Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 57(3); 2023 > Article
-
Original Article
Expression of specific microRNAs in tissue and plasma in colorectal cancer -
Allan Fellizar1,2
 , Vivencio Refuerzo3
, Vivencio Refuerzo3 , John Donnie Ramos1,4,5
, John Donnie Ramos1,4,5 , Pia Marie Albano1,4,5
, Pia Marie Albano1,4,5
-
Journal of Pathology and Translational Medicine 2022;57(3):147-157.
DOI: https://doi.org/10.4132/jptm.2022.02.19
Published online: May 3, 2022
1The Graduate School, University of Santo Tomas, Manila, Philippines
2Department of Pathology and Laboratories, Mariano Marcos Memorial Hospital and Medical Center, Batac, Philippines
3Department of Surgery, Mariano Marcos Memorial Hospital and Medical Center, Batac, Philippines
4Research Center for Natural and Applied Sciences, University of Santo Tomas, Manila, Philippines
5Department of Biological Sciences, College of Science, University of Santo Tomas, Manila, Philippines
- Corresponding Author: Allan L. Fellizar, RMT, MSMT, Department of Pathology and Laboratories, Molecular Biology Laboratory, Mariano Marcos Memorial Hospital and Medical Center, City of Batac, 2906, Ilocos Norte, Philippines, Tel: +63-917-8734007, Fax: +63-77-600-8000, E-mail: allan.fellizar.gs@ust.edu.ph
© 2023 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (https://creativecommons.org/licenses/by-nc/4.0) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- MicroRNAs (miRNA/miR) play significant roles in the regulation of cell differentiation, cell cycle progression, and apoptosis. They become dysregulated during carcinogenesis and are eventually released into the circulation, enabling their detection in body fluids. Thus, this study compared the miRNA expression in tissue and plasma samples of colorectal cancer (CRC) patients and clinically healthy controls and determined miRNA expression as a potential CRC biomarker.
-
Methods
- Using quantitative reverse transcription polymerase chain reaction (RT-qPCR), miR-21-5p, miR-29a-3p, miR-92a-3p, miR-135b-5p, miR-196b-5p, and miR-197-3p, expression was analyzed and compared between the malignant (n = 41) and the adjacent neoplasm free mucosal tissues (n = 41) of CRC patients. The findings were validated in plasma samples (n = 36) collected from the same CRC patients prior to surgery or any form of treatment and compared to plasma from their age and sex-matched controls (n = 36).
-
Results
- MiR-21-5p, miR-29a-3p, miR-92a-3p, and miR-196b-5p were upregulated and miR-135b-5p was downregulated in CRC malignant tissues compared to their expression in adjacent neoplasm-free tissue. This was further observed in the plasma of the same CRC cases compared to controls. MiR-92a-3p showed itself the most sensitive (0.93; p < .001) and most specific (0.95; p < .001) in detecting CRC in tissue. In plasma, miR-196b-5p was the most sensitive (0.97; p < .001) and specific (0.94; p < .001) in detecting CRC. Plasma miR-92a-3p and miR-196b-5p were the most sensitive (0.95; p < .001) and specific (0.94; p < .001) in the early detection of CRC.
-
Conclusions
- Results show that specific miRNAs dysregulated in malignant tissues are released and can be detected in the circulation, supporting their potential as non-invasive biomarkers of CRC.
- Study participants and samples
- Formalin fixed paraffin embedded (FFPE) tissue samples surgically removed from patients with histologically confirmed CRC and seen at the Mariano Marcos Memorial Hospital and Medical Center (MMMH-MC) in Ilocos Norte, Philippines from March 2018 to December 2018 were included in this study. Each FFPE tissue block positive for malignant cells was matched with neoplasm-free mucosal tissue removed from the same patient and designated as case and control, respectively. Successive 5-μm-thick tissue samples were sectioned using a microtome (Leica Biosystems, Wetzlar, Germany), with the outer sections mounted on glass slides and then stained with hematoxylin and eosin (H&E). The inner sections (approximately 5 mg) were aseptically collected in nuclease-free microcentrifuge tube. The H&E-stained slides were evaluated by a pathologist to ensure that the tissue samples designated as cases and controls were strictly positive and negative for cancer cells, respectively.
- Blood samples were also collected from the same CRC patients prior to their surgery or any form of treatment (cases). These were matched with blood samples from clinically healthy volunteers (controls) of the same age (± 2 years) and sex as the CRC patients. Controls were strictly not suspected of any type of malignancy at the time of physical or clinical assessment by a physician, had not undergone any colorectal resection except for sigmoid diverticular disease and had not been diagnosed with inflammatory bowel disease (IBD), chronic ulcerative colitis, or Crohn’s disease. All blood samples were collected in K2EDTA tubes (Becton Dickinson, Franklin Lakes, NJ, USA) and centrifuged immediately at 2,000 × g for 10 minutes. at 4°C. Hemolyzed blood samples and turbid plasma were excluded from the study. Plasma was separated and aliquoted into nuclease-free cryovials and then stored in a freezer at −80°C until analysis.
- RNA isolation from FFPE tissues and plasma samples
- Total RNA including miRNAs were isolated from FFPE tissues utilizing miRNeasy FFPE kit according to manufacturer’s instructions (miRNeasy FFPE Handbook June 2015, http://www.qiagen.com). FFPE tissue sections were deparaffinized and then digested by proteinase K, followed by heat treatment. After centrifugation, the supernatant was collected and treated with DNAse. After mixing with ethanol and buffer, the lysate was transferred into a RNeasy MinElute spin column where total RNA are bound. After washing twice, the RNA was eluted.
- Plasma miRNA was isolated using the miRNeasy serum/plasma kit total RNA (Qiagen, Hilden, Germany) according to manufacturer’s instructions. QIAzol was added to 100 μL of plasma, incubated, and miRNeasy serum/plasma spike-in control Caenorhabditis elegans miR-39 miRNA mimic (1 × 108 copies/μL) (Qiagen) was added to each sample. Aqueous and organic phase separation was achieved using chloroform. The aqueous phase was extracted after centrifugation and addition of absolute ethyl alcohol. The mixture was transferred to RNEasy mini spin columns and centrifuged. Buffers RWT and RPE were used respectively on two consecutive steps to wash the spin columns with centrifugation. The total RNA including miRNA was eluted using RNAse-free water applied directly at the center of the mini spin column silica membrane and centrifuged at full speed. The RNA eluates from tissue and plasma were stored at −20°C until subsequent analysis.
- Reverse transcription of miRNA and preamplification of cDNA
- MiRNAs from colorectal tissues and plasma were polyadenylated and reverse transcribed using the miScript II RT kit and miScript HiSpec buffer (Qiagen) following the manufacturer’s procedure. RNAse-free water (40 μL) was added to the synthesized cDNA (10 μL), and the mixture was aliquoted into PCR tubes and stored at −20°C until analysis.
- MiScript PreAMP PCR Kit (Qiagen) was used to pre-amplify the cDNA target templates following manufacturer instructions. MiR-16 was analyzed to determine the optimal dilution for real-time quantitative reverse transcription polymerase chain reaction (RT-qPCR). Samples that generated Ct values between 10 and 24 needed no further dilution for target miRNAs using RT-qPCR. Efficiency of reverse transcription was measured using miRTC assay. Efficient reverse transcription for the miRTC primer assay was set at Ct values between 14 and 20.
- Quantification of selected miRNAs by qRT-PCR
- SNORD61 and C. elegans miR-39 primers for normalization of tissue and plasma miRNA, respectively, and 10× miScript primer assay mixes for hsa-miR-21-5p, hsa-miR-196b-5p, hsa-miR-135b-5p, hsa-miR-92a-3p, hsa-29a-3p, and hsa197-3p were ordered from Qiagen. Two microliters of pre-amplified cDNA was pipetted into a 96-well PCR plate, and 23 μL of reaction mix (12.5 μL 2 × QuantiTect SYBR Green PCR master mix, 2.5 μL 10 × miScript universal primer, 2.5 μL 10 × miScript primer assay mix, and 5.5 μL RNAse-free water) was added. Amplification was performed under the following conditions: initial activation step at 95°C for 15 minutes, denaturation at 94°C for 15 seconds, annealing at 55°C for 30 seconds, and extension at 70°C for 30 seconds for 40 cycles. All assays were performed in duplicate.
- Data processing and analysis
- The relative expression levels of the miRNAs interrogated were normalized to those of SNORD61 and cel-miR-39 and determined by the 2−ΔΔCt method. Using GraphPad Prism 8.3 for Windows (GraphPad Software, San Diego, CA, USA, http://www.graphpad.com) and XLSTAT 2019.3.2 (Addinsoft, Paris, France), intergroup comparisons were analyzed by paired t tests and Mann-Whitney tests. Spearman correlation coefficient was used for correlation analysis. A p-value less than .05 (two-tailed) was considered statistically significant. The receiver operating characteristic (ROC) and the area under the ROC curve (AUC) with 95% confidence interval (CI) were generated, and the number of outliers per tissue and plasma group were determined by Robust Regression and Outlier Removal (ROUT) [23]. These discriminating performance calculations were carried out using GraphPad Prism 8.3 (GraphPad Software Inc., San Diego, CA, USA).
MATERIALS AND METHODS
- Clinicopathological characteristics of CRC patients
- A total of 41 FFPE malignant tissues and their corresponding adjacent neoplasm-free mucosal tissues (n = 41) were collected from histologically confirmed Filipino CRC patients. EDTA-treated plasma samples from the same CRC patients (n = 36) were collected prior to surgery or any form of treatment and from their age and sex-matched clinically healthy controls (n = 36). Table 1 shows the clinicopathologic characteristic of cases included in this study.
- Expression patterns of selected miRNAs in CRC tissue and plasma
- Table 2 shows the relative expression and fold regulation of the miRNAs in malignant colorectal tissue. MiR-21-5p, miR-29a-3p, miR-92a-3p, and miR-196b-5p were upregulated in CRC tissues in relation to their adjacent neoplasm-free tissue. MiR-92a-3p showed the highest fold change (FC = 7.41, p < .001) in CRC tissue, followed by miR-21-5p (FC = 4.23, p < .001) and miR-29a-3p (FC = 3.56, p < .001). MiR-135b-5p was downregulated (FC = 0.17, p < .001) in CRC tissue compared to its adjacent neoplasm-free tissue. The relative expression of miR-197-3p in CRC tissue was not significantly different (FC = 0.78, p = .676) in relation to its adjacent neoplasm-free tissue.
- To determine whether the dysregulated miRNAs in CRC tissue were released into the circulation, plasma from the same CRC patients was collected and tested for the specific miRNA under study prior to any form of treatment. Interestingly, expression patterns of miR-21-5p, miR-29a-3p, miR-92a-3p, miR-135b-5p, miR-196b-5p, and miR-197-3p in plasma of CRC cases were similar to plasma from clinically healthy individuals (Table 2). MiR-21-5p showed the highest expression (FC = 8.40, p < .001) in CRC plasma, followed by miR-196b-5p (FC = 4.75, p < .001) and miR-29a-3p (FC = 2.89, p < .001). Similar to CRC tissue, miR-135b-5p was downregulated (FC = 0.44, p < .001) in CRC plasma compared to clinically healthy controls. The relative expression of miR-197-3p in CRC plasma was not significantly different (FC = 1.32, p = .112) in relation to control plasma.
- To determine if the significant differences in miRNA expression levels in tissue and plasma were attributable to the presence of outliers, ROUT was performed. The residuals of the robust fit were analyzed to identify any outlier according to the false discovery rate approach for testing multiple comparisons. The ROUT coefficient was set at Q=1%. Table 2 shows that, upon outlier removal, the p-values between malignant and adjacent neoplasm-free tissue and CRC and clinically health control plasma remained significantly different.
- Ability of selected miRNAs to discriminate malignant from neoplasm-free mucosal tissue and plasma from CRC patients from clinically healthy controls
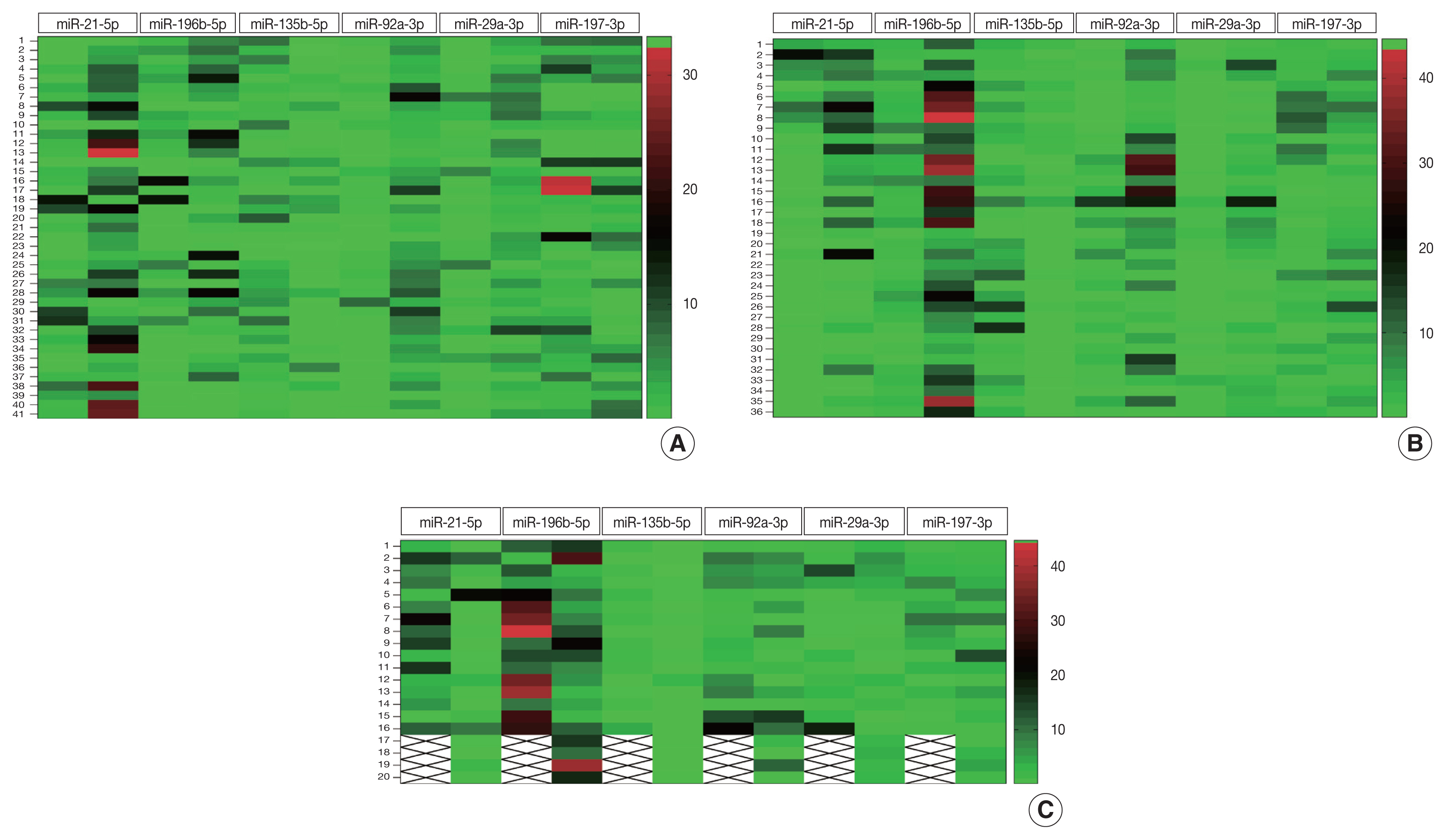
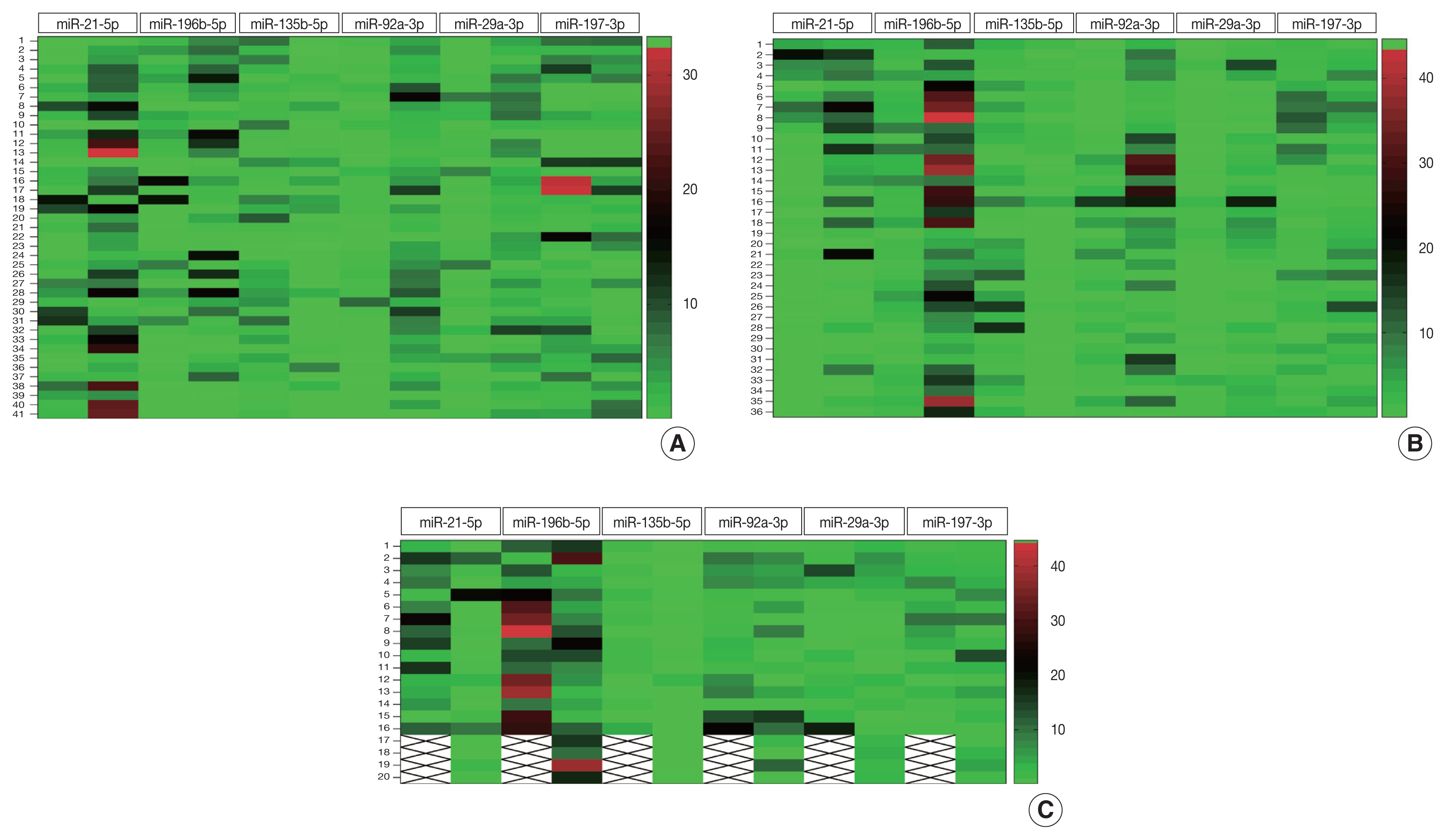
- To determine discriminatory ability of miRNAs under study, heatmap analysis was performed. Fig. 1A shows that majority of the CRC tissues exhibited greater color intensities than normal adjacent tissues with respect to miR-21-5p, miR-196b-5p, miR-92a-3p, and miR-29a-3p expression. For miR-135b-5p, the color intensity appeared lighter in CRC compared to normal adjacent tissue.
- Heatmap analysis (Fig. 1B) of plasma samples show that color intensity for miR-21-5p, miR-196b-5p, miR-92a-3p, and miR-29a-3p of CRC was stronger than color intensity for control plasma. For miR-135b-5p, the color intensity appeared lighter in CRC plasma than in control plasma. This color intensity pattern exhibited in CRC plasma is similar to that shown by malignant tissue.
- Relative expression of selected miRNAs in early and advanced stage CRC
- Relative expressions of miR-21-5p (p = .531), miR-29a-3p (p = .473), miR-92a-3p (p = .578), miR-135b-5p (p = .611), miR-196b-5p (p = .769), and miR-197-3p (p = .879) in tissues of patients with early-stage CRC were not significantly different from those in tissues of patients with advanced stage CRC (Table 3). Conversely, levels of all miRNAs studied were useful to discriminate between early and advanced stage CRC using plasma. The relative plasma expressions of miR-21-5p (p = .006), miR-29a-3p (p < .001), miR-92a-3p (p < .001), and miR-196b-5p (p < .001) were significantly higher in early-stage CRC compared to advanced-stage CRC (Table 3). In contrast, significantly higher expression of miR-135b-5p (p < .001) and miR-197-3p (p = .007) was observed in plasma samples of patients with advanced CRC compared to early stage. MiR-197-3p, which failed to discriminate CRC plasma from clinically healthy plasma, was able to distinguish between early and advanced stages of CRC (p = .007). Heatmap analysis (Fig. 1C) showed the potential of specific miRNAs in discriminating early (I/II) from advanced (III/IV) stage CRC. The mean color intensities of miR-21-5p, miR-196b-5p, miR-92a-3p, and miR-29-3p are greater in the early stages compared to late stages of CRC. MiR-135b-5p and miR-197-3p have a lower mean color intensity in early stages compared to late stages of CRC.
- Correlation of expression levels of miRNAs in tissues and plasma
- Results show that expression levels of miR-196b-5p (r = 0.02, p = .921), miR-135b-5p (r = 0.14, p = .430) and miR-197-3p (r = −0.24, p = .150) in tissue and plasma were not correlated. A weak positive correlation was observed with miR-21-5p (r = 0.33, p = .049) while miR-92a-3p (r = 0.54, p = .007) and miR-29-3p (r = 0.52, p = .048) expressions in CRC tissues and plasma presented a moderate correlation (Table 4).
- Diagnostic potential of selected miRNAs in detection of CRC in tissue and plasma
- AUC values of the ROC were computed for each miRNA to evaluate their diagnostic value in detection of CRC in tissues and plasma. In CRC tissue, miR-92a-3p obtained the highest AUC (0.89; 95% CI, 0.82 to 0.97), with a sensitivity of 0.93 and specificity of 0.95 (Table 5). In detecting malignancy using plasma samples, MiR-196b-5p showed the highest AUC (0.94; 95% CI, 0.89 to 0.99) with a sensitivity of 0.97 and specificity of 0.94 (Table 5). In early detection of CRC using plasma samples, miR-92a-3p showed the highest AUC (0.95; 95% CI, 0.88 to 0.99), with a sensitivity of 0.95 and specificity of 0.94 (Table 5).
RESULTS
- The identification of diagnostic molecular biomarkers is substantial in cancer research. Increasing evidence suggests that miRNAs play a vital role in regulating the development, differentiation, and progression of cancer. MiRNAs are well preserved in tissue even after formalin-fixation and paraffin-embedding. They can be efficiently extracted and assessed from tumors and body fluids such as serum, plasma, urine, stool, and saliva. The miRNAs analyzed in the current study have been identified in previous independent studies utilizing diverse study subjects (different races and populations) through sequencing and PCR techniques as candidate biomarkers for CRC. The differences and similarities in the expression patterns of these miRNAs in tissue and plasma across different study populations have been highlighted. Similarly, the current study determined the pattern of these miRNAs, thereby contributing to the current understanding of which specific signature miRNAs can be applied in the clinical setting.
- The current study shows that miR-21-5p, miR-29a-3p, miR-92a-3p, and miR-196b-5p were significantly overexpressed while miR-135b-5p was underexpressed in malignant compared to neoplasm-free colorectal tissues. The above specific miRNAs were also dysregulated in the plasma of CRC patients but not in their matched clinically healthy controls. However, miR-197-3p expression in tumor tissue and plasma of cancer patients was not significantly different from that in controls.
- MiR-21-5p plays an oncogenic role in the development and progression of CRC by modulating malignant processes such as proliferation, anti-apoptosis, cell cycle progression, and invasion in CRC cells through downregulation of PTEN protein expression [24]. Prostaglandin-endoperoxide synthase 2 (PGTS2) produces inflammatory mediator prostaglandin E2 (PGE2), which has been described as promoting colorectal tumor development. PGTS2-driven inflammatory responses induce tumor expression of miR-21-5p that can elevate PGE level by downregulating PGE2-metabolizing enzymes [25]. During CRC development, miR-21 also induces stemness by downregulating TGFβR2 (transforming growth factor beta receptor 2) and stimulates invasion and metastasis by suppressing the PDCD4 gene [26]. MiR-21 expression has been found to be upregulated in breast, lung, and gastric cancers including CRC and hematological malignancies [27]. This study agrees with previous findings that miR-21-5p is upregulated in tissues [27–29] and plasma or serum [29–32]. Mir-21-5p expression was 12 and 10 times higher in serum and stool, respectively, of CRC patients compared to those of healthy controls [31]. Meanwhile, Stiegelbauer et al. [32] observed a serological underexpression of miR-21-5p in CRC. Plasma miR-21-5p downregulation was also detected in invasive breast cancer [33].
- It has been observed that miR-196b-5p is upregulated in acute lymphoblastic leukemia but downregulated in glioblastoma, cervical cancer, and B-cell lymphoma. This miRNA has been experimentally validated to regulate CRC cell migration and metastasis through interaction with GALNT5 and HOXB7 genes [34]. Although they vary in expression levels, the miR-196 family of molecules is consistently overexpressed in oral cavity, esophageal, stomach, and intestinal cancer tissues [35]. Several investigations [36–38] support our findings that miR-196b expression levels in malignant tissues are significantly higher than in adjacent normal colorectal mucosa. MiR-196b-5p was also found to be elevated in serum exosomes of CRC and is associated with liver metastasis [35]. MiR-196b-5p is also more upregulated in serum of patients with CRC compared to that in adenoma patients or healthy individuals [36]. A few reports have shown that the miR-196 family can also act as tumor suppressors. For instance, miR-196a suppresses metastasis in breast cancers [37] and melanoma [38]. MiR-196-b has been found to be downregulated in different types of leukemia cells [39].
- Upregulation of miR-29a in tissue and plasma of CRC patients was noted in the present study. Similarly, Brunet Vega et al. [40] noted that miR-29a along with 10 other miRNAs were significantly increased in malignant colorectal tissue samples compared with non-cancerous adjacent mucosa. In the same study, serum level of miR-29a was overexpressed in CRC patients but not in healthy controls. In a genome-wide miRNA profiling conducted by Giraldez et al. [41], in 63 plasma samples from newly diagnosed CRC patients, miR-29a was confirmed to be upregulated compared with controls. In contrast, downregulation of miR-29a has been observed in human lung cancer tissues. Liu et al. [42] demonstrated that expression level of miR-29a was significantly downregulated in 38 pairs of lung cancer tissues compared to adjacent normal tissue. MiR-29a-3p promotes CRC metastasis by regulating MMP-2 gene and E-cadherin via the KLF-4 signaling pathway [43]. It has also been found to promote cell proliferation and epithelial-mesenchymal transition in breast cancer by targeting TET-1 [44]. Conversely, miR-29a functions as a tumor suppressor by targeting the MUC-1 in pancreatic cancer cell [45].
- The miR-92 family, one of the four families that belong to the miR-17-92 cluster, has been shown to regulate formation of vascular endothelial cells and blood vessels. Aberrant expression of the miR-92a family has been observed in breast, lung, gastric, prostate, and pancreatic cancers [46]. MiR-92a-3p is implicated as a key oncogenic component in the miR-17-92 cluster during colorectal tumorigenesis [47]. Mir-92a promotes metastasis by suppressing PTEN gene expression and activation of the PI3K/AKT pathway [48]. The current study showed miR-92a-3p to be overexpressed in malignant tissue and plasma of CRC patients. Our study revealed that miR-92a-3p obtained the highest sensitivity and specificity in detecting CRC in tissue. Tsuchida et al. [47] found in their study that miR-92 was more significantly upregulated in both colorectal adenoma and carcinoma compared to the other five miRNAs in the cluster. Along with miR-21 and miR-29a, miR-92a has been observed to be significantly upregulated in CRC tissues compared to normal colorectal mucosal tissue. Moreover, this study agrees with the results of Ng et al. [9] that miR-92a was upregulated in both plasma and tissue samples of CRC patients in comparison with healthy controls.
- MiR-135b-5p, an oncogene, has a tumor-promoting effect by enabling proliferation and inhibiting apoptosis of CRC cells through negative regulation of the TGF-β signaling pathway [49]. Magalhaes et al. [50] experimentally validated that downregulation of the APC gene caused by miR-135b-5p led to higher transduction signaling within the β-catenin/Wnt pathway, causing a greater proliferative capacity of malignant cells. APC modulates β-catenin as an essential part of the multi-protein complex that marks it for proteasomal degradation. MiR-135b-5p expression in CRC tissue and plasma was downregulated in this study. Zekri et al. [51] obtained similar findings, wherein serum miR-135b-5p and miR-454 were the only downregulated miRNAs in CRC and colonic polyp groups compared to the IBD group. In contrast, Bastaminejad et al. [52] demonstrated that expression levels of miR-135b-5p in serum and stool of CRC patients were 32 and 16 times higher compared to healthy controls, respectively. While miR-135b-5p is known to be an oncogenic miRNA, it might have different binding affinities due to its numerous potential target genes [53].
- Whether the miRNAs detected in circulation are products of tumor apoptosis or are secreted by the active tumor cell itself remains controversial. MicroRNAs released in the circulation are usually bound to Argonaute proteins, microvesicles, exosomes, or lipoproteins that offer different degrees of protection. Hence, circulating miRNAs exhibit half-lives ranging from minutes to hours, which might explain why their concentrations in circulation vary and why their expressions in the tumor and plasma of the same patient might not be correlated [54]. MiR-1224, for instance, is the most upregulated miRNA in CRC plasma samples, but its expression in its paired tumor sample is rather low. Several explanations have been offered regarding the non-correlation between tumor and plasma miRNA expression levels. Exosomal shifting between the tumor mass and fluid microenvironment is one possibility [55].
- Cancer-associated circulating miRNAs might have originated from immunocytes in the tumor microenvironment or from some other response mediated by affected organs or system [56,57]. Investigators suggest that tumor cells secrete a variety of miRNAs that act on immunocytes to modulate immune responses. In response, the immunocytes secrete miRNAs that either promote or inhibit tumor proliferation, migration, and apoptosis [58]. Lastly, technical issues such as sample type, biological and racial differences, and analytical and normalization methods can significantly contribute to the lack of or weak relational expression of specific microRNAs in tissue and plasma. Researchers found that serum samples can yield lower miRNA concentrations compared to plasma primarily due to the presence of cellular contaminants particularly from platelets.
- CRC has been considered an extremely heterogenous and dynamic disease characterized by multiple molecular pathways throughout its development. This can be attributed to the existence of cellular subpopulations between and within tumors of divergent genotype and phenotype expressions [59]. Chromosomal instability, microsatellite instability, aberrant DNA methylation, and DNA repair mechanisms are intricate processes involved in colorectal epithelial cell transformation and all confer on a tumor its distinctiveness [60]. This high degree of genomic diversity in CRC over time, including relationships between subpopulations within and between the tumor, inaccurately reflects the true molecular profile within the tumor tissue sample, forming a major obstacle in clinical practice [61]. Further, this tumor heterogeneity can introduce sampling bias, heightening the intricacies in validation of oncology biomarkers [62]. Molecular diagnostic studies that seek to recognize driver mutations within a tumor can be affected by the presence of multiple mutations within the same sample. Thus, it is imperative to prudently characterize tumor samples for research to avoid inexplicable results or outcomes.
- There are certain limitations that need to be considered when interpreting the findings of this study. First is the small sample size in the context of miRNA-based biomarker identification and validation. Due to limited budget, the current study was only able to analyze a limited number of samples. Thus, a follow-up study to validate the current findings in a prospective set of samples in a blinded manner is recommended. Moreover, the research team has, in a separate study, applied artificial intelligence (AI) in the analysis of the data, wherein a set of samples has been used as a training set to create the AI models, and the remaining samples were used as test specimens to determine the diagnostic potential of the miRNAs. Second, this study was a single-institution study focused on a specific population in the Philippines, whose diet and lifestyle might not represent those of other ethnic regions of the country. Recruitment of CRC patients and controls from different parts of the country is suggested to determine reproducibility of findings.
- In conclusion, this study presented a proof of concept that dysregulated miRNAs in CRC tissues are observed in the circulation, supporting their potential as non-invasive biomarkers for early detection of CRC. Results of this study agree with those of previous studies using samples from other populations. However, clinical translation of miRNA as a non-invasive circulating biomarker requires highly sensitive, specific, reproducible, reliable, and robust assays to enable its accurate and precise quantification in tissue, plasma, serum, and other human body fluids. Critical pre-analytical and analytical variables such as individual biological variability, sample type, sample storage, miRNA extraction and purification, detection method, and normalization all need to be optimized.
DISCUSSION
Acknowledgments
Ethics Statement
All procedures were performed in accordance with the ethical standards of institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards. This study complied with all applicable ethical guidelines approved by the Research Ethics Review Committee of the Mariano Marcos Memorial Hospital and Medical Center (MMMH-RERC-18-005, March 2, 2018). Study participants gave their written informed consent and were assured of the confidentiality of the results.
Availability of Data and Material
The datasets generated or analyzed during the study are available from the corresponding author on reasonable request.
Code Availability
Not applicable.
Author Contributions
Conceptualization: AF, PMA. Data Curation: AF, VR, PMA. Formal Analysis: AF, PMA. Funding Acquisition: PMA, AF. Investigation: AF, VR, PMA. Methodology: AF, VR, JDR, PMA. Project Administration: PMA. Resources: AF, VR, PMA. Supervision: PMA, JDR. Validation: AF, VR, JDR, PMA. Visualization: AF, VR, JDR, PMA. Writing—original draft: AF, VR, JDR, PMA. Writing—review and editing: AF, VR, JDR, PMA.
Conflicts of Interest
The authors declare that they have no potential conflicts of interest.
Funding Statement
This study was funded by the Commission on Higher Education (CHED) of the Philippines and the Philippine Association of Medical Technologists (PAMET)-Safeguard (Procter & Gamble Philippines, Inc.) through its Postgraduate Scholarship Program Dagdag Karunungan, Kinabukusan ng Kalusugan.
| miRNA | Mean expression in tissue | Mean expression in tissue after outlier removal | Fold change in tissue (n = 41) | Mean expression in plasma | Mean expression in plasma after outlier removal | Fold change in plasma (n = 36) | Fold regulation in tissue and plasma | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
|
|
|
||||||||||||
| Malignant tissue (n = 41) | Adjacent neoplasm-free tissue (n = 41) | p-value | Malignant tissuea | Adjacent neoplasm-free tissuea | p-value | CRC cases (n = 36) | Clinically healthy controls (n = 36) | p-value | CRC casesa | Clinically healthy controlsa | p-value | ||||
| miR-21-5p | 9.35 ± 8.28 | 5.49 ± 6.44 | < .001 | 8.76 ± 7.44 | 1.43 ± 1.25 | < .001 | 4.23 | 5.84 ± 6.56 | 2.05 ± 4.15 | < .001 | 5.82 ± 6.65 | 0.44 ± 0.47 | < .001 | 8.40 | Upregulated |
| miR-196b-5p | 4.39 ± 5.09 | 2.19 ± 3.52 | .026 | 2.61 ± 2.79 | 0.96 ± 1.06 | .010 | 2.45 | 16.60 ± 12.09 | 3.13 ± 2.31 | < .001 | 19.20 ± 13.65 | 2.58 ± 1.40 | < .001 | 4.75 | Upregulated |
| miR-92a-3p | 4.45 ± 3.56 | 0.79 ± 1.35 | < .001 | 3.38 ± 2.02 | 0.59 ± 0.57 | < .001 | 7.41 | 7.63 ± 8.29 | 2.24 ± 2.93 | < .001 | 5.18 ± 4.32 | 1.43 ± 1.11 | < .001 | 2.50 | Upregulated |
| miR-29a-3p | 3.41 ± 2.53 | 1.27 ± 1.84 | < .001 | 3.40 ± 2.52 | 0.56 ± 0.33 | < .001 | 3.56 | 3.15 ± 3.79 | 0.80 ± 0.86 | < .001 | 2.35 ± 1.71 | 0.59 ± 0.43 | < .001 | 2.89 | Upregulated |
| miR-135b-5p | 1.06 ± 1.64 | 2.64 ± 2.58 | < .001 | 0.32 ± 0.43 | 2.47 ± 2.35 | < .001 | 0.17 | 0.89 ± 0.99 | 3.98 ± 4.06 | < .001 | 0.74 ± 0.69 | 2.96 ± 2.23 | < .001 | 0.44 | Downregulated |
| miR-197-3p | 3.33 ± 3.36 | 4.80 ± 7.38 | .676 | 2.61 ± 2.79 | 2.30 ± 2.07 | .681 | 0.78 | 3.86 ± 3.04 | 3.45 ± 3.50 | .112 | 3.15 ± 1.89 | 2.88 ± 1.14 | .656 | 1.32 | Not significant |
- 1. Kavousipour S, Khademi F, Zamani M, Vakili B, Mokarram P. Novel biotechnology approaches in colorectal cancer diagnosis and therapy. Biotechnol Lett 2017; 39: 785-803. ArticlePubMedPDF
- 2. Moreno CC, Mittal PK, Sullivan PS, et al. Colorectal cancer initial diagnosis: screening colonoscopy, diagnostic colonoscopy, or emergent surgery, and tumor stage and size at initial presentation. Clin Colorectal Cancer 2016; 15: 67-73. ArticlePubMed
- 3. Sung H, Ferlay J, Siegel RL, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin 2021; 71: 209-49. ArticlePubMedPDF
- 4. Pawa N, Arulampalam T, Norton JD. Screening for colorectal cancer: established and emerging modalities. Nat Rev Gastroenterol Hepatol 2011; 8: 711-22. ArticlePubMedPDF
- 5. Ahmed FE, Ahmed NC. MicroRNAs as molecular markers for colon cancer: diagnostic screening in stool and blood. Med Res Innov 2017; 1: 1-20.
- 6. Ju J. miRNAs as biomarkers in colorectal cancer diagnosis and prognosis. Bioanalysis 2010; 2: 901-6. ArticlePubMedPMC
- 7. Lin J, Chuang CC, Zuo L. Potential roles of microRNAs and ROS in colorectal cancer: diagnostic biomarkers and therapeutic targets. Oncotarget 2017; 8: 17328-46. ArticlePubMedPMC
- 8. Nagy ZB, Wichmann B, Kalmar A, et al. Colorectal adenoma and carcinoma specific miRNA profiles in biopsy and their expression in plasma specimens. Clin Epigenetics 2017; 9: 22.ArticlePubMedPMCPDF
- 9. Ng EK, Chong WW, Jin H, et al. Differential expression of microRNAs in plasma of patients with colorectal cancer: a potential marker for colorectal cancer screening. Gut 2009; 58: 1375-81. ArticlePubMed
- 10. Wikberg ML, Myte R, Palmqvist R, van Guelpen B, Ljuslinder I. Plasma miRNA can detect colorectal cancer, but how early? Cancer Med 2018; 7: 1697-705. PubMedPMCPDF
- 11. Saini V, Dawar R, Suneja S, Gangopadhyay S, Kaur C. Can microRNA become next-generation tools in molecular diagnostics and therapeutics?: a systematic review. Egypt J Med Hum Genet 2021; 22: 4.ArticlePDF
- 12. Buhagiar A, Seria E, Borg M, Borg J, Ayers D. Overview of microRNAs as liquid biopsy biomarkers for colorectal cancer sub-type profiling and chemoresistance. Cancer Drug Resist 2021; 4: 934-45. ArticlePubMedPMC
- 13. Zuo Z, Jiang Y, Zeng S, et al. The value of microRNAs as the novel biomarkers for colorectal cancer diagnosis: a meta-analysis. Pathol Res Pract 2020; 216: 153130.ArticlePubMed
- 14. Yau TO, Tang CM, Harriss EK, Dickins B, Polytarchou C. Faecal microRNAs as a non-invasive tool in the diagnosis of colonic adenomas and colorectal cancer: a meta-analysis. Sci Rep 2019; 9: 9491.ArticlePubMedPMCPDF
- 15. Pardini B, Sabo AA, Birolo G, Calin GA. Noncoding RNAs in extracellular fluids as cancer biomarkers: the new frontier of liquid biopsies. Cancers (Basel) 2019; 11: 1170.ArticlePubMedPMC
- 16. Hibner G, Kimsa-Furdzik M, Francuz T. Relevance of microRNAs as potential diagnostic and prognostic markers in colorectal cancer. Int J Mol Sci 2018; 19: 2944.ArticlePubMedPMC
- 17. Carter JV, Galbraith NJ, Yang D, Burton JF, Walker SP, Galandiuk S. Blood-based microRNAs as biomarkers for the diagnosis of colorectal cancer: a systematic review and meta-analysis. Br J Cancer 2017; 116: 762-74. ArticlePubMedPMCPDF
- 18. Yi R, Li Y, Wang FL, Miao G, Qi RM, Zhao YY. MicroRNAs as diagnostic and prognostic biomarkers in colorectal cancer. World J Gastrointest Oncol 2016; 8: 330-40. ArticlePubMedPMC
- 19. Wang DD, Chen X, Yu DD, et al. miR-197: a novel biomarker for cancers. Gene 2016; 591: 313-9. ArticlePubMed
- 20. Zhi ML, Liu ZJ, Yi XY, Zhang LJ, Bao YX. Diagnostic performance of microRNA-29a for colorectal cancer: a meta-analysis. Genet Mol Res 2015; 14: 18018-25. ArticlePubMed
- 21. Zhang H, Li P, Ju H, et al. Diagnostic and prognostic value of microRNA-21 in colorectal cancer: an original study and individual participant data meta-analysis. Cancer Epidemiol Biomarkers Prev 2014; 23: 2783-92. ArticlePubMedPDF
- 22. Yang X, Zeng Z, Hou Y, et al. MicroRNA-92a as a potential biomarker in diagnosis of colorectal cancer: a systematic review and meta-analysis. PLoS One 2014; 9: e88745.ArticlePubMedPMC
- 23. Motulsky HJ, Brown RE. Detecting outliers when fitting data with nonlinear regression: a new method based on robust nonlinear regression and the false discovery rate. BMC Bioinformatics 2006; 7: 123.ArticlePubMedPMCPDF
- 24. Wu Y, Song Y, Xiong Y, et al. MicroRNA-21 (Mir-21) promotes cell growth and invasion by repressing tumor suppressor PTEN in colorectal cancer. Cell Physiol Biochem 2017; 43: 945-58. ArticlePubMedPDF
- 25. Mima K, Nishihara R, Yang J, et al. MicroRNA MIR21 (miR-21) and PTGS2 expression in colorectal cancer and patient survival. Clin Cancer Res 2016; 22: 3841-8. ArticlePubMedPMCPDF
- 26. Yu Y, Kanwar SS, Patel BB, et al. MicroRNA-21 induces stemness by downregulating transforming growth factor beta receptor 2 (TGFbetaR2) in colon cancer cells. Carcinogenesis 2012; 33: 68-76. ArticlePubMedPMC
- 27. Feng YH, Tsao CJ. Emerging role of microRNA-21 in cancer. Biomed Rep 2016; 5: 395-402. ArticlePubMedPMC
- 28. Yamada A, Horimatsu T, Okugawa Y, et al. Serum miR-21, miR-29a, and miR-125b are promising biomarkers for the early detection of colorectal neoplasia. Clin Cancer Res 2015; 21: 4234-42. ArticlePubMedPMCPDF
- 29. Bastaminejad S, Taherikalani M, Ghanbari R, Akbari A, Shabab N, Saidijam M. Investigation of microRNA-21 expression levels in serum and stool as a potential non-invasive biomarker for diagnosis of colorectal cancer. Iran Biomed J 2017; 21: 106-13. ArticlePubMedPMC
- 30. Almeida AL, Bernardes MV, Feitosa MR, et al. Serological under expression of microRNA-21, microRNA-34a and microRNA-126 in colorectal cancer. Acta Cir Bras 2016; 31(Suppl 1): 13-8. ArticlePubMed
- 31. Jurkovicova D, Smolkova B, Magyerkova M, et al. Down-regulation of traditional oncomiRs in plasma of breast cancer patients. Oncotarget 2017; 8: 77369-84. ArticlePubMedPMC
- 32. Stiegelbauer V, Vychytilova-Faltejskova P, Karbiener M, et al. miR-196b-5p regulates colorectal cancer cell migration and metastases through interaction with HOXB7 and GALNT5. Clin Cancer Res 2017; 23: 5255-66. ArticlePubMedPDF
- 33. Lu YC, Chang JT, Huang YC, et al. Combined determination of circulating miR-196a and miR-196b levels produces high sensitivity and specificity for early detection of oral cancer. Clin Biochem 2015; 48: 115-21. ArticlePubMed
- 34. Li X, Zhang G, Luo F, et al. Identification of aberrantly expressed miRNAs in rectal cancer. Oncol Rep 2012; 28: 77-84. PubMed
- 35. Wu J, Lin B, Yu S, et al. Exosomal miR-196b-5p is a potential diagnostic marker for colorectal cancer with metachronous liver metastasis. Transl Cancer Res 2018; 7: 1482-90. Article
- 36. Xu C, Gu L. The diagnostic effect of serum miR-196b as biomarker in colorectal cancer. Biomed Rep 2017; 6: 39-45. ArticlePubMedPMC
- 37. Li Y, Zhang M, Chen H, et al. Ratio of miR-196s to HOXC8 messenger RNA correlates with breast cancer cell migration and metastasis. Cancer Res 2010; 70: 7894-904. ArticlePubMedPMCPDF
- 38. Braig S, Mueller DW, Rothhammer T, Bosserhoff AK. MicroRNA miR-196a is a central regulator of HOX-B7 and BMP4 expression in malignant melanoma. Cell Mol Life Sci 2010; 67: 3535-48. ArticlePubMedPMCPDF
- 39. Bhatia S, Kaul D, Varma N. Potential tumor suppressive function of miR-196b in B-cell lineage acute lymphoblastic leukemia. Mol Cell Biochem 2010; 340: 97-106. ArticlePubMedPDF
- 40. Brunet Vega A, Pericay C, Moya I, et al. microRNA expression profile in stage III colorectal cancer: circulating miR-18a and miR-29a as promising biomarkers. Oncol Rep 2013; 30: 320-6. ArticlePubMed
- 41. Giraldez MD, Lozano JJ, Ramirez G, et al. Circulating microRNAs as biomarkers of colorectal cancer: results from a genome-wide profiling and validation study. Clin Gastroenterol Hepatol 2013; 11: 681-8. ArticlePubMed
- 42. Liu X, Lv X, Yang Q, Jin H, Zhou W, Fan Q. MicroRNA-29a functions as a tumor suppressor and increases cisplatin sensitivity by targeting NRAS in lung cancer. Technol Cancer Res Treat 2018; 17: 1533033818758905.ArticlePubMedPMCPDF
- 43. Tang W, Zhu Y, Gao J, et al. MicroRNA-29a promotes colorectal cancer metastasis by regulating matrix metalloproteinase 2 and E-cadherin via KLF4. Br J Cancer 2014; 110: 450-8. ArticlePubMedPMCPDF
- 44. Pei YF, Lei Y, Liu XQ. MiR-29a promotes cell proliferation and EMT in breast cancer by targeting ten eleven translocation 1. Biochim Biophys Acta 2016; 1862: 2177-85. ArticlePubMed
- 45. Trehoux S, Lahdaoui F, Delpu Y, et al. Micro-RNAs miR-29a and miR-330-5p function as tumor suppressors by targeting the MUC1 mucin in pancreatic cancer cells. Biochim Biophys Acta 2015; 1853: 2392-403. ArticlePubMed
- 46. Li M, Guan X, Sun Y, et al. miR-92a family and their target genes in tumorigenesis and metastasis. Exp Cell Res 2014; 323: 1-6. ArticlePubMed
- 47. Tsuchida A, Ohno S, Wu W, et al. miR-92 is a key oncogenic component of the miR-17-92 cluster in colon cancer. Cancer Sci 2011; 102: 2264-71. ArticlePubMed
- 48. Ke TW, Wei PL, Yeh KT, Chen WT, Cheng YW. MiR-92a promotes cell metastasis of colorectal cancer through PTEN-mediated PI3K/AKT pathway. Ann Surg Oncol 2015; 22: 2649-55. ArticlePubMedPDF
- 49. Li J, Liang H, Bai M, et al. miR-135b promotes cancer progression by targeting transforming growth factor beta receptor II (TGFBR2) in colorectal cancer. PLoS One 2015; 10: e0130194.ArticlePubMedPMC
- 50. Magalhaes L, Quintana LG, Lopes DCF, et al. APC gene is modulated by hsa-miR-135b-5p in both diffuse and intestinal gastric cancer subtypes. BMC Cancer 2018; 18: 1055.PubMedPMCPDF
- 51. Zekri AR, Youssef AS, Lotfy MM, et al. Circulating serum miRNAs as diagnostic markers for colorectal cancer. PLoS One 2016; 11: e0154130.ArticlePubMedPMC
- 52. Bastaminejad S, Taherikalani M, Ghanbari R, et al. Serum and stool miR-135b levels as a potential diagnostic biomarker for colorectal cancer. Clin Exp Invest 2020; 1: 1-6.
- 53. Uddin MN, Li M, Wang X. Identification of transcriptional markers and microRNA-mRNA regulatory networks in colon cancer by integrative analysis of mRNA and microRNA expression profiles in colon tumor stroma. Cells 2019; 8: 1054.ArticlePubMedPMC
- 54. Reichholf B, Herzog VA, Fasching N, Manzenreither RA, Sowemimo I, Ameres SL. Time-resolved small RNA sequencing unravels the molecular principles of microRNA homeostasis. Mol Cell 2019; 75: 756-68. ArticlePubMedPMC
- 55. Cojocneanu R, Braicu C, Raduly L, et al. Plasma and tissue specific miRNA expression pattern and functional analysis associated to colorectal cancer patients. Cancers (Basel) 2020; 12: 843.ArticlePubMedPMC
- 56. Nagy ZB, Bartak BK, Kalmar A, et al. Comparison of circulating miRNAs expression alterations in matched tissue and plasma samples during colorectal cancer progression. Pathol Oncol Res 2019; 25: 97-105. ArticlePubMedPDF
- 57. Ma R, Jiang T, Kang X. Circulating microRNAs in cancer: origin, function and application. J Exp Clin Cancer Res 2012; 31: 38.ArticlePubMedPMCPDF
- 58. De Rosa M, Rega D, Costabile V, et al. The biological complexity of colorectal cancer: insights into biomarkers for early detection and personalized care. Therap Adv Gastroenterol 2016; 9: 861-86. ArticlePubMedPMCPDF
- 59. Balboa E, Carracedo A, Barros F. The complexity of colorectal cancer biology: putting bricks on the path to personalized medicine. Colorectal cancer. In: Khan JS, ed. Rijeka: Intech, 2014; 434-66. Article
- 60. Nguyen HT, Duong HQ. The molecular characteristics of colorectal cancer: implications for diagnosis and therapy. Oncol Lett 2018; 16: 9-18. PubMedPMC
- 61. Diaz-Cano SJ. Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci 2012; 13: 1951-2011. ArticlePubMedPMC
- 62. Buikhuisen JY, Torang A, Medema JP. Exploring and modelling colon cancer inter-tumour heterogeneity: opportunities and challenges. Oncogenesis 2020; 9: 66.ArticlePubMedPMCPDF
REFERENCES
Figure & Data
References
Citations

- Liquid biopsy for the management of gastrointestinal cancers
Zeinab Salehnia, Delsuz Rezaee, Sajad Ehtiati, Mohammad Bakhtiari, Mohammad Amin Khalilzad, Sajad Najafi
Clinica Chimica Acta.2026; 578: 120474. CrossRef - Differential Circulating miRNA Responses to PM Exposure in Healthy and Diabetes Mellitus Patients: Implications for Lung Cancer Susceptibility
Moe Thi Thi Han, Nichakorn Satitpornbunpot, Naoomi Tominaga, Saranta Freeouf, Khanittha Punturee, Chidchamai Kewchareonwong, Busayamas Chewaskulyong, Ganjana Lertmemongkolchai, Ratchada Cressey
International Journal of Molecular Sciences.2026; 27(2): 613. CrossRef - Unlocking the power of non-coding RNAs: toward real-time cancer monitoring in precision oncology
Manon Chang, Thomas Papazyan, Elvire Pons-Tostivint, Delphine Fradin
Molecular Cancer.2026;[Epub] CrossRef - Diagnostic Performance of Circulating miRNA-92a and Related Circulating miRNA Panels for Colorectal Cancer: An Updated Systematic Review and Meta-Analysis
Rosmeri Handayani, Siti Aliyah, Muhammad Begawan Bestari, Reno Rudiman, Ninik Sukartini, Ida Parwati
OncoTargets and Therapy.2026; Volume 19: 1. CrossRef - Exosome-derived hsa-miR-21-5p stimulates malignant progression of colorectal cancer by regulating HRK
Shengyong You, Lingyun Gu, Huirong Wu, Cuihua Wu
Biochemical and Biophysical Research Communications.2026; 819: 153829. CrossRef - Except for Robust Outliers, Rapamycin Increases Lesion Burden in a Murine Model of Cerebral Cavernous Malformations
Roberto J. Alcazar-Felix, Robert Shenkar, Christian R. Benavides, Akash Bindal, Abhinav Srinath, Ying Li, Serena Kinkade, Tatiana Terranova, Evon DeBose-Scarlett, Rhonda Lightle, Dorothy DeBiasse, Hanadi Almazroue, Diana Vera Cruz, Sharbel Romanos, Aditya
Translational Stroke Research.2025; 16(3): 859. CrossRef - miR-135b: A key role in cancer biology and therapeutic targets
Yingchun Shao, Shuangshuang Zhang, Yuxin Pan, Zhan Peng, Yinying Dong
Non-coding RNA Research.2025; 12: 67. CrossRef - The role of tumor-associated fibroblast-derived exosomes in chemotherapy resistance of colorectal cancer and its application prospect
Meichen Liu, Teng-zheng Li, Congcong Xu
Biochimica et Biophysica Acta (BBA) - General Subjects.2025; 1869(6): 130796. CrossRef - A Systematic Review of MicroRNAs in Hemorrhagic Neurovascular Disease: Cerebral Cavernous Malformations as a Paradigm
Roberto J. Alcazar-Felix, Aditya Jhaveri, Javed Iqbal, Abhinav Srinath, Carolyn Bennett, Akash Bindal, Diana Vera Cruz, Sharbel Romanos, Stephanie Hage, Agnieszka Stadnik, Justine Lee, Rhonda Lightle, Robert Shenkar, Janne Koskimäki, Sean P. Polster, Romu
International Journal of Molecular Sciences.2025; 26(8): 3794. CrossRef - microRNA-196b-5p expression in cancer tissues is closely associated with clinical and pathological characteristics and prognosis of patients with non-small cell lung cancer
Wei Liu, Xiaomin Lu, Changgang Yang, Fengjun Ji, Xiaodan Wu
Journal of Cardiothoracic Surgery.2025;[Epub] CrossRef - Association of mir196a2 and mir146a polymorphisms and colorectal cancer risk: A meta-analysis
Zahraa isam jameel, Hanan Ali Kareem, Zahraa Mohammed Yahya, Ahmed Ali Hussein, zahraa abdel Kareem
Human Gene.2025; 45: 201448. CrossRef - Micro RNA in Colorectal Cancer—Potential Diagnostic and Prognostic Markers—An Updated Review
Weronika Pająk, Jakub Kleinrok, Joanna Pec, Karolina Michno, Jan Wojtas, Miłosz Badach, Barbara Teresińska, Jacek Baj
International Journal of Molecular Sciences.2025; 26(17): 8615. CrossRef - MicroRNAs in colorectal cancer: A comparative analysis of circulating and tissue microRNA levels
Iulia Andreea Pelisenco, Bogdan Trandafir, Anastasia-Maria Dobre, Andrei-Daniel Dragne, Vlad Herlea, Andrei Marian Niculae, Catalin Vasilescu, Mihail Eugen Hinescu, Elena Milanesi, Maria Dobre
World Journal of Gastrointestinal Oncology.2025;[Epub] CrossRef - Identification of miRNAs Present in Cell- and Plasma-Derived Extracellular Vesicles—Possible Biomarkers of Colorectal Cancer
Marzena Lenart, Izabela Siemińska, Rafał Szatanek, Anna Mordel, Antoni Szczepanik, Mateusz Rubinkiewicz, Maciej Siedlar, Monika Baj-Krzyworzeka
Cancers.2024; 16(13): 2464. CrossRef - The emerging role of blood-based biomarkers in early detection of colorectal cancer: A systematic review
Faris Shweikeh, Yuhao Zeng, Abdur Rahman Jabir, Erica Whittenberger, Saurav P. Kadatane, Yuting Huang, Mohamad Mouchli, Dani Ran Castillo
Cancer Treatment and Research Communications.2024; 42: 100872. CrossRef - The Role of Extracellular Vesicles in Colorectal Cancer
Yujian Xia, Chaoran Yu, Ernest Johann Helwig, Yousheng Li
Technology in Cancer Research & Treatment.2023;[Epub] CrossRef - HNRNPA2B1-Mediated MicroRNA-92a Upregulation and Section Acts as a Promising Noninvasive Diagnostic Biomarker in Colorectal Cancer
Yiling Li, Kexin Li, Xiaoying Lou, Yue Wu, Samuel Seery, Danfei Xu, Yuqing Pei, Benheng Qian, Yuxin Wu, Shuang Liang, Kui Wu, Wei Cui
Cancers.2023; 15(4): 1367. CrossRef - Role of salivary miRNAs in the diagnosis of gastrointestinal disorders: a mini-review of available evidence
Maria Oana Săsăran, Claudia Bănescu
Frontiers in Genetics.2023;[Epub] CrossRef - Exploring the Role of Circulating Cell-Free RNA in the Development of Colorectal Cancer
Chau-Ming Kan, Xiao Meng Pei, Martin Ho Yin Yeung, Nana Jin, Simon Siu Man Ng, Hin Fung Tsang, William Chi Shing Cho, Aldrin Kay-Yuen Yim, Allen Chi-Shing Yu, Sze Chuen Cesar Wong
International Journal of Molecular Sciences.2023; 24(13): 11026. CrossRef - Lynch Syndrome Biopathology and Treatment: The Potential Role of microRNAs in Clinical Practice
Serena Ascrizzi, Grazia Maria Arillotta, Katia Grillone, Giulio Caridà, Stefania Signorelli, Asad Ali, Caterina Romeo, Pierfrancesco Tassone, Pierosandro Tagliaferri
Cancers.2023; 15(15): 3930. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure

Fig. 1
| Characteristics | No (%) (n = 41) |
|---|---|
| Age | |
| < 60 | 18 (44) |
| > 60 | 23 (56) |
| Sex | |
| Male | 18 (44) |
| Female | 23 (56) |
| Location of tumor | |
| Rectum/cecum/rectosigmoid | 34 (83) |
| Sigmoid ascending/transverse and descending | 7 (17) |
| Histologic type | |
| Adenocarcinoma | 33 (80) |
| Mucinous cell carcinoma/signet ring cell carcinoma | 8 (20) |
| Histologic grade | |
| Well-differentiated | 26 (63) |
| Moderately to poorly differentiated | 15 (37) |
| TNM stage | |
| Stage I/II | 20 (49) |
| Stage III/IV | 21(51) |
| Lymph node metastasis | |
| N0–N1 | 29 (71) |
| N2 | 12 (29) |
| Distant metastasis | |
| M0–M1 | 28 (68) |
| Mx | 13 (32) |
| Tumor size | |
| < 5 cm | 18 (44) |
| > 5 cm | 23 (56) |
| Depth of tumor infiltration | |
| Muscularis propria | 18 (44) |
| Muscularis propria into the pericolic adipose tissue and outside | 23 (56) |
| Perineural invasion | |
| Absent | 39 (95) |
| Present | 2 (5) |
| miRNA | Mean expression in tissue | Mean expression in tissue after outlier removal | Fold change in tissue (n = 41) | Mean expression in plasma | Mean expression in plasma after outlier removal | Fold change in plasma (n = 36) | Fold regulation in tissue and plasma | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
|
|
| ||||||||||||
| Malignant tissue (n = 41) | Adjacent neoplasm-free tissue (n = 41) | p-value | Malignant tissue |
Adjacent neoplasm-free tissue |
p-value | CRC cases (n = 36) | Clinically healthy controls (n = 36) | p-value | CRC cases |
Clinically healthy controls |
p-value | ||||
| miR-21-5p | 9.35 ± 8.28 | 5.49 ± 6.44 | < .001 | 8.76 ± 7.44 | 1.43 ± 1.25 | < .001 | 4.23 | 5.84 ± 6.56 | 2.05 ± 4.15 | < .001 | 5.82 ± 6.65 | 0.44 ± 0.47 | < .001 | 8.40 | Upregulated |
| miR-196b-5p | 4.39 ± 5.09 | 2.19 ± 3.52 | .026 | 2.61 ± 2.79 | 0.96 ± 1.06 | .010 | 2.45 | 16.60 ± 12.09 | 3.13 ± 2.31 | < .001 | 19.20 ± 13.65 | 2.58 ± 1.40 | < .001 | 4.75 | Upregulated |
| miR-92a-3p | 4.45 ± 3.56 | 0.79 ± 1.35 | < .001 | 3.38 ± 2.02 | 0.59 ± 0.57 | < .001 | 7.41 | 7.63 ± 8.29 | 2.24 ± 2.93 | < .001 | 5.18 ± 4.32 | 1.43 ± 1.11 | < .001 | 2.50 | Upregulated |
| miR-29a-3p | 3.41 ± 2.53 | 1.27 ± 1.84 | < .001 | 3.40 ± 2.52 | 0.56 ± 0.33 | < .001 | 3.56 | 3.15 ± 3.79 | 0.80 ± 0.86 | < .001 | 2.35 ± 1.71 | 0.59 ± 0.43 | < .001 | 2.89 | Upregulated |
| miR-135b-5p | 1.06 ± 1.64 | 2.64 ± 2.58 | < .001 | 0.32 ± 0.43 | 2.47 ± 2.35 | < .001 | 0.17 | 0.89 ± 0.99 | 3.98 ± 4.06 | < .001 | 0.74 ± 0.69 | 2.96 ± 2.23 | < .001 | 0.44 | Downregulated |
| miR-197-3p | 3.33 ± 3.36 | 4.80 ± 7.38 | .676 | 2.61 ± 2.79 | 2.30 ± 2.07 | .681 | 0.78 | 3.86 ± 3.04 | 3.45 ± 3.50 | .112 | 3.15 ± 1.89 | 2.88 ± 1.14 | .656 | 1.32 | Not significant |
| miRNA | Mean expression of miRNA in tissue | p-value | Mean expression of miRNA in plasma | p-value | ||
|---|---|---|---|---|---|---|
|
|
| |||||
| Early stage (n = 20) | Advanced stage (n = 21) | Early stage (n = 16) | Advanced stage (n = 20) | |||
| miR-21-5p | 10.13 ± 10.87 | 8.02 ± 7.39 | .531 | 12.33 ± 5.33 | 7.14 ± 5.13 | .006 |
| miR-196b-5p | 3.58 ± 2.53 | 5.02 ± 5.05 | .769 | 10.76 ± 1.93 | 5.54 ± 2.35 | < .001 |
| miR-135b-5p | 1.05 ± 1.63 | 0.76 ± 1.26 | .611 | 2.34 ± 1.22 | 10.80 ± 3.76 | < .001 |
| miR-92a-3p | 12.51 ± 11.89 | 14.90 ± 14.59 | .578 | 9.92 ± 2.53 | 2.94 ± 2.95 | < .001 |
| miR-29a-3p | 5.30 ± 3.19 | 5.06 ± 4.14 | .473 | 6.67 ± 2.18 | 3.32 ± 2.18 | < .001 |
| miR-197-3p | 1.18 ± 0.73 | 1.50 ± 1.49 | .879 | 1.16 ± 1.03 | 2.60 ± 2.05 | .007 |
| Sample | miR-21-5p | miR-196b-5p | miR-135b-5p | miR-92a-3p | miR-29a-3p | miR-197-3p | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
|
|
|
|
| |||||||||||||
| Mean expression level | r | p-value | Mean expression level | r | p-value | Mean expression level | r | p-value | Mean expression level | r | p-value | Mean expression level | r | p-value | Mean expression level | r | p-value | |
| Tissue | 9.35 ± 8.28 | 0.33 | .049 | 4.39 ± 5.09 | 0.02 | .921 | 1.06 ± 1.64 | 0.14 | .430 | 4.45 ± 3.56 | 0.54 | .007 | 3.41 ± 2.53 | 0.52 | .048 | 3.33 ± 3.36 | −0.24 | .150 |
| Plasma | 5.84 ± 6.56 | 16.60 ± 12.09 | 0.89 ± 0.99 | 7.63 ± 8.29 | 3.15 ± 3.79 | 3.86 ± 3.04 | ||||||||||||
| miRNA | Diagnostic performance for CRC irrespective of cancer stage using tissue samples | Diagnostic performance for CRC irrespective of cancer stage using plasma samples | Diagnostic performance for early-stage CRC using plasma samples | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
|
|
|
| ||||||||||
| AUC | Cutoff point | Sensitivity | Specificity | AUC | Cutoff point | Sensitivity | Specificity | AUC | Cutoff point | Sensitivity | Specificity | |
| miR-21-5p | 0.78 | 3.56 | 0.80 | 0.83 | 0.78 | 3.89 | 0.81 | 0.83 | 0.77 | 3.69 | 0.80 | 0.81 |
| miR-196b-5p | 0.64 | 0.97 | 0.68 | 0.71 | 0.94 | 4.96 | 0.97 | 0.94 | 0.94 | 5.24 | 0.95 | 0.94 |
| miR-135b-5p | 0.76 | 2.32 | 0.80 | 0.83 | 0.83 | 1.45 | 0.86 | 0.89 | 0.84 | 2.05 | 0.85 | 0.88 |
| miR-92a-3p | 0.89 | 1.40 | 0.93 | 0.95 | 0.76 | 1.39 | 0.81 | 0.81 | 0.95 | 1.79 | 0.95 | 0.94 |
| miR-29a-3p | 0.81 | 1.23 | 0.85 | 0.83 | 0.83 | 1.21 | 0.86 | 0.83 | 0.87 | 0.41 | 0.90 | 0.88 |
| miR-197-3p | 0.52 | 2.41 | 0.56 | 0.63 | 0.60 | 2.06 | 0.61 | 0.67 | 0.76 | 2.41 | 0.80 | 0.81 |
The number of samples after the outlier removal from each miRNA varied depending on the number of samples (outlier) removed during the Robust Regression and Outlier Removal analysis.
miRNA (miR), microRNA; CRC, colorectal cancer.
miRNA (miR), microRNA; CRC, colorectal cancer.
miRNA (miR), microRNA; AUC, area under the receiver operating characteristic curve; CRC, colorectal cancer.

 E-submission
E-submission