spring: U.S. Food and Drug Administration, 2021 [cited 2022 Dec 1]. Available from: https://www.fda.gov/drugs/resources-information-approveddrugs/fda-approves-fam-trastuzumab-deruxtecan-nxki-her2-positive-gastric-adenocarcinomas.
- 100. Ajani JA, D’Amico TA, Bentrem DJ, et al. Gastric cancer, version 2.2022, NCCN Clinical Practice Guidelines in Oncology. J Natl Compr Canc Netw 2022; 20: 167-92. PubMed
- 101. Hofmann M, Stoss O, Shi D, et al. Assessment of a HER2 scoring system for gastric cancer: results from a validation study. Histopathology 2008; 52: 797-805. ArticlePubMed
- 102. Ahn S, Ahn S, Van Vrancken M, et al. Ideal number of biopsy tumor fragments for predicting HER2 status in gastric carcinoma resection specimens. Oncotarget 2015; 6: 38372-80. ArticlePubMedPMC
- 103. Lee HS, Kim WH, Kwak Y, et al. Molecular testing for gastrointestinal cancer. J Pathol Transl Med 2017; 51: 103-21. ArticlePubMedPMCPDF
- 104. Bartley AN, Washington MK, Colasacco C, et al. HER2 testing and clinical decision making in gastroesophageal adenocarcinoma: guideline from the College of American Pathologists, American Society for Clinical Pathology, and the American Society of Clinical Oncology. J Clin Oncol 2017; 35: 446-64. ArticlePubMed
- 105. Wolff AC, Hammond ME, Hicks DG, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol 2013; 31: 3997-4013. PubMed
- 106. Van Cutsem E, Bang YJ, Feng-Yi F, et al. HER2 screening data from ToGA: targeting HER2 in gastric and gastroesophageal junction cancer. Gastric Cancer 2015; 18: 476-84. ArticlePubMedPDF
- 107. Tsapralis D, Panayiotides I, Peros G, Liakakos T, Karamitopoulou E. Human epidermal growth factor receptor-2 gene amplification in gastric cancer using tissue microarray technology. World J Gastroenterol 2012; 18: 150-5. ArticlePubMedPMC
- 108. Liu W, Zhong S, Chen J, Yu Y. HER-2/neu overexpression is an independent prognostic factor for intestinal-type and early-stage gastric cancer patients. Journal of clinical gastroenterology 2012; 46: e31-7. ArticlePubMed
- 109. Lee HE, Park KU, Yoo SB, et al. Clinical significance of intratumoral HER2 heterogeneity in gastric cancer. European Journal of Cancer 2013; 49: 1448-57. ArticlePubMed
- 110. Kim MA, Lee HJ, Yang HK, Bang YJ, Kim WH. Heterogeneous amplification of ERBB2 in primary lesions is responsible for the discordant ERBB2 status of primary and metastatic lesions in gastric carcinoma. Histopathology 2011; 59: 822-31. ArticlePubMedPMCPDF
- 111. Peng Z, Zou J, Zhang X, et al. HER2 discordance between paired primary gastric cancer and metastasis: a meta-analysis. Chin J Cancer Res 2015; 27: 163-71. PubMedPMC
- 112. Kim JH, Kim MA, Lee HS, Kim WH. Comparative analysis of protein expressions in primary and metastatic gastric carcinomas. Hum Pathol 2009; 40: 314-22. ArticlePubMed
- 113. Fusco N, Rocco EG, Del Conte C, et al. HER2 in gastric cancer: a digital image analysis in pre-neoplastic, primary and metastatic lesions. Mod Pathol 2013; 26: 816-24. ArticlePubMedPDF
- 114. Geng Y, Chen X, Qiu J, et al. Human epidermal growth factor receptor-2 expression in primary and metastatic gastric cancer. Int J Clin Oncol 2014; 19: 303-11. ArticlePubMedPDF
- 115. Kochi M, Fujii M, Masuda S, et al. Differing deregulation of HER2 in primary gastric cancer and synchronous related metastatic lymph nodes. Diagn Pathol 2013; 8: 191.ArticlePubMedPMCPDF
- 116. Ieni A, Barresi V, Rigoli L, Caruso RA, Tuccari G. HER2 status in premalignant, early, and advanced neoplastic lesions of the stomach. Dis Markers 2015; 2015: 234851.ArticlePubMedPMCPDF
- 117. Zhao W, Sun L, Dong G, Wang X, Jia Y, Tong Z. Receptor conversion impacts outcomes of different molecular subtypes of primary breast cancer. Ther Adv Med Oncol 2021; 13: 17588359211012982.ArticlePubMedPMCPDF
- 118. Garrido-Ramos MA. Satellite DNA: an evolving topic. Genes (Basel) 2017; 8: 230.ArticlePubMedPMC
- 119. Luchini C, Bibeau F, Ligtenberg MJL, et al. ESMO recommendations on microsatellite instability testing for immunotherapy in cancer, and its relationship with PD-1/PD-L1 expression and tumour mutational burden: a systematic review-based approach. Ann Oncol 2019; 30: 1232-43. ArticlePubMedPDF
- 120. Li K, Luo H, Huang L, Luo H, Zhu X. Microsatellite instability: a review of what the oncologist should know. Cancer Cell Int 2020; 20: 16.ArticlePubMedPMCPDF
- 121. Jiricny J. Postreplicative mismatch repair. Cold Spring Harb Perspect Biol 2013; 5: a012633.ArticlePubMedPMC
- 122. Ma J, Setton J, Lee NY, Riaz N, Powell SN. The therapeutic significance of mutational signatures from DNA repair deficiency in cancer. Nat Commun 2018; 9: 3292.ArticlePubMedPMCPDF
- 123. Peltomaki P. Role of DNA mismatch repair defects in the pathogenesis of human cancer. J Clin Oncol 2003; 21: 1174-9. ArticlePubMed
- 124. Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 2014; 513: 202-9. ArticlePubMedPMCPDF
- 125. Cristescu R, Lee J, Nebozhyn M, et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med 2015; 21: 449-56. ArticlePubMedPDF
- 126. Setia N, Agoston AT, Han HS, et al. A protein and mRNA expression-based classification of gastric cancer. Mod Pathol 2016; 29: 772-84. ArticlePubMedPDF
- 127. Ahn S, Lee SJ, Kim Y, et al. High-throughput protein and mRNA expression-based classification of gastric cancers can identify clinically distinct subtypes, concordant with recent molecular classifications. Am J Surg Pathol 2017; 41: 106-15. ArticlePubMed
- 128. Choi YY, Kim H, Shin SJ, et al. Microsatellite instability and programmed cell death-ligand 1 expression in stage II/III gastric cancer: post hoc analysis of the CLASSIC randomized controlled study. Ann Surg 2019; 270: 309-16. PubMed
- 129. Guastadisegni C, Colafranceschi M, Ottini L, Dogliotti E. Microsatellite instability as a marker of prognosis and response to therapy: a meta-analysis of colorectal cancer survival data. Eur J Cancer 2010; 46: 2788-98. ArticlePubMed
- 130. Kim ST, Cristescu R, Bass AJ, et al. Comprehensive molecular characterization of clinical responses to PD-1 inhibition in metastatic gastric cancer. Nat Med 2018; 24: 1449-58. ArticlePubMedPDF
- 131. Le DT, Uram JN, Wang H, et al. PD-1 blockade in tumors with mismatch-repair deficiency. N Engl J Med 2015; 372: 2509-20. PubMedPMC
- 132. Le DT, Durham JN, Smith KN, et al. Mismatch repair deficiency predicts response of solid tumors to PD-1 blockade. Science 2017; 357: 409-13. PubMedPMC
- 133. Ratti M, Lampis A, Hahne JC, Passalacqua R, Valeri N. Microsatellite instability in gastric cancer: molecular bases, clinical perspectives, and new treatment approaches. Cell Mol Life Sci 2018; 75: 4151-62. ArticlePubMedPMCPDF
- 134. Jang M, Kwon Y, Kim H, et al. Microsatellite instability test using peptide nucleic acid probe-mediated melting point analysis: a comparison study. Bmc Cancer 2018; 18: 1218.ArticlePubMedPMCPDF
- 135. Murphy KM, Zhang S, Geiger T, et al. Comparison of the microsatellite instability analysis system and the Bethesda panel for the determination of microsatellite instability in colorectal cancers. J Mol Diagn 2006; 8: 305-11. ArticlePubMedPMC
- 136. Berg KD, Glaser CL, Thompson RE, Hamilton SR, Griffin CA, Eshleman JR. Detection of microsatellite instability by fluorescence multiplex polymerase chain reaction. J Mol Diagn 2000; 2: 20-8. ArticlePubMedPMC
- 137. Deschoolmeester V, Baay M, Wuyts W, et al. Detection of microsatellite instability in colorectal cancer using an alternative multiplex assay of quasi-monomorphic mononucleotide markers. J Mol Diagn 2008; 10: 154-9. ArticlePubMedPMC
- 138. Shia J, Ellis NA, Klimstra DS. The utility of immunohistochemical detection of DNA mismatch repair gene proteins. Virchows Arch 2004; 445: 431-41. ArticlePubMedPDF
- 139. McCarthy AJ, Capo-Chichi JM, Spence T, et al. Heterogenous loss of mismatch repair (MMR) protein expression: a challenge for immunohistochemical interpretation and microsatellite instability (MSI) evaluation. J Pathol Clin Res 2019; 5: 115-29. ArticlePubMedPDF
- 140. Renkonen E, Zhang Y, Lohi H, et al. Altered expression of MLH1, MSH2, and MSH6 in predisposition to hereditary nonpolyposis colorectal cancer. J Clin Oncol 2003; 21: 3629-37. ArticlePubMed
- 141. Graham RP, Kerr SE, Butz ML, et al. Heterogenous MSH6 loss is a result of microsatellite instability within MSH6 and occurs in sporadic and hereditary colorectal and endometrial carcinomas. Am J Surg Pathol 2015; 39: 1370-6. ArticlePubMed
- 142. Jansson A, Arbman G, Zhang H, Sun XF. Combined deficiency of hMLH1, hMSH2, hMSH3 and hMSH6 is an independent prognostic factor in colorectal cancer. Int J Oncol 2003; 22: 41-9. ArticlePubMed
- 143. Ruszkiewicz A, Bennett G, Moore J, et al. Correlation of mismatch repair genes immunohistochemistry and microsatellite instability status in HNPCC-associated tumours. Pathology 2002; 34: 541-7. ArticlePubMed
- 144. Fukayama M, Abe H, Kunita A, et al. Thirty years of Epstein-Barr virus-associated gastric carcinoma. Virchows Arch 2020; 476: 353-65. ArticlePubMedPDF
- 145. Chang MS, Kim DH, Roh JK, et al. Epstein-Barr virus-encoded BARF1 promotes proliferation of gastric carcinoma cells through regulation of NF-kappaB. J Virol 2013; 87: 10515-23. ArticlePubMedPMCPDF
- 146. Hoebe EK, Le Large TY, Greijer AE, Middeldorp JM. BamHI-A rightward frame 1, an Epstein-Barr virus-encoded oncogene and immune modulator. Rev Med Virol 2013; 23: 367-83. ArticlePubMedPMCPDF
- 147. zur Hausen A, Brink AA, Craanen ME, Middeldorp JM, Meijer CJ, van den Brule AJ. Unique transcription pattern of Epstein-Barr virus (EBV) in EBV-carrying gastric adenocarcinomas: expression of the transforming BARF1 gene. Cancer Res 2000; 60: 2745-8. PubMed
- 148. Kang GH, Lee S, Kim WH, et al. Epstein-barr virus-positive gastric carcinoma demonstrates frequent aberrant methylation of multiple genes and constitutes CpG island methylator phenotypepositive gastric carcinoma. Am J Pathol 2002; 160: 787-94. ArticlePubMedPMC
- 149. Strong MJ, Xu G, Coco J, et al. Differences in gastric carcinoma microenvironment stratify according to EBV infection intensity: implications for possible immune adjuvant therapy. PLoS Pathog 2013; 9: e1003341.ArticlePubMedPMC
- 150. Gu L, Chen M, Guo D, et al. PD-L1 and gastric cancer prognosis: a systematic review and meta-analysis. PLoS One 2017; 12: e0182692.ArticlePubMedPMC
- 151. Park JH, Kim EK, Kim YH, et al. Epstein-Barr virus positivity, not mismatch repair-deficiency, is a favorable risk factor for lymph node metastasis in submucosa-invasive early gastric cancer. Gastric Cancer 2016; 19: 1041-51. ArticlePubMedPDF
- 152. Cheng Y, Zhou X, Xu K, Huang J, Huang Q. Very low risk of lymph node metastasis in Epstein-Barr virus-associated early gastric carcinoma with lymphoid stroma. BMC Gastroenterol 2020; 20: 273.ArticlePubMedPMCPDF
- 153. Yoon CJ, Chang MS, Kim DH, et al. Epstein-Barr virus-encoded miR-BART5-5p upregulates PD-L1 through PIAS3/pSTAT3 modulation, worsening clinical outcomes of PD-L1-positive gastric carcinomas. Gastric Cancer 2020; 23: 780-95. ArticlePubMedPDF
- 154. Lee HS, Chang MS, Yang HK, Lee BL, Kim WH. Epstein-barr virus-positive gastric carcinoma has a distinct protein expression profile in comparison with epstein-barr virus-negative carcinoma. Clin Cancer Res 2004; 10: 1698-705. ArticlePubMedPDF
- 155. Longnecker RM, Kieff E, Cohen JI. Esptein-Barr virus. In: Knipe DM, Howley PM, eds. Fields virology. 6th ed. Philadelphia: Lippincott Williams & Wilkins, 2013; 1898-959.
- 156. Boger C, Kruger S, Behrens HM, et al. Epstein-Barr virus-associated gastric cancer reveals intratumoral heterogeneity of PIK3CA mutations. Ann Oncol 2017; 28: 1005-14. ArticlePubMedPMC
- 157. Ribas A. Tumor immunotherapy directed at PD-1. N Engl J Med 2012; 366: 2517-9. ArticlePubMed
- 158. Kwak Y, Seo AN, Lee HE, Lee HS. Tumor immune response and immunotherapy in gastric cancer. J Pathol Transl Med 2020; 54: 20-33. ArticlePubMedPDF
- 159. Fuchs CS, Doi T, Jang RW, et al. Safety and efficacy of pembrolizumab monotherapy in patients with previously treated advanced gastric and gastroesophageal junction cancer: phase 2 clinical KEYNOTE-059 trial. JAMA Oncol 2018; 4: e180013.ArticlePubMedPMC
- 160. Shitara K, Ozguroglu M, Bang YJ, et al. Pembrolizumab versus paclitaxel for previously treated, advanced gastric or gastro-oesophageal junction cancer (KEYNOTE-061): a randomised, open-label, controlled, phase 3 trial. Lancet 2018; 392: 123-33. PubMed
- 161. Janjigian YY, Shitara K, Moehler M, et al. First-line nivolumab plus chemotherapy versus chemotherapy alone for advanced gastric, gastro-oesophageal junction, and oesophageal adenocarcinoma (CheckMate 649): a randomised, open-label, phase 3 trial. Lancet 2021; 398: 27-40. ArticlePubMedPMC
- 162. DAKO Agilent Technologies. Interpretation manual: gastric or gastroesophageal junction adenocarcinoma. PD-L1 IHC 22C3 pharmDx interpretation manual: gastric or gastroesophageal juction adenocarcinoma. Santa Clara: DAKO Agilent Technologies, 2018.
- 163. Yang JH, Kim H, Roh SY, et al. Discordancy and changes in the pattern of programmed death ligand 1 expression before and after platinum-based chemotherapy in metastatic gastric cancer. Gastric Cancer 2019; 22: 147-54. ArticlePubMedPDF
- 164. Zhou KI, Peterson B, Serritella A, et al. Spatial and temporal heterogeneity of PD-L1 expression and tumor mutational burden in gastroesophageal adenocarcinoma at baseline diagnosis and after chemotherapy. Clin Cancer Res 2020; 26: 6453-63. ArticlePubMedPMCPDF
- 165. Son SM, Woo CG, Kim DH, et al. Distinct tumor immune microenvironments in primary and metastatic lesions in gastric cancer patients. Sci Rep 2020; 10: 14293.ArticlePubMedPMCPDF
- 166. Marchio C, Scaltriti M, Ladanyi M, et al. ESMO recommendations on the standard methods to detect NTRK fusions in daily practice and clinical research. Ann Oncol 2019; 30: 1417-27. ArticlePubMedPDF
- 167. Catenacci DV, Rasco D, Lee J, et al. Phase I escalation and expansion study of bemarituzumab (FPA144) in patients with advanced solid tumors and FGFR2b-selected gastroesophageal adenocarcinoma. J Clin Oncol 2020; 38: 2418-26. ArticlePubMedPMC
- 168. Maron SB, Alpert L, Kwak HA, et al. Targeted therapies for targeted populations: anti-EGFR treatment for EGFR-amplified gastroesophageal adenocarcinoma. Cancer Discov 2018; 8: 696-713. ArticlePubMedPMCPDF
- 169. Lee J, Kim ST, Kim K, et al. Tumor genomic profiling guides patients with metastatic gastric cancer to targeted treatment: the VIKTORY Umbrella Trial. Cancer Discov 2019; 9: 1388-405. ArticlePubMedPDF
- 170. Smyth EC, Cafferkey C, Loehr A, et al. Genomic loss of heterozygosity and survival in the REAL3 trial. Oncotarget 2018; 9: 36654-65. ArticlePubMedPMC
- 171. Nakamura Y, Kawazoe A, Lordick F, Janjigian YY, Shitara K. Biomarker-targeted therapies for advanced-stage gastric and gastro-oesophageal junction cancers: an emerging paradigm. Nat Rev Clin Oncol 2021; 18: 473-87. ArticlePubMedPDF
- 172. van Grieken NC, Aoyama T, Chambers PA, et al. KRAS and BRAF mutations are rare and related to DNA mismatch repair deficiency in gastric cancer from the East and the West: results from a large international multicentre study. Br J Cancer 2013; 108: 1495-501. ArticlePubMedPMCPDF
- 173. Wen Z, Xiong D, Zhang S, et al. Case report: RAB10-ALK: a novel ALK fusion in a patient with gastric cancer. Front Oncol 2021; 11: 645370.ArticlePubMedPMC
- 174. Lee J, Lee SE, Kang SY, et al. Identification of ROS1 rearrangement in gastric adenocarcinoma. Cancer 2013; 119: 1627-35. ArticlePubMed
- 175. Fancello L, Gandini S, Pelicci PG, Mazzarella L. Tumor mutational burden quantification from targeted gene panels: major advancements and challenges. J Immunother Cancer 2019; 7: 183.ArticlePubMedPMCPDF
- 176. Lee KW, Van Cutsem E, Bang YJ, et al. Association of tumor mutational burden with efficacy of pembrolizumab+/-chemotherapy as first-line therapy for gastric cancer in the phase III KEYNOTE-062 study. Clin Cancer Res 2022; 28: 3489-98. ArticlePubMedPDF
- 177. Kang SY, Kim DG, Ahn S, Ha SY, Jang KT, Kim KM. Comparative analysis of microsatellite instability by next-generation sequencing, MSI PCR and MMR immunohistochemistry in 1942 solid cancers. Pathol Res Pract 2022; 233: 153874.ArticlePubMed
- 178. Ratovomanana T, Cohen R, Svrcek M, et al. Performance of nextgeneration sequencing for the detection of microsatellite instability in colorectal cancer with deficient DNA mismatch repair. Gastroenterology 2021; 161: 814-26. ArticlePubMed
- 179. Ascierto PA, Bifulco C, Palmieri G, Peters S, Sidiropoulos N. Preanalytic variables and tissue stewardship for reliable next-generation sequencing (NGS) clinical analysis. J Mol Diagn 2019; 21: 756-67. ArticlePubMed
- 180. Koh J, Lee KW, Nam SK, et al. Development and validation of an easy-to-implement, practical algorithm for the identification of molecular subtypes of gastric cancer: prognostic and therapeutic implications. Oncologist 2019; 24: e1321-30. ArticlePubMedPMCPDF
- 181. Ramos M, Pereira MA, Amorim LC, et al. Gastric cancer molecular classification and adjuvant therapy: is there a different benefit according to the subtype? J Surg Oncol 2020; 121: 804-13. ArticlePubMedPDF
REFERENCES
Figure & Data
References
Citations

- GAST-NET: A multi-modal and multi-task deep learning framework for preoperative prediction of perineural invasion and prognostic risk in gastric cancer
Shidi Miao, Hexiang Dong, Jinyang Feng, Yuyang Jiang, Mengzhuo Sun, Zengyao Liu, Qiujun Wang, Xuemei Ding, Ruitao Wang
International Journal of Medical Informatics.2026; 212: 106348. CrossRef - Poorly cohesive carcinoma diffusely involving the whole gastrointestinal tract: a case report
Wei Gao, Yusheng Yang, Xinyi Hu, Yujuan Shi, Kai Liu, Minmin Gu, Jing Wang
Discover Oncology.2026;[Epub] CrossRef - Comprehensive Overview of Gastric Cancer Immunohistochemistry: Key Biomarkers, Advanced Detection Methods, and Perspectives
Bogdan Oprea
Medicina.2026; 62(4): 683. CrossRef - Tumor Budding in Gastric Carcinoma: Beyond Counting Cells at the Invasive Front—A Review of Current Evidence and Biological Perspectives
Catalin-Bogdan Satala, Gabriela Gurau, Alina-Mihaela Gurau, Gabriela Patrichi, Daniela Mihalache
International Journal of Molecular Sciences.2026; 27(9): 3787. CrossRef - Spatial and Temporal Tumor Heterogeneity in Gastric Cancer: Discordance of Predictive Biomarkers
Hye Seung Lee
Journal of Gastric Cancer.2025; 25(1): 192. CrossRef - PD-L1 as a Biomarker in Gastric Cancer Immunotherapy
Yunjoo Cho, Soomin Ahn, Kyoung-Mee Kim
Journal of Gastric Cancer.2025; 25(1): 177. CrossRef - Korean Gastric Cancer Association-Led Nationwide Survey on Surgically Treated Gastric Cancers in 2023
Dong Jin Kim, Jeong Ho Song, Ji-Hyeon Park, Sojung Kim, Sin Hye Park, Cheol Min Shin, Yoonjin Kwak, Kyunghye Bang, Chung-sik Gong, Sung Eun Oh, Yoo Min Kim, Young Suk Park, Jeesun Kim, Ji Eun Jung, Mi Ran Jung, Bang Wool Eom, Ki Bum Park, Jae Hun Chung, S
Journal of Gastric Cancer.2025; 25(1): 115. CrossRef - A Comprehensive and Comparative Review of Global Gastric Cancer Treatment Guidelines: 2024 Update
Sang Soo Eom, Keun Won Ryu, Hye Sook Han, Seong-Ho Kong
Journal of Gastric Cancer.2025; 25(1): 153. CrossRef - Korea, Japan, Europe, and the United States: Why are guidelines for gastric cancer different?
Emily E. Stroobant, Seong-Ho Kong, Maria Bencivenga, Takahiro Kinoshita, Tae-Han Kim, Takeshi Sano, Giovanni de Manzoni, Han-Kwang Yang, Yuko Kitagawa, Vivian E. Strong
Gastric Cancer.2025; 28(4): 559. CrossRef - Can the Japanese guidelines for endoscopic submucosal dissection be safely applied to Korean gastric cancer patients? A multicenter retrospective study based on the Korean Gastric Cancer Association nationwide survey
Hayemin Lee, Mi Ryeong Park, Junhyun Lee
Annals of Surgical Treatment and Research.2025; 109(2): 81. CrossRef - Double optimal transport for differential gene regulatory network inference with unpaired samples
Mengyu Li, Bencong Zhu, Cheng Meng, Xiaodan Fan, Laura Cantini
Bioinformatics.2025;[Epub] CrossRef - A Randomized Controlled Trial to Evaluate the Effect of Fibrin Glue on Bleeding after Gastric Endoscopic Submucosal Dissection
Tae-Se Kim, Tae-Jun Kim, Yang Won Min, Hyuk Lee, Byung-Hoon Min, Jun Haeng Lee, Poong-Lyul Rhee, Jae J. Kim
Gut and Liver.2025; 19(5): 677. CrossRef - Diagnostic accuracy of stereomicroscopy assessment of invasion depth in ex vivo specimens of early gastric cancer
Jing Wang, Lin Chang, Dong-Feng Niu, Yan Yan, Chang-Qi Cao, Shi-Jie Li, Qi Wu
World Journal of Gastroenterology.2025;[Epub] CrossRef - SMMILe enables accurate spatial quantification in digital pathology using multiple-instance learning
Zeyu Gao, Anyu Mao, Yuxing Dong, Hannah Clayton, Jialun Wu, Jiashuai Liu, ChunBao Wang, Kai He, Tieliang Gong, Chen Li, Mireia Crispin-Ortuzar
Nature Cancer.2025; 6(12): 2025. CrossRef - Genomic and Transcriptomic Characterization of Gastric Cancer with Bone Metastasis
Sujin Oh, Soo Kyung Nam, Keun-Wook Lee, Hye Seung Lee, Yujun Park, Yoonjin Kwak, Kyu Sang Lee, Ji-Won Kim, Jin Won Kim, Minsu Kang, Young Suk Park, Sang-Hoon Ahn, Yun-Suhk Suh, Do Joong Park, Hyung Ho Kim
Cancer Research and Treatment.2024; 56(1): 219. CrossRef - Microscopic tumor mapping of post-neoadjuvant therapy pancreatic cancer specimens to predict post-surgical recurrence: A prospective cohort study
Yeshong Park, Yeon Bi Han, Jinju Kim, MeeYoung Kang, Boram Lee, Eun Sung Ahn, Saemi Han, Haeryoung Kim, Hee-Young Na, Ho-Seong Han, Yoo-Seok Yoon
Pancreatology.2024; 24(4): 562. CrossRef - Effect of Neoadjuvant Chemotherapy on Tumor-Infiltrating Lymphocytes in Resectable Gastric Cancer: Analysis from a Western Academic Center
Elliott J. Yee, Danielle Gilbert, Jeffrey Kaplan, Sachin Wani, Sunnie S. Kim, Martin D. McCarter, Camille L. Stewart
Cancers.2024; 16(7): 1428. CrossRef - Interpretation of PD-L1 expression in gastric cancer: summary of a consensus meeting of Korean gastrointestinal pathologists
Soomin Ahn, Yoonjin Kwak, Gui Young Kwon, Kyoung-Mee Kim, Moonsik Kim, Hyunki Kim, Young Soo Park, Hyeon Jeong Oh, Kyoungyul Lee, Sung Hak Lee, Hye Seung Lee
Journal of Pathology and Translational Medicine.2024; 58(3): 103. CrossRef - Expression of claudin 18.2 in poorly cohesive carcinoma and its association with clinicopathologic parameters in East Asian patients
Moonsik Kim, Byung Woog Kang, Jihyun Park, Jin Ho Baek, Jong Gwang Kim
Pathology - Research and Practice.2024; 263: 155628. CrossRef - Clinicopathological analysis of claudin 18.2 focusing on intratumoral heterogeneity and survival in patients with metastatic or unresectable gastric cancer
T.-Y. Kim, Y. Kwak, S.K. Nam, D. Han, D.-Y. Oh, S.-A. Im, H.S. Lee
ESMO Open.2024; 9(12): 104000. CrossRef - Pathological Interpretation of Gastric Tumors in Endoscopic Submucosal Dissection
Jung Yeon Kim
Journal of Digestive Cancer Research.2023; 11(1): 15. CrossRef - Histopathology of Gastric Cancer
Baek-hui Kim, Sung Hak Lee
The Korean Journal of Helicobacter and Upper Gastrointestinal Research.2023; 23(2): 143. CrossRef - Endoscopic submucosal dissection hands-on training with artificial mucosal layer EndoGEL
Tae-Se Kim, Jun Haeng Lee
Journal of Innovative Medical Technology.2023; 1(1): 5. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure














Fig. 1.
Fig. 2.
Fig. 3.
Fig. 4.
Fig. 5.
Fig. 6.
Fig. 7.
Fig. 8.
Fig. 9.
Fig. 10.
Fig. 11.
Fig. 12.
Fig. 13.
Fig. 14.
| Standard and Conditional data elements | |||
|---|---|---|---|
| Gastrectomy (specimen) type |
|||
| □ Total gastrectomy | |||
| □ Distal (subtotal) gastrectomy | |||
| □ Proximal gastrectomy | |||
| □ Wedge resection | |||
| □ Others ( _______________ ) | |||
| Gross type |
|||
| □ EGC type | |||
| □ EGC type I/IIa/IIb/IIc/III | |||
| □ Mixed EGC type ( ____________ ) | |||
| □ AGC type | |||
| □ Borrmann type 1/2/3/4/unclassifiable | |||
| □ Others ( _______________ ) | |||
| Residual with previous treatment |
|||
| □ Residual | |||
| □ Previous treatment | |||
| □ Chemotherapy | |||
| □ Chemoradiotherapy | |||
| □ Endoscopic mucosal resection | |||
| □ Endoscopic submucosal dissection | |||
| □ Unknown | |||
| □ Others ( ____________ ) | |||
| Tumor focality |
|||
| □ Single | |||
| □ Multiple | |||
| Tumor location |
|||
| □ Involvement | |||
| □ Esophagus/Upper/Middle/Lower third of the stomach/Duodenum | |||
| □ Center | |||
| □ Cardia/Fundus/Body/Antrum/Pylorus | |||
| □ Lesser curvature/Greater curvature/Anterior wall/Posterior wall | |||
| □ Others ( ____________ ) | |||
| Tumor size |
|||
| One largest dimension | |||
| □ ___ cm | |||
| Tumor size |
|||
| Secondary or tertiary tumor dimensions | |||
| □ ___ × ___ cm | |||
| □ ___ × ___ × ___ cm | |||
| Histologic type |
|||
| According to the principles described in “Histologic classification” section | |||
| □ WHO | |||
| □ Lauren | |||
| Tumor regression grade |
|||
| □ Grade 0: Complete response (no viable cancer cells) | |||
| □ Grade 1: Near complete response (single cells or rare small groups of cancer cells) | |||
| □ Grade 2: Partial response (residual cancer with evident tumor regression, but more than single cells or rare small groups of cancer cells) | |||
| □ Grade 3: Poor or no response (extensive residual cancer with no evident tumor regression) | |||
| Lymph node tumor regression |
|||
| □ Not identified | |||
| □ Present | |||
| Depth of invasion (pT) |
|||
| □ Invades lamina propria (pT1a) | |||
| □ Invades muscularis mucosae (pT1a) | |||
| □ Invades submucosa (sm1/sm2/sm3) (pT1b) | |||
| □ Invades proper muscle (pT2) | |||
| □ Invades subserosa (pT3) | |||
| □ Invades serosa (visceral peritoneum) (pT4a) | |||
| □ Directly invades adjacent structure (pT4b) | |||
| specify ( _____________ ) | |||
| Resection margin |
|||
| □ Proximal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Distal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Circumferential resection margin |
|||
| Applied in EGJ or cardia cancer | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Regional lymph node metastasis |
|||
| At least 16 regional lymph nodes should be assessed | |||
| □ no metastasis in ____ regional lymph nodes | |||
| □ metastasis in ___ out of ___ regional lymph nodes | |||
| Extranodal tumor extension |
|||
| □ Not identified | |||
| □ Present | |||
| Isolated tumor cell clusters |
|||
| Applied in incidentally identified tumor cell cluster less than 0.2 mm in greatest dimension with no other regional lymph node metastasis (pN0) | |||
| □ Present [pN0 (i+)] | |||
| Lymphovascular invasion |
|||
| □ Not identified | |||
| □ Present | |||
| Venous invasion |
|||
| Applied when identified in large vessels with an identifiable smooth muscle layer or elastic lamina | |||
| □ Not identified | |||
| □ Present | |||
| Perineural invasion |
|||
| □ Not identified | |||
| □ Present | |||
| Pre-existing adenoma |
|||
| Used if the carcinoma is within the adenoma | |||
| □ Tubular/Tubulovillous/Villous adenoma | |||
| □ Low grade dysplasia/High grade dysplasia | |||
| Associated findings |
|||
| □ Tumor perforation | |||
| □ Serosal (peritoneal, mesenteric) seeding | |||
| □ Distant metastasis | |||
| Other organ, specify: ______________ | |||
| Distant lymph node | |||
| Separate lesions |
|||
| □ Peptic ulcer | |||
| □ Adenoma | |||
| □ GIST | |||
| □ Others ( ____________ ) | |||
| Standard and Conditional data elements | |||
|---|---|---|---|
| Specimen size |
|||
| □ ___ × ___ cm | |||
| Gross type of tumor |
|||
| Same as method of surgical specimen | |||
| Tumor size |
|||
| One largest dimension | |||
| □ _____ cm | |||
| Histologic type |
|||
| According to the principles described in “Histologic classification” section | |||
| □ WHO | |||
| □ Lauren | |||
| Histologic components |
|||
| All morphologic components of tumor cell may be described | |||
| Depth of invasion (pT) |
|||
| □ Invades lamina propria (pT1a) | |||
| □ Invades muscularis mucosae (pT1a) | |||
| □ Invades submucosa (submucosal depth: _____ mm or μm) | |||
| □ Invades proper muscle (pT2) | |||
| Depth of invasion (pT) |
|||
| In case of submucosa invasion, the invasion width can be additionally described | |||
| □ invades submucosa (submucosal depth: _____ mm or μm) (submucosal width: _____ mm) | |||
| Resection margin |
|||
| □ Lateral margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Deep margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Resection margin |
|||
| □ Proximal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Distal margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Anterior margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Posterior margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| □ Deep margin | |||
| □ Free from carcinoma (safety margin, ___ cm) | |||
| □ Involved by carcinoma | |||
| Ulceration |
|||
| □ Absent | |||
| □ Present | |||
| Ulceration |
|||
| □ Absent | |||
| □ Non-significant (≤ 4 mm) | |||
| □ Significant (> 4 mm) | |||
| Cases with adenoma components |
|||
| □ Absent | |||
| □ Present | |||
| specify: ______________ | |||
| En bloc resection |
|||
| □ Yes | |||
| □ No (piecemeal/tearing) | |||
| Lymphatic invasion |
|||
| □ Not identified | |||
| □ Present | |||
| Venous invasion |
|||
| □ Not identified | |||
| □ Present | |||
| Histopathologic classification | ||
|---|---|---|
| WHO classification | ||
| □ Tubular adenocarcinoma | ||
| □ Tubular adenocarcinoma, well differentiated | ||
| □ Tubular adenocarcinoma, moderately differentiated | ||
| □ Tubular adenocarcinoma, poorly differentiated | ||
| □ Papillary adenocarcinoma | ||
| □ Mucinous adenocarcinoma | ||
| □ Poorly cohesive carcinoma | ||
| □ Poorly cohesive carcinoma, signet-ring cell type | ||
| □ Poorly cohesive carcinoma, not otherwise specified | ||
| □ Mixed adenocarcinoma | ||
| □ Adenocarcinoma with lymphoid stroma | ||
| □ Hepatoid adenocarcinoma | ||
| □ Micropapillary adenocarcinoma | ||
| □ Adenocarcinoma of fundic-gland type | ||
| □ Undifferentiated carcinoma | ||
| □ Squamous cell carcinoma | ||
| □ Adenosquamous carcinoma | ||
| □ Gastroblastoma | ||
| □ Others (specify: ______________) | ||
| Lauren classification | ||
| □ Intestinal | ||
| □ Diffuse | ||
| □ Indeterminate | ||
| □ Mixed | ||
| Molecular markers | ||
|---|---|---|
| All molecular markers are “conditional data element” | ||
| HER2 immunohistochemistry | ||
| □ Negative (0/1+) | ||
| □ Equivocal (2+) | ||
| □ Positive (3+) | ||
| □ Undetermined (explain): | ||
| HER2 (ERBB2) in situ hybridization | ||
| Number of invasive cancer cells counted: ______ cells | ||
| □ Using dual-probe assay | ||
| □ HER2 (ERBB2)/CEP17 ratio: ______ | ||
| □ Average number of HER2 (ERBB2) signals per cancer cell: ______ | ||
| □ Average number of CEP17 signals per cancer cell: ______ | ||
| □ Using single-probe assay | ||
| □ Average number of HER2 (ERBB2) signals per cancer cell: ______ | ||
| Summary: Negative/Positive for HER2 (ERBB2) gene amplification | ||
| □ Undetermined (explain): | ||
| Microsatellite instability (MSI) | ||
| Summary: | ||
| □ Microsatellite stable (MSS) | ||
| □ Microsatellite instability-low (MSI-L) | ||
| □ Microsatellite instability-high (MSI-H) | ||
| □ Undetermined (explain) |
||
| DNA mismatch repair immunohistochemistry | ||
| MLH1: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| MSH2: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| PMS2: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| MSH6: | ||
| □ Positive (retained expression) | ||
| □ Negative (loss of expression) | ||
| □ Undetermined (explain): | ||
| Summary: | ||
| □ DNA mismatch repair deficiency (was/was not) observed | ||
| □ Because it is difficult to determine DNA mismatch repair deficiency, PCR-based testing and/or NGS for MSI is recommended. | ||
| In situ hybridization for Epstein-Barr virus–encoded small RNAs | ||
| □ Positive [diffuse/heterogenous (focal and/or mixed intensity)] |
||
| □ Negative | ||
| Summary: Epstein-Barr virus–associated gastric carcinoma | ||
| PD-L1 immunohistochemistry | ||
| PD-L1 [Antibody (22C3 PharmDx/22C3 conc. Ventana/28-8 PharmDx/others:______)]: | ||
| □ CPS = ________ | ||
EGC, early gastric cancer; AGC, advanced gastric cancer; WHO, World Health Organization; EGJ, esophagogastric junction. Standard data elements; Conditional data elements.
WHO, World Health Organization. Standard data elements; Conditional data elements.
WHO, World Health Organization.
HER2, human epidermal growth factor receptor 2; CEP17, centromeric region of chromosome 17; MLH1, mutL homolog 1; MSH2, mutS homolog 2; PMS2, PMS1 homolog 2; MSH6, mutS homolog 6; PCR, polymerase chain reaction; NGS, next-generation sequencing; PD-L1, programmed death ligand 1; CPS, combined positive score. Because it is difficult to determine MSI status, mismatch repair immunohistochemistry and/or NGS is recommended; Checking the signal pattern is optional; The term “Epstein-Barr virus–associated gastric carcinoma” applies to positive cases.

 E-submission
E-submission