Articles
- Page Path
- HOME > J Pathol Transl Med > Volume 47(3); 2013 > Article
-
Original Article
Cytological Evaluation and REBA HPV-ID HPV Testing of Newly Developed Liquid-Based Cytology, EASYPREP: Comparison with SurePath - Youn Soo Lee, Gyungyub Gong1, Jin Hee Sohn2, Ki Sung Ryu3, Jung Hun Lee4, Shin Kwang Khang1, Kyung-Ja Cho1, Yong-Man Kim5, Chang Suk Kang
-
Korean Journal of Pathology 2013;47(3):265-274.
DOI: https://doi.org/10.4132/KoreanJPathol.2013.47.3.265
Published online: June 25, 2013
Department of Hospital Pathology, The Catholic University of Korea College of Medicine, Seoul, Korea.
1Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea.
2Department of Pathology, Kangbuk Samsung Hospital, Sungkyunkwan University School of Medicine, Seoul, Korea.
3Department of Obstetrics and Gynecology, The Catholic University of Korea College of Medicine, Seoul, Korea.
4Department of Obstetrics and Gynecology, Kangbuk Samsung Hospital, Sungkyunkwan University School of Medicine, Seoul, Korea.
5Department of Gynecology and Obstetrics, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea.
- Corresponding Author: Chang Suk Kang, M.D. Department of Hospital Pathology, The Catholic University of Korea College of Medicine, 10 63-ro, Yeongdeungpo-gu, Seoul 150-713, Korea. Tel: +82-2-3779-1312, Fax: +82-2-783-6648, cskang@catholic.ac.kr
© 2013 The Korean Society of Pathologists/The Korean Society for Cytopathology
This is an Open Access article distributed under the terms of the Creative Commons Attribution Non-Commercial License (http://creativecommons.org/licenses/by-nc/3.0/) which permits unrestricted non-commercial use, distribution, and reproduction in any medium, provided the original work is properly cited.
Abstract
-
Background
- The objective of this study was to evaluate a newly-developed EASYPREP liquid-based cytology method in cervicovaginal specimens and compare it with SurePath.
-
Methods
- Cervicovaginal specimens were prospectively collected from 1,000 patients with EASYPREP and SurePath. The specimens were first collected by brushing for SurePath and second for EASYPREP. The specimens of both methods were diagnosed according to the Bethesda System. Additionally, we performed to REBA HPV-ID genotyping and sequencing analysis for human papillomavirus (HPV) on 249 specimens.
-
Results
- EASYPREP and SurePath showed even distribution of cells and were equal in cellularity and staining quality. The diagnostic agreement between the two methods was 96.5%. Based on the standard of SurePath, the sensitivity, specificity, positive predictive value, and negative predictive value of EASYPREP were 90.7%, 99.2%, 94.8%, and 98.5%, respectively. The positivity of REBA HPV-ID was 49.4% and 95.1% in normal and abnormal cytological samples, respectively. The result of REBA HPV-ID had high concordance with sequencing analysis.
-
Conclusions
- EASYPREP provided comparable results to SurePath in the diagnosis and staining quality of cytology examinations and in HPV testing with REBA HPV-ID. EASYPREP could be another LBC method choice for the cervicovaginal specimens. Additionally, REBA HPV-ID may be a useful method for HPV genotyping.
- Subjects and specimen collection
- The cervicovaginal cytology specimens were obtained from 1,000 patients with their informed consent at three hospitals between May 1 and August 31, 2012 after approval from the Institutional Review Board. The specimens were prospectively collected with the EASYPREP and SurePath methods at the same time for comparison. Each cervicovaginal sample was obtained using cervexbrush (SurePath) and cytobrush (EASYPREP). Initially, samples from the first brushing were placed into SurePath preservation solution and then samples from the second brushing were placed into EASYPREP preservation solution. The cytology specimens of both EASYPREP and SurePath methods were screened by three cytotechnologists and were subsequently diagnosed by two pathologists. For both the EASYPREP and SurePath methods, the cytological diagnosis was determined according to The Bethesda System with access to other relevant patient background information. The diagnoses of sensitivity, specificity, positive predictive value (PPV), negative predictive value (NPV) and smear characteristics using the EASYPREP method were compared with the results of the SurePath method.
- EASYPREP process
- The specimen vials were processed in the automated smearing system following the manufacturer's instructions. All preparation procedures for LBC were processed automatically in the EASYPREP after vortexing and then transferred onto EASYPREP. In the EASYPREP 5 mL of gradient density reagent and 5 mL of preserved specimen were dispensed successively into the centrifugal tube, centrifuged for 3 minutes at 1,000 rpm and the non-diagnostic debris in the supernatant (including mucus, red blood cells and excess inflammatory cells from the sample) were removed by aspiration. The sample was then centrifuged for 7 minutes at 2,000 rpm and the residual gradient density reagent removed by aspiration. The pelleted cells were resuspended with EASYPREP suspension buffer, mixed and transferred to an EASYPREP slide chamber mounted on the EASYPREP slide. The cells were sedimented by gravity and attached to the slide by electronic charge for 10-20 minutes. After cells were completely attached onto the slide, excessive smeared cells were removed by repeating the washing step 3-4 times. Staining and coverslipping of specimen slides were performed according to laboratory procedures, and slides were examined under a microscope by trained cytotechnologists and pathologists.
- Reverse blot hybridization assay for HPV DNA using EASYPREP
- EASYPREP cell samples were preserved at room temperature. Thereafter, 1 mL of specimen from the remnant cell suspension was used to extract HPV viral DNA within a week after preparing specimens for diagnosis.
- REBA HPV-ID is a system for HPV genotyping that can identify the 18 high-risk groups (HPV genotypes 16, 26, 18, 31, 33, 35, 39, 45, 51, 52, 53, 56, 58, 59, 66, 68, 69, 73), 1 probable high-risk group (HPV genotype 34), and the 13 low-risk groups (HPV genotypes 6, 11, 40 42, 43, 44, 54, 70, 72, 81, 84, 87). DNA preparation was performed using the supplied DNA extraction solution according to the manufacturer's instructions. Specimens of 1 mL were centrifuged at 13,000 rpm and 4℃ for 5 minutes and the supernatant was discarded. Sterile distilled water (1.2 mL) was used to adjust the pellet volume. After vortexing, the homogenate was centrifuged at 13,000 rpm and 4℃ for 5 minutes and the supernatant was discarded. The resulting pellet was washed 2-3 times by homogenization as described above. The DNA extraction solution (50 µL; supplied by YD Diagnostics Corp.) was added to the pellet. The sample was immediately boiled at 100℃ for 10 minutes on a heat block and centrifuged at 17,590×g for 3 minutes (25℃). The resulting supernatant was transferred to a new 1.5 mL tube. A sample 3-5 µL of the supernatant was taken to be used as a template in the simultaneous PCR amplification of the HPV and β-globin gene. For HPV genotyping, PCR products were denatured by the addition of a denaturation solution after PCR amplification of the target region. The denatured DNA was added into each well in order to adjust the supplied membrane strip onto a provided blotting tray with hybridization solution and hybridization was processed at the appropriate temperature for 30 minutes. The hybridization solution was removed and the membrane was washed twice in a pre-heated washing solution. The alkaline-phosphatase enzyme reaction was performed for 30 minutes and chromogenic reaction for 10 minutes until the color was detected. The membrane was washed twice with distilled water and completely dried. The developed strips were placed onto the data sheet provided.
- Immunocytochemistry
- All EASYPREP slides for immunocytochemical analysis were processed with the EASYPREP processor according to the manufacturer's instructions. For immunocytochemical staining, slides were first fixed with spray fixative and then rinsed in 50% ethanol for 30 minutes. Antigen retrieval was first conducted for 20 minutes at 99℃ by heating with the microwave processor. Slides remained in the Coplin jar with distilled water at room temperature for 30 minutes. After blocking endogenous peroxidase activity, the slides were incubated with primary antibody for 30 minutes. Immunostaining with p63 (1:200, Zeta, Sierra Madre, CA, USA) and Ki-67 (1:50, MIB-1, Dako, Glostrup, Denmark) was performed using the standard biotin streptavidin detection system (Dako, Carpinteria, CA, USA). 3,3-Diaminobenzidine was used as the chromogen and the slides were counter-stained with hematoxylin. All stains were performed on the Dako Autostainer.
- Sequencing analysis
- Sequencing analysis was performed for all specimens that were examined by REBA HPV-ID. For sequencing analysis, nested PCR was performed to amplify the target regions (GP5 and GP6).
- Statistical analysis
- Diagnosis between EASYPREP and SurePath methods were compared by the Pearson chi-square test. All p-values were 2-tailed; p<0.05 were considered statistically significant. Analyses were conducted using SPSS ver. 13.0 (SPSS Inc., Chicago, IL, USA).
MATERIALS AND METHODS
- Smear characteristics
- SurePath and EASYPREP thin layer slides were very similar in their overall appearance in the 17 mm in diameter round observation field of EASYPREP and 11 mm observation field of SurePath. The gross areas of EASYPREP and SurePath were 213.7 mm2 and 103.8 mm2, respectively. EASYPREP was 2.1 times larger than SurePath. Both preparations displayed a uniform layer of cells across their respective cellular areas. Mucus and red cells were removed, and the number of leukocytes was reduced or distributed randomly throughout the preparation. The individual cells showed even staining intensity. Air-dried artifacts and obscuring and overlapping cellular material and debris were largely eliminated. The inflammatory cells did not overlap with the epithelial cells, which allowed for easier visualization of epithelial cells, diagnostically relevant cells and infectious organisms. The red blood cells were cytolyzed at the smear specimens. Lactobacilli were confirmed outside of the cells as well as in the cytoplasm of epithelial cells. Candida infections were detected in 0.8% of specimens. There were no significant differences between EASYPREP and SurePath in the quality of preservation, cellularity, infectious organisms and specimen adequacy. EASYPREP and SurePath methods provided evenly distributed thin layers of cells. Total smeared cells of EASYPREP were calculated to average 129,585. The total squamous cells of SurePath were 78,406 on average. The amount of smeared cells of EASYPREP was 1.7 times higher than SurePath. The cell density (643 cells) under the 200× magnification field of EASYPREP was 1.2 times lower than that of SurePath (800 cells). Cellularity and staining quality were the same in the EASYPREP and SurePath methods.
- Diagnosis
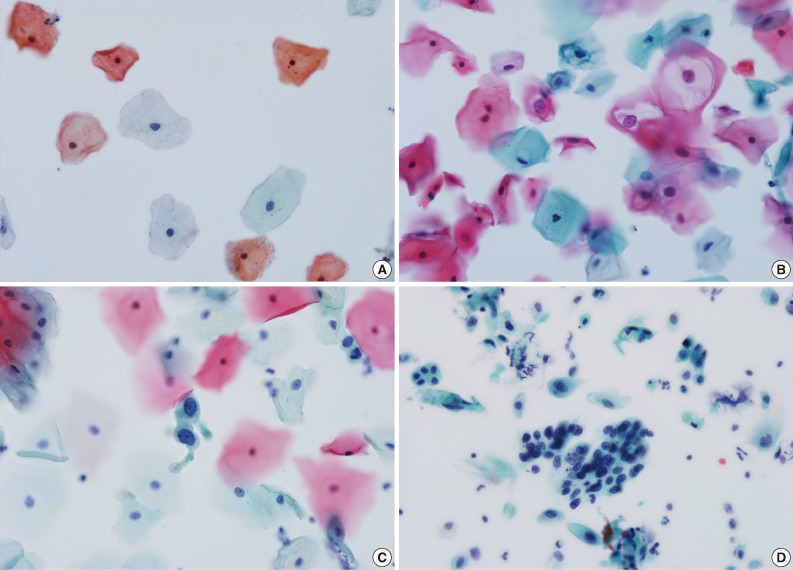
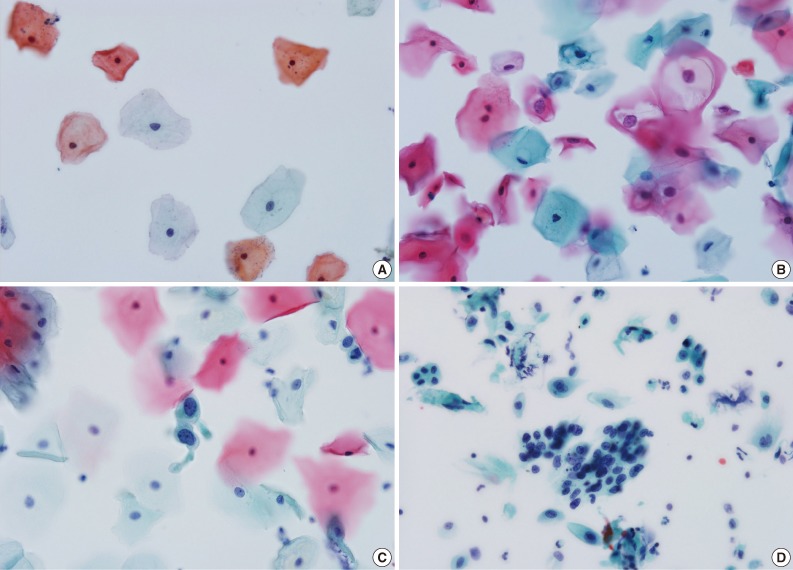
- Table 1 presents the results of cytological diagnosis performed on 1,000 cases using EASYPREP and SurePath methods. The mean age was 45±2.5 years. The diagnosis results using the EASYPREP technique were negative for intraepithelial lesion or malignancy (NILM) in 86.6% of cases, atypical squamous cells of undetermined significance (ASC-US) in 4.3%, atypical squamous cells, cannot exclude high-grade squamous intraepithelial lesion (ASC-H) in 0.6%, low grade squamous intraepithelial lesion (LSIL) in 5.9% and high grade squamous intraepithelial lesion (HSIL) in 2.0% of cases. The squamous cell carcinoma (SCC) was detected in 0.4%. The atypical glandular cells of undetermined significance (AGUS) and adenocarcinoma were detected in 0.1%, respectively. The cytological appearance for each diagnosis when using EASYPREP is presented in Fig. 1. The proportion of diagnostic results did not differ significantly between the two methods (p=0.087). The negative rate (86.6%) was slightly higher in the EASYPREP method and the detection rate of abnormal cells (14.0%) was slightly higher in the SurePath method. Table 2 shows diagnostic agreement between the EASYPREP and SurePath methods. Based on the standard of the SurePath method, the sensitivity, specificity, PPV, and NPV of EASYPREP were 90.7%, 99.2%, 94.8%, and 98.5%, respectively. The concordance rate of diagnosis was 96.5% in total, 99.2% in NILM, 80.0% in abnormal diagnosis cases more than ASC-US and 92.9% in abnormal diagnosis cases more than LSIL. The concordance of diagnosis with the two methods is presented in Table 3. Although the numbers are small, the 2 methods agreed exactly in AGUS and adenocarcinoma (1 case, 0.1%, respectively). Among the 860 cases of NILM diagnosed using SurePath, five cases were reported as ASC-US, one case as LSIL and one case as HSIL when using EASYPREP. Among 48 cases interpreted as ASC-US using SurePath, nine were reported as NILM, one as ACS-H and four as LSIL. Eight cases were interpreted as ASC-H when using SurePath, two of which were reported as NILM and two of which were reported as HSIL. A total of 59 cases were interpreted as LSIL when using SurePath, one of which was reported as NILM, four as ACS-US and one as HSIL. Among the 17 cases interpreted as HSIL when using SurePath, one was reported as ASC-H and one as LSIL. Six cases were interpreted as SCC when using SurePath, one of which was reported as NILM and one as HSIL. There was no inadequate specimen in either the EASYPREP or SurePath.
- Immunohistochemistry
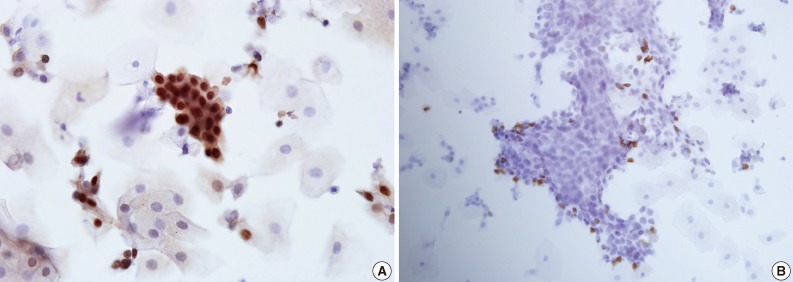
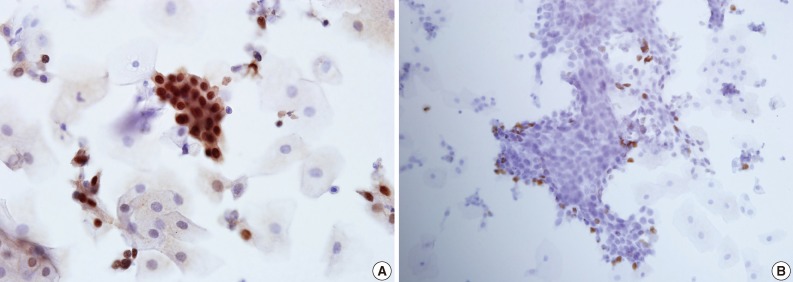
- In all 5 different cases, dysplastic cells arranged in groups and reactive squamous cells demonstrated positive nuclear staining for p63 and Ki-67 (Fig. 2).
- Reverse blot hybridization assay for HPV DNA genotyping using EASYPREP samples
- HPV genotyping was performed on 249 specimens examined cytologically using the REBA HPV-ID. Tables 4 and 5 show the results of REBA-HPV DNA genotyping using EASYPREP samples. Out of 249 cases, 123 (49.4%) cases were HPV positive and 126 (50.6%) cases were HPV negative. The positivity rate of the REBA HPV-ID for 41 abnormal cytological samples was 95.1%. The positivity rate of the REBA HPV-ID was 100.0% for SCC, 90.0% for HSIL, 100.0% for LSIL, 83.3% for ASC-H, and 100.0% for ASC-US. Based on the standard of REBA HPV-ID genotyping using SurePath samples, the sensitivity, specificity, PPV, and NPV of REBA HPV-ID genotyping using EASYPREP samples were 95.1%, 59.6%, 31.7%, and 98.4%, respectively. The 123 positive cases consisted of 84 cases of NILM, 12 of ASC-US, 5 of ASC-H, 9 of LSIL, 9 of HSIL, and 4 of SCC. The REBA HPV-ID positivity rate with normal cytological samples was 40.4% (84/208). Forty-nine (58.3%) cases had a previous clinical history including LSIL, HSIL and invasive cancer among the 84 normal cytology samples with HPV positivity. Among the patients who were HPV positive (123 cases), 44.7% (55/123) had single type infections and 55.3% had multiple HPV types (data not shown). HPV-16 was the most prevalent genotype observed in 22 cases (8.8%). The five most common HPV genotypes detected using REBA HPV-ID were HPV 16, 52, 18, 33, and 53 which were found in specimens that had an abnormal cytological diagnosis. The five most prevalent HPV genotypes detected using the REBA HPV-ID in the specimens with normal cytology were HPV 18, 54, 32, 66, and 45 (Table 5).
- Sequencing analysis for HPV using EASYPREP samples
- Sequencing analysis was performed to determine the accuracy and reliability of REBA HPV-ID with all specimens (249 cases) that were tested by REBV HPV-ID. The detection rate of HPV with sequencing analysis wa s 46.6% (116/249) (Table 6). The positivity rate of HPV with sequencing analysis for 41 abnormal cytological samples was 95.1%. The positivity rate of HPV with sequencing analysis was 100.0% for SCC, 90.0% for HSIL, 100.0% for LSIL, 83.3% for ASC-H, and 100.0% for ASC-US. HPV-16 was the most prevalent genotype observed in 18 (7.2%) cases. The HPV positivity rate with sequencing analysis with normal cytological samples was 37.0% (77/208). Based on the standard of the SurePath method, the sensitivity, specificity, PPV, and NPV of sequencing analysis were 95.1%, 63.0%, 33.6%, and 98.5%, respectively. The 116 positive cases consisted of 77 cases of NILM, 12 of ASC-US, 5 of ASC-H, 9 of LSIL, 9 of HSIL, and 4 of SCC (Table 6). The REBV HPV-ID had high concordance with sequencing analysis as considered by the gold standard of molecular diagnostics. As shown in Table 7, the sensitivity, specificity, PPV, and NPV of REBA HPV-ID except for specimens of 22 mixed infections in the sequencing analysis were 100.0%, 94.7%, 93.1%, and 100.0%, respectively. Seven samples could not be interpreted by sequencing analysis because they were not amplified using HPV-specific primer pairs. The cytological diagnosis of these samples was NILM. Table 8 shows the type-specific HPV results of REBA HPV-ID and sequencing analysis.
RESULTS
- In this study, we analyzed the cytological characteristics using the EASYPREP method, a newly developed LBC technique and performed HPV DNA genotyping using residual cell suspensions.
- EASYPREP is the world's first fully automated thin layer cell preparation processor using centrifugation. The entire operation process of the system is controlled automatically and is displayed to the user by the monitor. The automated smearing processors generally use the filter-membrane method. However, the filtering method is problematic when the physical morphology of cells changes while adhering to the slide, and in the loss of cells not transferring to the slide. Instead of using a filter, EASYPREP uses the centrifugal sedimentation and specific gravity which partially remove non-diagnostic debris (including mucus and red blood cells) and excess inflammatory cells from the sample.
- SurePath also utilizes centrifugation but is a semi-automated processor in which preparation process is performed manually and can cause differences in slide quality depending on the operators.10 Because EASYPREP is a fully automated walk-away processor, the operator does not have to be concerned about the slide quality after placing the consumables and specimens in the processor. Furthermore various types of processing software are installed in EASYPREP in accordance with the specimen characteristics allowing for objective cell preparation, which is an advantage for maintaining quality control. Moreover EASYPREP's sample loading capacity is 64 specimens. A characteristic of the EASYPREP method is the composition of the cytology-specific preservation vial, which can be directly used as a centrifuge tube. The surface of the glass slide is pretreated in order to have a positive charge for optimal cell adhesion.
- In this study, the cells processed by the EASYPREP method were evenly distributed in a monolayer without overlapping at the observation field. There were no significant differences in cellular distribution in the EASYPREP and SurePath methods. The gross areas of EASYPREP and SurePath were 213.7 mm2 and 103.8 mm2, respectively, EASYPREP was 2.1 times larger than SurePath. The total smeared cells of EASYPREP and SurePath were calculated as 129,585 and 78,406 on average, respectively. Because the total smeared cell count of EASYPREP was 1.6 times higher than SurePath, the possibility to detect abnormal cells could be high.11 The cell density (643 cells) under the 200× magnification field of EASYPREP was 1.2 times lower than of SurePath (800 cells). Although there was no significant statistical difference, the cell density of the EASYPREP method was less than the SurePath method due to the larger circle of smeared area in the EASYPREP method.
- The proportion of diagnostic results using the EASYPREP method did not differ significantly from the SurePath method. The negative rate (86.6%) was slightly higher in the EASYPREP method than in SurePath (86.0%). The detection rate of abnormal cells (14.0%) was slightly higher in the SurePath method than EASYPREP method (13.4%). However, there were five cases of ASC-US, one case of LSIL and one case of HSIL when using EASYPREP among the 860 cases of NILM in SurePath. The patient diagnosed as LSIL in EASYPREP and as NILM in SurePath did not perform a follow up cervicovaginal cytology examination and biopsy. One case diagnosed as HSIL in EASYPREP and as NILM in SurePath was diagnosed as cervical intraepithelial neoplasia 2 by punch biopsy.
- Additionally, there were 11 cases diagnosed as ASC-US or ASC-H, one case as LSIL and one case as SCC in SurePath among the 866 cases of NILM in EASYPREP. One case diagnosed as LSIL in SurePath and as NILM in EASYPREP had positive HPV testing results and NILM on five follow-up cervicovaginal cytology examinations without biopsy. One case diagnosed as SCC in SurePath and as NILM in EASYPREP was diagnosed as SCC by punch biopsy. The authors detected necrotic materials only without atypical squamous cells on review of the EASYPREP slide.
- Based on the standard of the SurePath method, the sensitivity, specificity, PPV, and NPV of the EASYPREP method were 90.7%, 99.2%, 94.8%, and 98.5%, respectively. The concordance rate of diagnosis was 96.5% in total and 99.2% in NILM, 80.0% in abnormal diagnosis cases more than ASC-US and 92.9% in abnormal diagnosis cases more than LSIL. The disagreement of diagnosis between the two methods and the slightly low sensitivity of the EASYPREP method was possibly due to bias in specimen collection process in this study.
- Several reports regarding LBC showed that sensitivity in detecting abnormal squamous and glandular lesion was better than conventional cytology procedures due to even distribution and staining quality and short observation time on the restricted field.2-5 Additionally, the number of unsatisfactory specimens was decreased. However, several reports showed similar performance characteristics between conventional cytology and LBC.12-14 Nevertheless, LBC has advantages for cervical smear readers with an easy microscopic examination due to short observation time and even distribution.15
- Moreover, the same EASYPREP specimen was used to assess additional immunocytochemistry. The positive nuclear staining for Ki-67 and p63 was successfully detected in this study, showing the EASYPREP specimen can be used for immunocytochemistry.
- The merit of LBC is that the samples used for LBC can be used in HPV testing.7 In this study, HPV genotyping using EASYPREP samples was performed by REBA HPV-ID. The positivity rate of the REBA HPV-ID was 49.4% in total and 95.1% in abnormal cytology samples. Using REBA HPV-ID in the current study, the prevalence of HPV was found in patients with ASC-US (100.0%), ASC-H (83.3%), LSIL (100.0%), HSIL (90.0%), and SCC (100.0%). The REBA HPV-ID positivity rate with normal cytology samples was 40.4%. Forty-nine (58.3%) cases among the normal cytology samples with HPV positivity had previous clinical history including LSIL, HSIL, and invasive cancer. In this study, the REBA HPV-ID positive rate was lower than previously reported (49.4% vs 72.8%), but higher in abnormal cytology samples (95.1% vs 80.9%).16 Subsequent sequencing analysis was performed and confirmed that the specimens which were positive by REBA HPV-ID did contain HPV sequences. The detection rate (46.6%) of HPV with sequencing analysis was lower than REBA HPV-ID (49.4%). The REBA HPV-ID had high concordance with sequencing analysis as considered by the gold standard of molecular diagnostics. Based on the standard of sequencing analysis, the sensitivity, specificity, PPV, and NPV of REBA HPV-ID (except for 22 mixed infection specimens) in sequencing analysis were 100.0%, 94.7%, 93.1%, and 100.0%, respectively. The performance results of the REBA HPV-ID were more sensitive and specific than other HPV genotype testing in terms of detection rates for samples with abnormal and normal cytology.16
- In conclusion, the EASYPREP method provided comparable results to the SurePath method in diagnosis and staining quality and the samples could be used for immunocytochemistry and HPV testing. The EASYPREP method is a feasible cytological procedure and could be another choice of LBC method for the cervicovaginal specimens. The REBA HPV-ID might be a very useful diagnostic method for HPV genotyping using EASYPREP residual samples.
DISCUSSION
- 1. National Cancer Statistics, Seoul, Korea [Internet]. Goyang: National Cancer Information Center, 2012; cited Jun 1. Available from: http://www.cancer.go.kr/ncic/.
- 2. Doyle B, O'Farrell C, Mahoney E, Turner L, Magee D, Gibbons D. Liquid-based cytology improves productivity in cervical cytology screening. Cytopathology 2006; 17: 60-64. ArticlePubMed
- 3. Strander B, Andersson-Ellström A, Milsom I, Rådberg T, Ryd W. Liquid-based cytology versus conventional Papanicolaou smear in an organized screening program: a prospective randomized study. Cancer 2007; 111: 285-291. ArticlePubMed
- 4. Schledermann D, Ejersbo D, Hoelund B. Improvement of diagnostic accuracy and screening conditions with liquid-based cytology. Diagn Cytopathol 2006; 34: 780-785. ArticlePubMed
- 5. Siebers AG, Klinkhamer PJ, Grefte JM, et al. Comparison of liquid-based cytology with conventional cytology for detection of cervical cancer precursors: a randomized controlled trial. JAMA 2009; 302: 1757-1764. ArticlePubMed
- 6. Sahebali S, Depuydt CE, Boulet GA, et al. Immunocytochemistry in liquid-based cervical cytology: analysis of clinical use following a cross-sectional study. Int J Cancer 2006; 118: 1254-1260. ArticlePubMed
- 7. Jamison J, Wilson RT, Carson J. The evaluation of human papillomavirus genotyping in cervical liquid-based cytology specimens: using the Roche Linear Array HPV genotyping assay. Cytopathology 2009; 20: 242-248. ArticlePubMed
- 8. Renshaw AA, Young NA, Birdsong GG, et al. Comparison of performance of conventional and ThinPrep gynecologic preparations in the College of American Pathologists Gynecologic Cytology Program. Arch Pathol Lab Med 2004; 128: 17-22. ArticlePubMedPDF
- 9. Fremont-Smith M, Marino J, Griffin B, Spencer L, Bolick D. Comparison of the SurePath liquid-based Papanicolaou smear with the conventional Papanicolaou smear in a multisite direct-to-vial study. Cancer 2004; 102: 269-279. ArticlePubMed
- 10. Hoda RS, Loukeris K, Abdul-Karim FW. Gynecologic cytology on conventional and liquid-based preparations: a comprehensive review of similarities and differences. Diagn Cytopathol 2013; 41: 257-278. ArticlePubMed
- 11. Studeman KD, Ioffe OB, Puszkiewicz J, Sauvegeot J, Henry MR. Effect of cellularity on the sensitivity of detecting squamous lesions in liquid-based cervical cytology. Acta Cytol 2003; 47: 605-610. ArticlePubMed
- 12. Shin E, Park JK, Park NW, et al. Cervical cytologic smears in Pap solution vs ThinPrep: smear characteristics and diagnostic agreement. Korean J Pathol 2011; 45: 621-625. Article
- 13. Arbyn M, Bergeron C, Klinkhamer P, Martin-Hirsch P, Siebers AG, Bulten J. Liquid compared with conventional cervical cytology: a systematic review and meta-analysis. Obstet Gynecol 2008; 111: 167-177. PubMed
- 14. Siebers AG, Klinkhamer PJ, Grefte JM, et al. Comparison of liquid-based cytology with conventional cytology for detection of cervical cancer precursors: a randomized controlled trial. JAMA 2009; 302: 1757-1764. ArticlePubMed
- 15. Dowie R, Stoykova B, Crawford D, et al. Liquid-based cytology can improve efficiency of cervical smear readers: evidence from timing surveys in two NHS cytology laboratories. Cytopathology 2006; 17: 65-72. ArticlePubMed
- 16. Kim S, Lee D, Park S, et al. REBA HPV-ID® for efficient genotyping of human papillomavirus in clinical samples from Korean patients. J Med Virol 2012; 84: 1248-1253. ArticlePubMed
REFERENCES
Values are presented as number (%).
NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS | AC | Total | |
|---|---|---|---|---|---|---|---|---|---|
| SurePath | 860 | 48 | 8 | 59 | 17 | 6 | 1 | 1 | 1,000 |
| Concordant cases in EASYPREP | 853 | 34 | 4 | 53 | 15 | 4 | 1 | 1 | 965 |
| Concordance rate (%) | 99.2 | 70.8 | 50.0 | 89.8 | 88.2 | 66.7 | 100 | 100 | 96.5 |
NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma; PPV, positive predictive value; NPV, negative predictive value.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma; PPV, positive predictive value; NPV, negative predictive value.
|
Reference standard (sequencing analysis) |
||||||
|---|---|---|---|---|---|---|
| HPV (+) | HPV (-) | Total | PPV | NPV | ||
| REBA HPV-ID | HPV (+) | 94 | 7 | 101 | 93.1% | - |
| HPV (-) | 0 | 126 | 126 | - | 100.0% | |
| Total | 94 | 133 | 227 | - | - | |
| Sensitivity 100.0% | Specificity 94.7% | - | - | - | ||
Figure & Data
References
Citations

- Virome capture sequencing for comprehensive HPV genotyping in cervical samples
Thanayod Sasivimolrattana, Sasiprapa Liewchalermwong, Wasun Chantratita, Insee Sensorn, Arkom Chaiwongkot, Parvapan Bhattarakosol
Science Progress.2025;[Epub] CrossRef - High-Risk Human Papillomavirus Detection via Cobas® 4800 and REBA HPV-ID® Assays
Sasiprapa Liewchalermwong, Shina Oranratanaphan, Wichai Termrungruanglert, Surang Triratanachat, Patou Tantbirojn, Nakarin Kitkumthorn, Parvapan Bhattarakosol, Arkom Chaiwongkot
Viruses.2022; 14(12): 2713. CrossRef - Evaluation of nuclear chromatin using grayscale intensity and thresholded percentage area in liquid‐based cervical cytology
Hyekyung Lee, Myungein Han, Taejo Yoo, Chanho Jung, Hyun‐Jin Son, Migyung Cho
Diagnostic Cytopathology.2018; 46(5): 384. CrossRef - Comparison of EASYPREP® and SurePath® in thyroid fine‐needle aspiration
Yosep Chong, Ki Hyun Baek, Jee Young Kim, Tae‐Jung Kim, Eun Jung Lee, Chang Suk Kang
Diagnostic Cytopathology.2016; 44(4): 283. CrossRef
 PubReader
PubReader ePub Link
ePub Link-
 Cite this Article
Cite this Article
- Cite this Article
-
- Close
- Download Citation
- Close
- Figure


Fig. 1
Fig. 2
| SurePath | EASYPREP | |
|---|---|---|
| NILM | 860 (86.0) | 866 (86.6) |
| ASC-US | 48 (4.8) | 43 (4.3) |
| ASC-H | 8 (0.8) | 6 (0.6) |
| LSIL | 59 (5.9) | 59 (5.9) |
| HSIL | 17 (1.7) | 20 (2.0) |
| SCC | 6 (0.6) | 4 (0.4) |
| AGUS | 1 (0.1) | 1 (0.1) |
| AC | 1 (0.1) | 1 (0.1) |
| 1,000 | 1,000 |
| EASYPREP |
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS | AC | Total | ||
| SurePath | NILM | 853 | 5 | 0 | 1 | 1 | 0 | 0 | 0 | 860 |
| ASC-US | 9 | 34 | 1 | 4 | 0 | 0 | 0 | 0 | 48 | |
| ASC-H | 2 | 0 | 4 | 0 | 2 | 0 | 0 | 0 | 8 | |
| LSIL | 1 | 4 | 0 | 53 | 1 | 0 | 0 | 0 | 59 | |
| HSIL | 0 | 0 | 1 | 1 | 15 | 0 | 0 | 0 | 17 | |
| SCC | 1 | 0 | 0 | 0 | 1 | 4 | 0 | 0 | 6 | |
| AGUS | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0 | 1 | |
| AC | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 1 | |
| Total | 866 | 43 | 6 | 59 | 20 | 4 | 1 | 1 | 1,000 | |
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS | AC | Total | |
|---|---|---|---|---|---|---|---|---|---|
| SurePath | 860 | 48 | 8 | 59 | 17 | 6 | 1 | 1 | 1,000 |
| Concordant cases in EASYPREP | 853 | 34 | 4 | 53 | 15 | 4 | 1 | 1 | 965 |
| Concordance rate (%) | 99.2 | 70.8 | 50.0 | 89.8 | 88.2 | 66.7 | 100 | 100 | 96.5 |
| REBA HPV-ID | SurePath |
PPV (%) | NPV (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS/AC | Total | |||
| HPV (+) | 84 | 12 | 5 | 9 | 9 | 4 | 0 | 123 | 31.7 (39/123) | 98.4 (124/126) |
| HPV (-) | 124 | 0 | 1 | 0 | 1 | 0 | 0 | 126 | ||
| Total | 208 | 12 | 6 | 9 | 10 | 4 | 0 | 249 | ||
| HPV (+) rate (%) | 40.4 | 100.0 | 83.3 | 100.0 | 90.0 | 100.0 | 0.0 | 49.4 | ||
| REBA HPV-ID | HPV genotype | SurePath |
Total | Prevalence (%) (n=249) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS/AC | ||||
| High-risk | 16 | 6 | 1 | 4 | 1 | 6 | 4 | 0 | 22 | 8.8 |
| 18 | 13 | 3 | 1 | 0 | 0 | 0 | 0 | 17 | 6.8 | |
| 26 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.0 | |
| 31 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.4 | |
| 33 | 0 | 0 | 0 | 1 | 3 | 0 | 0 | 4 | 1.6 | |
| 35 | 1 | 0 | 0 | 1 | 0 | 0 | 0 | 2 | 0.8 | |
| 39 | 4 | 0 | 0 | 3 | 0 | 0 | 0 | 7 | 2.8 | |
| 45 | 9 | 0 | 0 | 0 | 0 | 0 | 0 | 9 | 3.6 | |
| 51 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.0 | |
| 52 | 6 | 4 | 0 | 1 | 0 | 0 | 0 | 11 | 4.4 | |
| 53 | 6 | 1 | 0 | 3 | 0 | 0 | 0 | 10 | 4.0 | |
| 56 | 4 | 0 | 0 | 1 | 0 | 0 | 0 | 5 | 2.0 | |
| 58 | 2 | 1 | 0 | 1 | 1 | 0 | 0 | 5 | 2.0 | |
| 66 | 11 | 2 | 0 | 1 | 0 | 0 | 0 | 14 | 5.6 | |
| 59/68 | 4 | 3 | 0 | 1 | 0 | 0 | 0 | 8 | 3.2 | |
| 69 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 1.2 | |
| 73 | 0 | 1 | 0 | 0 | 0 | 0 | 0 | 1 | 0.4 | |
| Probable high-risk | 34 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0 | 0.0 |
| Low-risk | 6 | 4 | 0 | 0 | 0 | 0 | 0 | 0 | 4 | 1.6 |
| 11 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.4 | |
| 32 | 12 | 1 | 0 | 0 | 1 | 0 | 0 | 14 | 5.6 | |
| 40 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 1.2 | |
| 42 | 3 | 0 | 0 | 1 | 0 | 0 | 0 | 4 | 1.6 | |
| 43 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.4 | |
| 44 | 3 | 0 | 0 | 0 | 0 | 0 | 0 | 3 | 1.2 | |
| 54 | 13 | 0 | 0 | 0 | 0 | 0 | 0 | 13 | 5.2 | |
| 70 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0.8 | |
| 72 | 2 | 0 | 0 | 0 | 0 | 0 | 0 | 2 | 0.8 | |
| 81/87 | 1 | 1 | 0 | 1 | 0 | 0 | 0 | 3 | 1.2 | |
| 84 | 1 | 0 | 0 | 0 | 0 | 0 | 0 | 1 | 0.4 | |
| Other | 15 | 2 | 1 | 0 | 0 | 0 | 0 | 18 | 7.2 | |
| Total | 131 | 20 | 6 | 16 | 11 | 4 | 0 | 261 | ||
| Sequencing analysis | SurePath |
PPV (%) | NPV (%) | |||||||
|---|---|---|---|---|---|---|---|---|---|---|
| NILM | ASC-US | ASC-H | LSIL | HSIL | SCC | AGUS/AC | Total | |||
| HPV (+) | 77 | 12 | 5 | 9 | 9 | 4 | 0 | 116 | 33.6 (39/116) | 98.5 (131/133) |
| HPV (-) | 131 | 0 | 1 | 0 | 1 | 0 | 0 | 133 | ||
| Total | 208 | 12 | 6 | 9 | 10 | 4 | 0 | 249 | ||
| HPV (+) rate (%) | 37.0 | 100.0 | 83.3 | 100.0 | 90.0 | 100.0 | 0.0 | 46.6 | ||
| Reference standard (sequencing analysis) |
||||||
|---|---|---|---|---|---|---|
| HPV (+) | HPV (-) | Total | PPV | NPV | ||
| REBA HPV-ID | HPV (+) | 94 | 7 | 101 | 93.1% | - |
| HPV (-) | 0 | 126 | 126 | - | 100.0% | |
| Total | 94 | 133 | 227 | - | - | |
| Sensitivity 100.0% | Specificity 94.7% | - | - | - | ||
| MolecuTech REBA HPV-ID result | Sequencing analysis result |
Prevalence (n = 227, %) | |||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| High-risk |
Low-risk |
Other type | HPV (-) | Total | |||||||||||||||||||||||||||||||
| 16 | 18 | 26 | 31 | 33 | 35 | 39 | 45 | 51 | 52 | 53 | 56 | 58 | 66 | 59/68 | 69 | 73 | 34 | 6 | 11 | 32 | 40 | 42 | 43 | 44 | 54 | 70 | 72 | 84 | 81/87 | ||||||
| High-risk | 16 | 17 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | 18 | 7.2 |
| 18 | - | 5 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | 6 | 2.4 | |
| 26 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 31 | - | - | - | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 0.4 | |
| 33 | - | - | - | - | 4 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 4 | 1.6 | |
| 35 | - | - | - | - | - | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 0.4 | |
| 39 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 2 | 2 | 0.8 | |
| 45 | - | - | - | - | - | - | - | 7 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 8 | 3.2 | |
| 51 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 52 | - | - | - | - | - | - | - | - | - | 5 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | 6 | 2.4 | |
| 53 | - | - | - | - | - | - | - | - | - | - | 5 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | - | 6 | 2.4 | |
| 56 | - | - | - | - | - | - | - | - | - | 1 | - | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 2 | 0.8 | |
| 58 | - | - | - | - | - | - | - | - | - | - | - | - | 5 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 5 | 2.0 | |
| 66 | - | - | - | - | - | - | - | - | - | - | - | - | - | 3 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 1 | 5 | 2.0 | |
| 59/68 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 69 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 0.4 | |
| 73 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 34 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| Low-risk | 6 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 |
| 11 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 32 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | - | - | - | - | - | - | - | - | 2 | - | 3 | 1.2 | |
| 40 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | - | - | - | - | - | - | - | 1 | - | 2 | 0.8 | |
| 42 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 43 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 44 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 0 | 0.0 | |
| 54 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 4 | - | - | - | - | 2 | - | 6 | 2.4 | |
| 70 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | - | - | - | - | - | 1 | 0.4 | |
| 72 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 2 | - | - | - | - | 2 | 0.8 | |
| 84 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 1 | 1 | - | - | 2 | 0.8 | |
| 81/87 | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 2 | - | - | 2 | 0.8 | |
| Other type | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 15 | 3 | 18 | 7.2 | |
| HPV (-) | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | 126 | 126 | 50.6 | |
| Total | 17 | 5 | 0 | 1 | 4 | 1 | 0 | 7 | 0 | 6 | 5 | 1 | 5 | 3 | 0 | 1 | 0 | 0 | 0 | 0 | 1 | 1 | 0 | 0 | 0 | 4 | 1 | 2 | 1 | 4 | 24 | 133 | 227 | - | |
Values are presented as number (%). NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma; PPV, positive predictive value; NPV, negative predictive value.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma.
HPV, human papillomavirus; NILM, negative for intraepithelial lesion or malignancy; ASC-US, atypical squamous cell of undetermined significance; ASC-H, atypical squamous cell cannot exclude high grade lesion; LSIL, low grade squamous intraepithelial lesion; HSIL, high grade squamous intraepithelial lesion; SCC, squamous cell carcinoma; AGUS, atypical glandular cells of undetermined significance; AC, adenocarcinoma; PPV, positive predictive value; NPV, negative predictive value.
HPV, human papillomavirus; PPV, positive predictive value; NPV, negative predictive value.
HPV, human papillomavirus.

 E-submission
E-submission