Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 47(1); 2013 > Article
-
Original Article
Comparison of Direct Sequencing, PNA Clamping-Real Time Polymerase Chain Reaction, and Pyrosequencing Methods for the Detection ofEGFR Mutations in Non-small Cell Lung Carcinoma and the Correlation with Clinical Responses to EGFR Tyrosin - Hyun Ju Lee1,2, Xianhua Xu1, Hyojin Kim1, Yan Jin1, Pingli Sun1, Ji Eun Kim3, Jin-Haeng Chung1
-
Korean Journal of Pathology 2013;47(1):52-60.
DOI: https://doi.org/10.4132/KoreanJPathol.2013.47.1.52
Published online: February 25, 2013
1Department of Pathology, Seoul National University Bundang Hospital, Seoul National University College of Medicine, Seongnam, Korea.
2Department of Pathology, Soonchunhyang University Cheonan Hospital, Soonchunhyang University College of Medicine, Cheonan, Korea.
3Department of Pathology, Boramae Medical Center, Seoul National University College of Medicine, Seoul, Korea.
- Corresponding Author: Jin-Haeng Chung, M.D., Ph.D. Department of Pathology, Seoul National University Bundang Hospital, Seoul National University College of Medicine, 82 Gumi-ro 173beon-gil, Bundang-gu, Seongnam 463-707, Korea. Tel: +82-31-787-7713, Fax: +82-31-787-4012, chungjh@snu.ac.kr, Ji Eun Kim, M.D., Ph.D. Department of Pathology, Boramae Medical Center, Seoul National University College of Medicine, 20 Boramae-ro 5-gil, Dongjak-gu, Seoul 156-707, Korea. Tel: +82-2-870-2642, Fax: +82-2-831-0261, npol181@snu.ac.kr
© 2013 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Figure & Data
References
Citations

- Recent Trends of Lung Cancer in Korea
Jae Guk Lee, Ho Cheol Kim, Chang-Min Choi
Tuberculosis and Respiratory Diseases.2021; 84(2): 89. CrossRef - Predictive value of KRAS mutation and excision repair cross-complementing 1 (ERCC1) protein overexpression in patients with colorectal cancer administered FOLFOX regimen
Sun Min Park, Sung Bong Choi, Yoon Suk Lee, In Kyu Lee
Asian Journal of Surgery.2021; 44(5): 715. CrossRef - Recent advances in diagnostic technologies in lung cancer
Hye Jung Park, Sang Hoon Lee, Yoon Soo Chang
The Korean Journal of Internal Medicine.2020; 35(2): 257. CrossRef - Afatinib is effective in the treatment of lung adenocarcinoma with uncommon EGFR p.L747P and p.L747S mutations
Sheng-Kai Liang, Jen-Chung Ko, James Chih-Hsin Yang, Jin-Yuan Shih
Lung Cancer.2019; 133: 103. CrossRef - Biomarker Testing for Patients With Advanced Non–Small Cell Lung Cancer: Real-World Issues and Tough Choices
Nathan A. Pennell, Maria E. Arcila, David R. Gandara, Howard West
American Society of Clinical Oncology Educational Book.2019; (39): 531. CrossRef - Evaluation of EGFR mutations in NSCLC with highly sensitive droplet digital PCR assays
Xi‑Wen Jiang, Wei Liu, Xiao‑Ya Zhu, Xiao‑Xie Xu
Molecular Medicine Reports.2019;[Epub] CrossRef - Peptide Nucleic Acid Clamping and Direct Sequencing in the Detection of Oncogenic Alterations in Lung Cancer: Systematic Review and Meta-Analysis
Jae-Uk Song, Jonghoo Lee
Yonsei Medical Journal.2018; 59(2): 211. CrossRef - Distribution of KRAS, DDR2, and TP53 gene mutations in lung cancer: An analysis of Iranian patients
Zahra Fathi, Seyed Ali Javad Mousavi, Raheleh Roudi, Farideh Ghazi, Sumitra Deb
PLOS ONE.2018; 13(7): e0200633. CrossRef - EGFR T790M mutation testing within the osimertinib AURA Phase I study
Simon Dearden, Helen Brown, Suzanne Jenkins, Kenneth S. Thress, Mireille Cantarini, Rebecca Cole, Malcolm Ranson, Pasi A. Jänne
Lung Cancer.2017; 109: 9. CrossRef - Molecular Testing of Lung Cancers
Hyo Sup Shim, Yoon-La Choi, Lucia Kim, Sunhee Chang, Wan-Seop Kim, Mee Sook Roh, Tae-Jung Kim, Seung Yeon Ha, Jin-Haeng Chung, Se Jin Jang, Geon Kook Lee
Journal of Pathology and Translational Medicine.2017; 51(3): 242. CrossRef - Mutations of the Epidermal Growth Factor Receptor Gene in Triple-Negative Breast Cancer
Aeri Kim, Min Hye Jang, Soo Jung Lee, Young Kyung Bae
Journal of Breast Cancer.2017; 20(2): 150. CrossRef - Double primary lung adenocarcinoma diagnosed by epidermal growth factor receptor mutation status
Oh Jung Kwon, Min Hyeok Lee, Sung Ju Kang, Seul Gi Kim, In Beom Jeong, Ji Yun Jeong, Eun Jung Cha, Do Yeun Cho, Young Jin Kim, Ji Woong Son
Yeungnam University Journal of Medicine.2017; 34(2): 270. CrossRef - Generation of lung cancer cell lines harboring EGFR T790M mutation by CRISPR/Cas9-mediated genome editing
Mi-Young Park, Min Hee Jung, Eun Young Eo, Seokjoong Kim, Sang Hoon Lee, Yeon Joo Lee, Jong Sun Park, Young Jae Cho, Jin Haeng Chung, Cheol Hyeon Kim, Ho Il Yoon, Jae Ho Lee, Choon-Taek Lee
Oncotarget.2017; 8(22): 36331. CrossRef - Comparison of EGFR mutation detection between the tissue and cytology using direct sequencing, pyrosequencing and peptide nucleic acid clamping in lung adenocarcinoma: Korean multicentre study
Kyueng-Whan Min, Wan-Seop Kim, Se Jin Jang, Yoo Duk Choi, Sunhee Chang, Soon Hee Jung, Lucia Kim, Mee Sook Roh, Choong Sik Lee, Jung Weon Shim, Mi Jin Kim, Geon Kook Lee
QJM.2016; 109(3): 167. CrossRef - Epidermal Growth Factor Receptor Mutation and Anaplastic Lymphoma Kinase Gene Fusion: Detection in Malignant Pleural Effusion by RNA or PNA Analysis
Yi-Lin Chen, Chung-Ta Lee, Cheng-Chan Lu, Shu-Ching Yang, Wan-Li Chen, Yang-Cheng Lee, Chung-Hsien Yang, Shu-Ling Peng, Wu-Chou Su, Nan-Haw Chow, Chung-Liang Ho, Javier S Castresana
PLOS ONE.2016; 11(6): e0158125. CrossRef - IDH Mutation Analysis in Ewing Sarcoma Family Tumors
Ki Yong Na, Byeong-Joo Noh, Ji-Youn Sung, Youn Wha Kim, Eduardo Santini Araujo, Yong-Koo Park
Journal of Pathology and Translational Medicine.2015; 49(3): 257. CrossRef - Immunohistochemical demonstration of alteration of β-catenin during tumor metastasis by different mechanisms according to histology in lung cancer
XIANHUA XU, JI EUN KIM, PING-LI SUN, SEOL BONG YOO, HYOJIN KIM, YAN JIN, JIN-HAENG CHUNG
Experimental and Therapeutic Medicine.2015; 9(2): 311. CrossRef - Detection of EGFR-TK Domain–activating Mutations in NSCLC With Generic PCR-based Methods
Rajendra B. Shahi, Sylvia De Brakeleer, Jacques De Grève, Caroline Geers, Peter In’t Veld, Erik Teugels
Applied Immunohistochemistry & Molecular Morphology.2015; 23(3): 163. CrossRef - Frequent aerogenous spread with decreased E-cadherin expression of ROS1- rearranged lung cancer predicts poor disease-free survival
Yan Jin, Ping-Li Sun, Soo Young Park, Hyojin Kim, Eunhyang Park, Gilhyang Kim, Sukki Cho, Kwhanmien Kim, Choon-Taek Lee, Jin-Haeng Chung
Lung Cancer.2015; 89(3): 343. CrossRef - Membranous Insulin-like Growth Factor-1 Receptor (IGF1R) Expression Is Predictive of Poor Prognosis in Patients with Epidermal Growth Factor Receptor (EGFR)-Mutant Lung Adenocarcinoma
Eunhyang Park, Soo Young Park, Hyojin Kim, Ping-Li Sun, Yan Jin, Suk Ki Cho, Kwhanmien Kim, Choon-Taek Lee, Jin-Haeng Chung
Journal of Pathology and Translational Medicine.2015; 49(5): 382. CrossRef - Peptide Nucleic Acid Clamping Versus Direct Sequencing for the Detection of EGFR Gene Mutation in Patients with Non-small Cell Lung Cancer
Seong-Hoon Yoon, Yoo-Duk Choi, In-Jae Oh, Kyu-Sik Kim, Hayoung Choi, Jinsun Chang, Hong-Joon Shin, Cheol-Kyu Park, Young-Chul Kim
Cancer Research and Treatment.2015; 47(4): 661. CrossRef - Analysis of Mutations in Epidermal Growth Factor Receptor Gene in Korean Patients with Non-small Cell Lung Cancer: Summary of a Nationwide Survey
Sang Hwa Lee, Wan Seop Kim, Yoo Duk Choi, Jeong Wook Seo, Joung Ho Han, Mi Jin Kim, Lucia Kim, Geon Kook Lee, Chang Hun Lee, Mee Hye Oh, Gou Young Kim, Sun Hee Sung, Kyo Young Lee, Sun Hee Chang, Mee Sook Rho, Han Kyeom Kim, Soon Hee Jung, Se Jin Jang
Journal of Pathology and Translational Medicine.2015; 49(6): 481. CrossRef - Novel EGFR mutation-specific antibodies for lung adenocarcinoma: Highly specific but not sensitive detection of an E746_A750 deletion in exon 19 and an L858R mutation in exon 21 by immunohistochemistry
An Na Seo, Tae-In Park, Yan Jin, Ping-Li Sun, Hyojin Kim, Hyun Chang, Jin-Haeng Chung
Lung Cancer.2014; 83(3): 316. CrossRef - Simultaneous diagnostic platform of genotyping EGFR, KRAS, and ALK in 510 Korean patients with non‐small‐cell lung cancer highlights significantly higher ALK rearrangement rate in advanced stage
Tae‐Jung Kim, Chan Kwon Park, Chang Dong Yeo, Kihoon Park, Chin Kook Rhee, Jusang Kim, Seung Joon Kim, Sang Haak Lee, Kyo‐Young Lee, Hyoung‐Kyu Yoon
Journal of Surgical Oncology.2014; 110(3): 245. CrossRef - Epidermal growth factor receptor mutations and anaplastic lymphoma kinase rearrangements in lung cancer with nodular ground-glass opacity
Sung-Jun Ko, Yeon Joo Lee, Jong Sun Park, Young-Jae Cho, Ho Il Yoon, Jin-Haeng Chung, Tae Jung Kim, Kyung Won Lee, Kwhanmien Kim, Sanghoon Jheon, Hyojin Kim, Jae Ho Lee, Choon-Taek Lee
BMC Cancer.2014;[Epub] CrossRef - Cytoplasmic YAP Expression is Associated with Prolonged Survival in Patients with Lung Adenocarcinomas and Epidermal Growth Factor Receptor Tyrosine Kinase Inhibitor Treatment
Ping-Li Sun, Ji Eun Kim, Seol Bong Yoo, Hyojin Kim, Yan Jin, Sanghoon Jheon, Kwhanmien Kim, Choon Taek Lee, Jin-Haeng Chung
Annals of Surgical Oncology.2014; 21(S4): 610. CrossRef - Sensitive methods for detection of the S768R substitution in exon 18 of the DDR2 gene in patients with central nervous system metastases of non-small cell lung cancer
Marcin Nicoś, Tomasz Powrózek, Paweł Krawczyk, Bożena Jarosz, Beata Pająk, Marek Sawicki, Krzysztof Kucharczyk, Tomasz Trojanowski, Janusz Milanowski
Medical Oncology.2014;[Epub] CrossRef - Clinicopathologic and prognostic significance of c-MYC copy number gain in lung adenocarcinomas
A N Seo, J M Yang, H Kim, S Jheon, K Kim, C T Lee, Y Jin, S Yun, J-H Chung, J H Paik
British Journal of Cancer.2014; 110(11): 2688. CrossRef - KRASMutation Detection in Non-small Cell Lung Cancer Using a Peptide Nucleic Acid-Mediated Polymerase Chain Reaction Clamping Method and Comparative Validation with Next-Generation Sequencing
Boram Lee, Boin Lee, Gangmin Han, Mi Jung Kwon, Joungho Han, Yoon-La Choi
Korean Journal of Pathology.2014; 48(2): 100. CrossRef - Guideline Recommendations forEGFRMutation Testing in Lung Cancer: Proposal of the Korean Cardiopulmonary Pathology Study Group
Hyo Sup Shim, Jin-Haeng Chung, Lucia Kim, Sunhee Chang, Wan-Seop Kim, Geon Kook Lee, Soon-Hee Jung, Se Jin Jang
Korean Journal of Pathology.2013; 47(2): 100. CrossRef - Immunohistochemical Classification of Primary and Secondary Glioblastomas
Kyu Sang Lee, Gheeyoung Choe, Kyung Han Nam, An Na Seo, Sumi Yun, Kyung Ju Kim, Hwa Jin Cho, Sung Hye Park
Korean Journal of Pathology.2013; 47(6): 541. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure

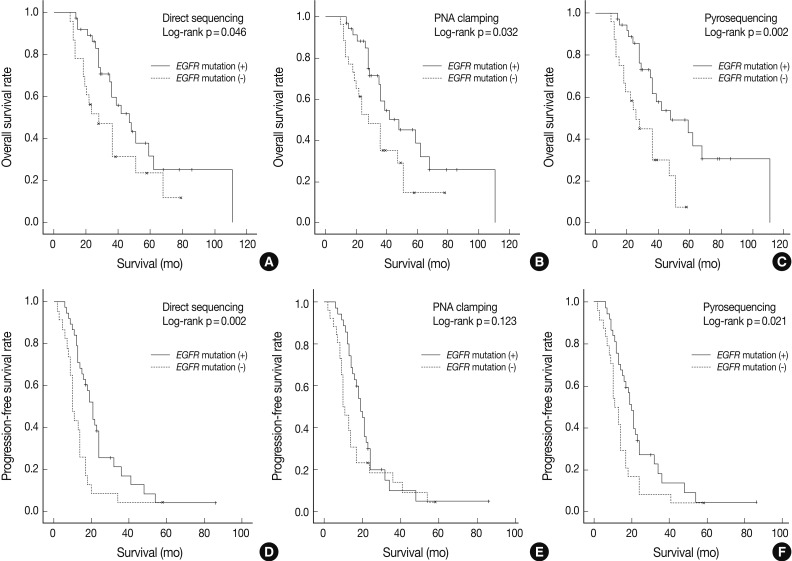
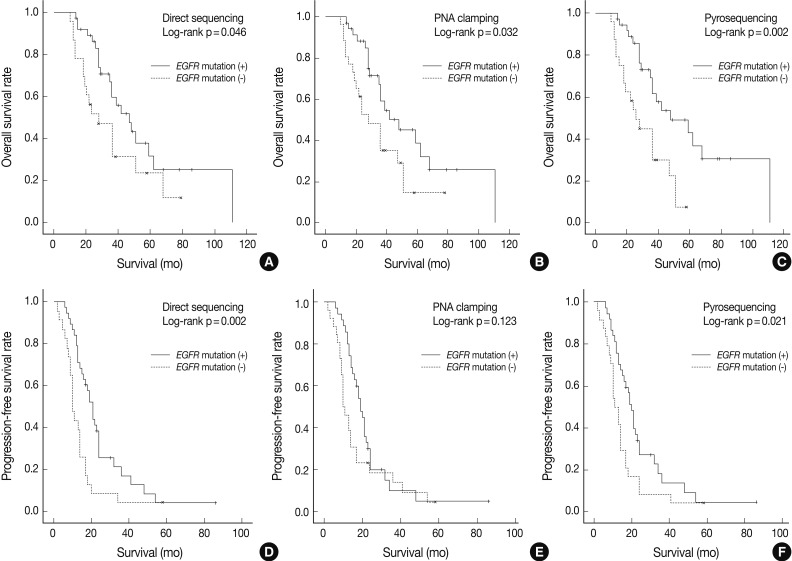
Fig. 1
| Case No. | Direct sequencing | PNA clamping | Pyrosequencing | TKI Response | Sex | Smoking | Histology |
|---|---|---|---|---|---|---|---|
| 1 | G719X, L861Q | G719X, L861Q | G719X | SD | F | N | ADC |
| 2 | Wild | Wild | Exon 19 del | PR | M | C | ASC |
| 3 | Wild | Wild | Exon 19 del | PD | F | N | ADC |
| 4 | Wild | G719X | Wild | SD | M | C | ADC |
| 5 | Wild | Exon 19 del | Exon 19 del | PR | M | C | ADC |
| 6 | Wild | S768I | Wild | PD | F | N | ADC |
| 7 | Wild | L858R | L858R | PR | M | N | ADC |
| 8 | Exon 19 del | Wild | Wild | PD | F | N | ADC |
| 9 | Exon 19 del | Wild | Wild | PD | M | FS | ASC |
| 10 | Exon 20 duplication | Wild | Wild | PD | F | N | ADC |
| 11 | R776H | Wild | Wild | PD | M | N | LCC |
| 12 | L858R | L858R | Wild | PD | M | N | ADC |
| 13 | L858R | Wild | L858R | PR | F | N | ADC |
| 14 | L858R | Wild | L858R | SD | F | N | ADC |
| 15 | L858R | Wild | L858R | PD | F | N | ADC |
| Case No. | Exon | Alteration | Direct sequencing |
TKI response | |
|---|---|---|---|---|---|
| Nucleotide alteration | Amino acid alteration | ||||
| 8 | 19 | Deletion | 2239-2263del | L747-755Adel | PD |
| 9 | 19 | Deletion | 2253-2276del | S752_I759del | PD |
| 10 | 20 | Duplication | dup 2311-2319 AACCCCCAC | D770_N771insNPH | PD |
| 11 | 20 | Point mutation | 2327G>A | R776H | PD |
| EGFR mutation | TKI response | Sex | Smoking | Histology |
|---|---|---|---|---|
| Exon19 (n = 18, 29%) | CR (n = 2,3%) | F | N | ADC |
| PR (n = 3) | F | N | ADC | |
| F | N | SCC | ||
| M | C | ADC | ||
| SD (n = 9) | F (n = 5) | N | ADC | |
| M (n = 2) | FS | ADC | ||
| M (n = 2) | C | ADC | ||
| PD (n = 5) | F (n = 3) | N | ADC | |
| M (n = 2) | C | ADC | ||
| Exon21 (n = 11, 18%) | PR (n = 5) | F (n = 3) | N | ADC |
| M | N | ADC | ||
| M | C | ADC | ||
| SD | M | C | ADC | |
| PD (n = 5) | F (n = 3) | N | ADC | |
| F (n = 2) | C | ADC | ||
| Wild (n = 17, 28%) | PR | M | FS | ADC |
| SD (n = 1,3%) | F | N | ADC | |
| PD (n = 15) | F (n = 4) | N | ADC | |
| F | N | SCC | ||
| F | C | ADC | ||
| F | C | LCNEC | ||
| M | N | SCC | ||
| M (n = 2) | FS | ADC | ||
| M | C | ADC | ||
| M (n = 2) | C | SCC | ||
| M (n = 2) | C | SarCa |
| n (%) | Direct sequencing EGFR mutation |
PNA clamping EGFR mutation |
Pyrosequencing EGFR mutation |
|||||||
|---|---|---|---|---|---|---|---|---|---|---|
| (+) | (-) | p-value | (+) | (-) | p-value | (+) | (-) | p-value | ||
| Total | 61 (100) | 38 (62) | 23 (38) | 35 (57) | 26 (43) | 37 (61) | 24 (39) | |||
| Sex | ||||||||||
| Female | 35 (57) | 25 (71) | 10 (29) | 0.113 | 21 (60) | 14 (40) | 0.794 | 24 (69) | 11 (31) | 0.188 |
| Male | 26 (43) | 13 (50) | 13 (50) | 14 (54) | 12 (46) | 13 (50) | 13 (50) | |||
| Smoking history |
||||||||||
| Never | 36 (59) | 26 (72) | 10 (28) | 0.066 |
22 (61) | 14 (39) | 0.600 |
24 (67) | 12 (33) | 0.294 |
| Former | 6 (10) | 3 (50) | 3 (50) | 2 (33) | 4 (67) | 2 (33) | 4 (67) | |||
| Current | 19 (31) | 9 (47) | 10 (53) | 11 (58) | 8 (42) | 11 (58) | 8 (42) | |||
| Histology | ||||||||||
| ADC | 50 (82) | 35 (70) | 15 (30) | 0.014 |
34 (68) | 16 (32) | <0.001 |
35 (70) | 15 (30) | 0.002 |
| SCC | 5 (8) | 1 (20) | 4 (80) | 1 (20) | 4 (80) | 1 (20) | 4 (80) | |||
| Others | 6 (10) | 2 (33) | 4 (67) | 0 | 6 (100) | 1 (17) | 5 (83) | |||
| Tumor size (cm) | ||||||||||
| ≤ 3 | 26 (43) | 21 (81) | 5 (19) | 0.016 | 19 (73) | 7 (27) | 0.040 | 21 (81) | 5 (19) | 0.008 |
| > 3 | 35 (57) | 17 (49) | 18 (51) | 16 (46) | 19 (54) | 16 (46) | 19 (54) | |||
| Operation method | ||||||||||
| Biopsy | 1 (2) | 1 (100) | 0 | 1.000 | 1 (100) | 0 | 1.000 | 1 (100) | 0 | 1.000 |
| Resection | 60 (98) | 37 (62) | 23 (38) | 34 (57) | 26 (43) | 36 (60) | 24 (40) | |||
| EGFR TKI response | ||||||||||
| CR | 1 (2) | 1 (100) | 0 | 0.537 |
1 (100) | 0 | 0.122 |
1 (100) | 0 | 0.005 |
| PR | 13 (21) | 9 (69) | 4 (31) | 10 (77) | 3 (23) | 12 (92) | 1 (8) | |||
| SD | 14 (23) | 12 (86) | 2 (14) | 12 (86) | 2 (14) | 12 (86) | 2 (14) | |||
| PD | 33 (54) | 16 (48) | 17 (52) | 12 (36) | 21 (64) | 12 (36) | 21 (64) | |||
| n (%) | Objective response |
p-value | Disease control rate |
p-value | Median TTP (mo) | p-value | MST (mo) | p-value | |
|---|---|---|---|---|---|---|---|---|---|
| Total | 61 (100) | 14 (23) | 28 (46) | 16.0 | 30.0 | ||||
| Sex | |||||||||
| Female | 35 (57) | 7 (20) | 0.553 | 15 (43) | 0.613 | 18.0 | 0.440 | 29.0 | 0.445 |
| Male | 26 (43) | 7 (27) | 13 (50) | 13.5 | 35.0 | ||||
| Smoking history |
|||||||||
| Never | 36 (59) | 9 (25) | 0.762 |
17 (47) | 1.000 |
17.0 | 0.300 |
32.0 | 0.795 |
| Former | 6 (10) | 1 (17) | 3 (50) | 19.5 | 44.5 | ||||
| Current | 19 (31) | 4 (21) | 8 (42) | 11.0 | 28.0 | ||||
| Histology | |||||||||
| ADC | 50 (82) | 12 (24) | 0.726 |
26 (52) | 0.051 |
17.5 | 0.043 |
30.0 | 0.100 |
| SCC | 5 (8) | 1 (20) | 1 (20) | 11.0 | 28.0 | ||||
| Others | 6 (10) | 1 (17) | 1 (17) | 10.0 | 28.0 | ||||
| Direct sequencing | |||||||||
| EGFR mutation (+) | 38 (62) | 10 (26) | 0.537 | 22 (58) | 0.019 | 20.0 | 0.008 | 34.5 | 0.428 |
| EGFR mutation (-) | 23 (38) | 4 (17) | 6 (26) | 10.0 | 24.0 | ||||
| PNA clamping | |||||||||
| EGFR mutation (+) | 35 (57) | 11 (31) | 0.122 | 23 (66) | 0.001 | 19.0 | 0.020 | 34.0 | 0.606 |
| EGFR mutation (-) | 26 (43) | 3 (12) | 5 (23) | 10.5 | 24.0 | ||||
| Pyrosequencing | |||||||||
| EGFR mutation (+) | 37 (61) | 13 (35) | 0.005 | 25 (68) | <0.001 | 19.0 | 0.018 | 35.0 | 0.294 |
| EGFR mutation (-) | 24 (39) | 1 (4) | 3 (13) | 12.0 | 25.0 |
Never smokers were defined as patients who had a lifetime smoking exposure of <100 cigarettes and former smokers were defined as patients who had stopped smoking at least 1 yr before diagnosis; Comparison between never smokers and others; Comparison between adenocarcinoma and nonadenocarcinoma; Comparison between CR, PR and SD, PD.
Objective response: complete response or partial response; Disease control rate: complete response or partial response or stable disease; Never smokers were defined as patients who had a lifetime smoking exposure of <100 cigarettes and former smokers were defined as patients who had stopped smoking at least 1 yr before diagnosis; Comparison between never smokers and others; Comparison between adenocarcinoma and nonadenocarcinoma.

 E-submission
E-submission